Abstract
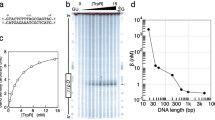
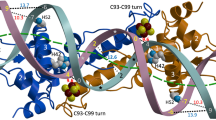
The crystal structure of the met represser–operator complex shows two dimeric repressor molecules bound to adjacent sites 8 base pairs apart on an 18-base-pair DNA fragment. Sequence specificity is achieved by insertion of double-stranded antiparallel protein β-ribbons into the major groove of B-form DNA, with direct hydrogen-bonding between amino-acid side chains and the base pairs. The repressor also recognizes sequence-dependent distortion or flexibility of the operator phosphate backbone, conferring specificity even for inaccessible base pairs.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout
Similar content being viewed by others
References
Steitz, T. A. Q. Rev. Biophys. 23, 205–280 (1990).
Freemont, P. S., Lane, A. N. & Sanderson, M. R. Biochem. J. 278, 1–23 (1991).
Harrison, S. C. & Aggarwal, A. K. A. Rev. Biochem. 59, 933–969 (1990).
Brennan, R. G. Curr. Opin. struct. Biol. 1, 80–88 (1991).
Harrison, S. C. Nature 353, 715–719 (1991).
Jordan, S. R. & Pabo, C. O. Science 242, 893–899 (1988).
Clarke, N. D. et al. Science 254, 267–270 (1991).
Brennan, R. G., Roderick, S. L., Takeda, Y. & Matthews, B. W. Proc. natn. Acad. Sci. U.S.A. 87, 8165–8169 (1990).
Aggarwal, A. K., Rodgers, D. W., Drottar, M., Ptashne, M. & Harrison, S. C. Science 242, 899–907 (1988).
Mondragon, A. & Harrison, S. C. J. molec. Biol. 219, 321–334 (1991).
Luisi, B. F. & Sigler, P. B. Biochim. biophys. Acta 1048, 113–126 (1990).
Otwinowski, Z. et al. Nature 335, 321–329 (1988).
Schultz, S. C., Shields, G. C. & Steitz, T. A. Science 253, 1001–1007 (1991).
Kissinger, C. R. et al. Cell 63, 579–590 (1990).
Wolberger, C. et al. Cell 67, 517–528 (1991).
Pavletich, N. P. & Pabo, C. O. Science 252, 809–817 (1991).
Luisi, B. F. et al. Nature 352, 497–505 (1991).
Old, I. G. et al. Prog. Biophys. molec. Biol. 56, 145–184 (1991).
Rafferty, J. B. et al. Nature 341, 705–710 (1989).
Belfaiza, J. et al. Proc. natn. Acad. Sci. U.S.A. 83, 867–871 (1986).
Phillips, S. E. V. et al. Nature 341, 711–715 (1989).
Davidson, B. E. & Saint-Girons, I. Molec. Microbiol. 3, 1639–1648 (1989).
He, Y. Y. et al. Nature 359, 431–433 (1992).
Saint-Girons, I. et al. J. biol. Chem. 261, 10936–10940 (1986).
Saenger, W. Principles of Nucleic Acid Structure 226 (Springer, New York, 1984).
Rhodes, D. & Klug, A. Nature 286, 573–578 (1980).
Klug, A. et al. J. molec. Biol. 131, 669–680 (1979).
Yoon, C. et al. Proc. natn. Acad. Sci. U.S.A. 85, 6332–6336 (1988).
Quintana, J. R. et al. J. molec. Biol. 225, 379–395 (1992).
Lee, B. & Richards, F. M. J. molec. Biol. 55, 379–400 (1971).
Seeman, N. C., Rosenberg, J. M. & Rich, A. Proc. natn. Acad. Sci. U.S.A. 73, 804–808 (1976).
Drew, H. R. & Travers, A. A. J. molec. Biol. 186, 773–790 (1985).
Breg, J. N. et al. Nature 346, 586–589 (1990).
Brown, B. M., Bowie, J. U. & Sauer, R. T. Biochemistry 29, 11189–11195 (1990).
Knight, K. L. et al. J. biol. Chem. 264, 3639–3642 (1989).
Bowie, J. U. & Sauer, R. T. J. molec. Biol. 211, 5–6 (1990).
Carter, C. W. Jr & Kraut, J. Proc. natn. Acad. Sci. U.S.A. 71, 283–287 (1974).
Church, G. M., Sussman, J. L. & Kim, S.-H. Proc. natn. Acad. Sci. U.S.A. 74, 1458–1462 (1977).
The SERC (UK) Collaborative Computing Project No. 4, a Suite of Programs for Protein Crystallography (Distributed from Daresbury Laboratory, Warrington WA4 4AD, UK, 1979).
Fitzgerald, P. M. D. J. appl. Crystallogr. 21, 273–288 (1988).
Sim, G. A. Acta crystallogr. 13, 511–512 (1960).
Jones, T. A. J. appl. Crystallogr. 11, 268–272 (1978).
Brünger, A. T., Kuriyan, J. & Karplus, M. Science 235, 458–460 (1985).
Tronrud, D. E., Ten Eyck, L. F. & Matthews, B. W. Acta crystallogr. A43, 489–501 (1987).
Priestle, J. P. J. appl. Crystallogr. 21, 572–576 (1988).
Flower, D. R. J. molec. Graphics 9, 257–258 (1991).
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Somers, W., Phillips, S. Crystal structure of the met represser–operator complex at 2.8 Å resolution reveals DNA recognition by β-strands. Nature 359, 387–393 (1992). https://doi.org/10.1038/359387a0
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/359387a0
This article is cited by
-
Unraveling the role of quorum sensing-dependent metabolic homeostasis of the activated methyl cycle in a cooperative population of Burkholderia glumae
Scientific Reports (2019)
-
Exploring comprehensive within-motif dependence of transcription factor binding in Escherichia coli
Scientific Reports (2015)
-
Modeling of the structure and interactions of the B. anthracis antitoxin, MoxX: deletion mutant studies highlight its modular structure and repressor function
Journal of Computer-Aided Molecular Design (2011)
-
THAP proteins target specific DNA sites through bipartite recognition of adjacent major and minor grooves
Nature Structural & Molecular Biology (2010)
-
Segrosome structure revealed by a complex of ParR with centromere DNA
Nature (2007)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.