Abstract
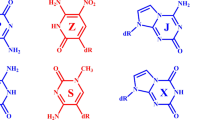
TRIPLE helices result from interaction between single- and double-stranded nucleic acids. Their formation is a possible mechanism for recombination of homologous gene sequences in nature and provides, inter alia, a basis for artificial control of gene activity. Triple-helix motifs have been extensively studied by a variety of techniques, but few high-resolution structural data are available. The only triplet structures characterized so far by X-rav diffraction were in protein–DNA complexes1,2 studied at about 3 Å resolution. We report here the X-ray analysis of a DNA nonamer, d(GCGAATTCG), to a resolution of 2.05 Å, in which the extended crystal structure contains (C · G)*G triplets as a fragment of triple helix. The guanosine-containing chains are in a parallel orientation. This arrangement is a necessary feature of models for homologous recombination which results ultimately in replacement of one length of DNA by another of similar sequence. The present-structure agrees with many published predictions of triplex organization, and provides an accurate representation of an element that allows sequence-specific association between single- and double-stranded nucleic acids.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout
Similar content being viewed by others
References
Schultz, S. C., Shields, G. C. & Steitz, T. A. Science 253, 1002–1007 (1991).
Luisi, B. F. et al. Nature 352, 497–505 (1991).
Ouali, M. et al. J. Am. chem Soc. 115, 4264–4270 (1993).
Piriou, J. M., Ketterlé, C., Gabarro-Arpa, J., Cognet, J. A. H. & Le Bret, M. Biophys. Chem. 50, 323–343 (1994).
Allen, F. H. et al. J. chem. Inf. Comput. Sci. 31, 187–204 (1991).
Zhurkin, V. B., Raghunathan, G., Ulyanov, N. B., Camerini-Otero, R. D. & Jernigan, R. L. J. molec. Biol. 239, 181–200 (1994).
Cheng, Y.-K. & Pettitt, B. M. J. Am. chem. Soc. 114, 4465–4474 (1992).
Dickerson, R. E. et al. EMBO J. 8, 1–4 (1989).
Wing, R. et al. Nature 287, 755–758 (1981).
Navaza, J. Acta crystallogr. A50, 157–163 (1994).
Brünger, A. T., Kuriyan, J. & Karplus, M. Science 235, 458–460 (1987).
Bernstein, F. C. et al. J. molec. Biol. 112, 535–542 (1977).
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Van Meervelt, L., Vlieghe, D., Dautant, A. et al. High-resolution structure of a DNA helix forming (C·G)*G base triplets. Nature 374, 742–744 (1995). https://doi.org/10.1038/374742a0
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/374742a0
This article is cited by
-
Structure and stability of different triplets involving artificial nucleobases: clues for the formation of semisynthetic triple helical DNA
Scientific Reports (2023)
-
The structure of r(UUCGCG) has a 5′-UU-overhang exhibiting Hoogsteen-like trans U•U base pairs
Nature Structural Biology (1996)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.