Summary
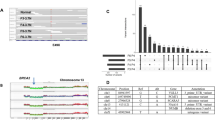
We describe a survey of genetic changes by comparative genomic hybridization (CGH) in 11 human breast cancer cell lines recently established in our laboratory. The most common gains took place at 8q (73%), 1q (64%), 7q (64%), 3q (45%) and 7p (45%), whereas losses were most frequent at Xp (54%), 8p (45%), 18q (45%) and Xq (45%). Many of the cell lines displayed prominent, localized DNA amplifications by CGH. One-third of these loci affected breast cancer oncogenes, whose amplifications were validated with specific probes: 17q12 (two cell lines with ERBB2 amplifications), 11q13 (two with cyclin-D1), 8p11–p12 (two with FGFR1) and 10q25 (one with FGFR2). Gains and amplifications affecting 8q were the most common genetic alterations in these cell lines with the minimal, common region of involvement at 8q22–q23. No high-level MYC (at 8q24) amplifications were found in any of the cell lines. Two-thirds of the amplification sites took place at loci not associated with established oncogenes, such as 1q41–q43, 7q21–q22, 7q31, 8q23, 9p21–p23, 11p12–p14, 15q12–q14, 16q13–q21, 17q23, 20p11–p12 and 20q13. Several of these locations have not been previously reported and may harbour important genes whose amplification is selected for during cancer development. In summary, this set of breast cancer cell lines displaying prominent DNA amplifications should facilitate discovery and functional analysis of genes and signal transduction pathways contributing to breast cancer development.
Similar content being viewed by others
Article PDF
Change history
16 November 2011
This paper was modified 12 months after initial publication to switch to Creative Commons licence terms, as noted at publication
References
Adnane, J, Gaudray, P, Dionne, CA, Crumley, G, Jaye, M, Schlessinger, J, Jeanteur, P, Bimbaum, D & Theillet, C (1991). BEK and FLG, two receptors to members of the FGF family, are amplified in subsets of human breast cancers. Oncogene 6: 659–663.
Anzick, SL, Kononen, J, Walker, RL, Azorsa, DO, Tanner, MM, Guan, XY, Sauter, G, Kallioniemi, OP, Trent, JM & Meltzer, PS (1997). AIB1, a steroid receptor coactivator amplified in breast and ovarian cancer. Science 277: 965–968.
Bischoff, JR, Anderson, L, Zhu, Y, Mossie, K, Ng, L, Souza, B, Schryver, B, Flanagan, P, Clairvoyant, F, Ginther, C, Chan, CS, Novotny, M, Slamon, DJ & Plowman, GD (1998). A homologue of Drosophila aurora kinase is oncogenic and amplified in human colorectal cancers. EMBO J 17: 3052–3065.
Collins, C, Rommens, JM, Kowbel, D, Godfrey, T, Tanner, M, Hwang, SI, Polikoff, D, Nonet, G, Cochran, J, Myambo, K, Jay, KE, Froula, J, Cloutier, T, Kuo, WL, Yaswen, P, Dairkee, S, Giovanola, J, Hutchinson, GB, Isola, J, Kallioniemi, OP, Palazzolo, M, Martin, C, Ericsson, C, Pinkel, D, Albertson, D, Li, W-B & Gray, JW (1998). Positional cloning of ZNF217 and NABC1: genes amplified at 20q13.2 and overexpressed in breast carcinoma. Proc Natl Acad Sci USA 95: 8703–8708.
Devilee, P & Cornelisse, CJ (1994). Somatic genetic changes in human breast cancer. Biochem et Biophys Acta 1198: 113–130.
Ethier, SP (1996). Human breast cancer cell lines as models of growth regulation and disease progression. J Mamm Gland Biol Neoplas 1: 111–121.
Ethier, SP, Summerfelt, RM, Cundiff, KC & Asch, BB (1990). The influence of growth factors on the proliferative potential of normal and primary breast cancer-derived human breast epithelial cells. Breast Cancer Res Treat 17: 221–230.
Ethier, SP, Mahacek, ML, Gullick, WJ, Frank, TJ & Weber, BL (1993). Differential isolation of normal luminal mammary epithelial cells and breast cancer cells from primary and metastatic sites using selective media. Cancer Res 53: 627–635.
Fischer, U, Muller, HW, Sattler, HP, Feiden, K, Zang, KD & Meese, E (1995). Amplification of MET in gliomas. Genes Chromosomes Cancer 12: 63–65.
Flanagan, L, VanWeelden, K, Ammerman, C, Ethier, SP & Welsh, J (1999). SUM-159PT cells: a novel estrogen independent human breast cancer model system. Breast Cancer Res Treat, (in press)
Forozan, F, Karhu, R, Kononen, J, Kallioniemi, A & Kallioniemi, OP (1997). Genome screening by comparative genomic hybridization. Trends Genet 13: 405–409.
Garcia, R, Yu, CL, Hudnall, A, Nelson, KL, Smithgall, T, Fujita, DJ, Ethier, SP & Jove, R (1997). Constitutive activation of STAT3 in fibroblasts transformed by diverse oncoproteins and in breast carcinoma cells. Cell Growth Diff 8: 1267–1276.
Geradts, J & Wilson, PA (1996). High frequency of aberrant p16 (INK4A) expression in human breast cancer. Am J Pathol 149: 15–20.
Guan, XY, Meltzer, PS, Dalton, WS & Trent, JM (1994). Identification of cryptic sites of DNA sequence amplification in human breast cancer by chromosome microdissection. Nat Genet 8: 155–161.
Houldsworth, J, Cordon-Cardo, C, Ladanyi, M, Kelsen, DP & Chaganti, RS (1990). Gene amplification in gastric and esophageal adenocarcinomas. Cancer Res 50: 6417–6422.
Ignatoski, KM & Ethier, SP (1999). Constitutive activation of pp125fak in newly isolated human breast cancer cell lines. Breast Cancer Res Treat, (in press)
Kallioniemi, A, Kallioniemi, OP, Piper, J, Tanner, M, Stokke, T, Chen, L, Smith, HS, Pinkel, D, Gray, JW & Waldman, FM (1994). Detection and mapping of amplified DNA sequences in breast cancer by comparative genomic hybridization. Proc Natl Acad Sci USA 91: 2156–2160.
Karhu, R, Kähkönen, M, Kuukasjärvi, T, Pennanen, S, Tirkkonen, M & Kallioniemi, O (1997). Quality control of CGH: impact of metaphase chromosomes and the dynamic range of hybridization. Cytometry 28: 198–205.
Kononen, J, Bubendorf, L, Kallioniemi, A, Barlund, M, Schraml, P, Leighton, S, Torhorst, J, Mihatsch, MJ, Sauter, G & Kallioniemi, O-P (1998). Tissue microarray for high-throughput molecular profiling of tumor specimens. Nature Med 4: 844–847.
Kuukasjarvi, T, Karhu, R, Tanner, M, Kahkonen, M, Schaffer, A, Nupponen, N, Pennanen, S, Kallioniemi, A, Kallioniemi, OP & Isola, J (1997). Genetic heterogeneity and clonal evolution underlying development of asynchronous metastasis in human breast cancer. Cancer Res 57: 1597–1604.
Mahacek, ML, Beer, DG, Frank, TS & Ethier, SP (1993). Finite proliferative lifespan in vitro of a human breast cancer cell strain isolated from a metstatic lymph node. Breast Cancer Res Treat 28: 267–276.
Ram, TG, Kokeny, KE, Dilts, CA & Ethier, SP (1995). Mitogenic activity of neu differentiation factor/heregulin mimics that of epidermal growth factor and insulin-like growth factor-I in human mammary epithelial cells. J Cell Physiol 163: 589–596.
Ram, TG, Dilts, CA, Dziubinski, ML, Pierce, LJ & Ethier, SP (1996). Insulin-like growth factor and epidermal growth factor independence in human mammary carcinoma cells with c-erbB-2 gene amplification and progressively elevated levels of tyrosine phosphorylated erbB-2. Mol Carcinogen 15: 227–238.
Ried, T, Just, KE, Holtgreve-Grez, H, du Manoir, S, Speicher, MR, Schrock, E, Latham, C, Blegen, H, Zetterberg, A, Cremer, T & Auer, G (1995). Comparative genomic hybridization of formalin-fixed, paraffin-embedded breast tumors reveals different patterns of chromosomal gains and losses in fibroadenomas and diploid and aneuploid carcinomas. Cancer Res 55: 5415–5423.
Sartor, CI, Dziubinski, ML, Yu, CL, Jove, R & Ethier, SP (1997). Role of epidermal growth factor receptor and STAT3 activation in autonomous proliferation of SUM-102PT human breast cancer cells. Cancer Res 57: 978–987.
Sen, S, Zhou, H & White, RA (1997). A putative serine/threonine kinase encoding gene BTAK on chromosome 20q13 is amplified and overexpressed in human breast cancer cell lines. Oncogene 14: 2195–2200.
Tanner, M, Tirkkonen, M, Kallioniemi, A, Collins, C, Stokke, T, Karhu, R, Kowbel, D, Shadrawan, F, Hintz, M, Kuo, WL, Waldman, F, Kallioniemi, A, Isola, J & Kallioniemi, OP (1994). Increased copy number at 20q13 in breast cancer: defining the critical region and exclusion of candidate genes. Cancer Res 54: 2457–2460.
Tanner, MM, Tirkkonen, M, Kallioniemi, A, Isola, J, Kuukasjarvi, T, Collins, C, Kowbel, D, Guan, XY, Trent, J, Gray, JW, Meltzer, P & Kallioniemi, OP (1996). Independent amplification and frequent co-amplification of three nonsyntenic regions on the long arm of chromosome 20 in human breast cancer. Cancer Res 56: 3441–3445.
Tirkkonen, M, Tanner, M, Karhu, R, Kallioniemi, A, Isola, J & Kallioniemi, OP (1998). Molecular cytogenetics of primary breast cancer by CGH. Genes Chromosomes Cancer 21: 177–184.
Wullich, B, Muller, HW, Fischer, U, Zang, KD & Meese, E (1993). Amplified met gene linked to double minutes in human glioblastomas. Eur J Cancer 29A: 1991–1995.
Author information
Authors and Affiliations
Rights and permissions
From twelve months after its original publication, this work is licensed under the Creative Commons Attribution-NonCommercial-Share Alike 3.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-sa/3.0/
About this article
Cite this article
Forozan, F., Veldman, R., Ammerman, C. et al. Molecular cytogenetic analysis of 11 new breast cancer cell lines. Br J Cancer 81, 1328–1334 (1999). https://doi.org/10.1038/sj.bjc.6695007
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.bjc.6695007
Keywords
This article is cited by
-
ADAM8 is expressed widely in breast cancer and predicts poor outcome in hormone receptor positive, HER-2 negative patients
Cancer Cell International (2023)
-
Comparative transcriptional analyses of preclinical models and patient samples reveal MYC and RELA driven expression patterns that define the molecular landscape of IBC
npj Breast Cancer (2022)
-
Robust inflammatory breast cancer gene signature using nonparametric random forest analysis
Breast Cancer Research (2021)
-
TGF-β-induced DACT1 biomolecular condensates repress Wnt signalling to promote bone metastasis
Nature Cell Biology (2021)
-
Development and implementation of the SUM breast cancer cell line functional genomics knowledge base
npj Breast Cancer (2020)