Abstract
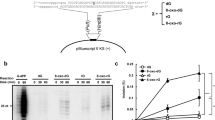
WHEN present in single-stranded DNA, palindromic or quasi-palindromic sequences have the potential to form complex secondary structures, including hairpins, which may facilitate inter-strand misalignment of direct repeats and be responsible for diverse types of replication-based mutations, including deletions, additions, frameshifts and duplications1–5. In regions of palindromic symmetry, specific deletion events may involve the formation of a hairpin or other DNA secondary structures which can stabilize the misalignment of direct repeats1,2. One model suggests that these deletions occur during DNA replication by slippage of the template strand and misalignment with the progeny strand6,7. The concurrent DNA replication model, involving an asymmetric dimeric DNA polymerase III complex which replicates the leading and lagging strands8, has significant implications for mutagenesis. The intermittent looping of the lagging strand template, and the fact that the lagging strand template may contain a region of single-stranded DNA the length of an Okazaki fragment, provides an opportunity for DNA secondary-structure formation and misalignment. Here we report our design of a palindromic fragment to create an 'asymmetric palindromic insert' in the chloram-phenicol acetyltransferase gene of plasmid pBR325. The frequency with which the insert was deleted in Escherichia coli depends on the orientation of the gene in the plasmid. Our results suggest that replication-dependent deletion between direct repeats may occur preferentially in the lagging strand.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout
Similar content being viewed by others
References
Glickman, B. W. & Ripley, L. S. Proc. natn. Acad. Sci. U.S.A. 81, 4046–4050 (1984).
Ripley, L. S. & Glickman, B. W. in Cellular Responses to DNA Damage, UCLA Symposia on Molecular and Cellular Biology, Vol 10 (ed. N. R. Cozzarelli) 521–540 (Liss, Inc. New York, N.Y. 1983).
Drake, J. W., Glickman, B. W. & Ripley, L. S. Am. Scient. 71, 621–630 (1983).
Schaaper, R. M., Danforth, B. N. & Glickman, B. W. J. molec. Biol. 189, 273–284 (1986).
Ripley, L. S. Proc. natn. Acad. Sci. U.S.A. 79, 4128–4132 (1982).
Streisinger, G. et al. Cold Spring Harb. Symp. quant. Biol. 31, 77–84 (1966).
Albertini, A. M., Hofer, M. Calos, M. P. & Miller, J. H. Cell 29, 319–328 (1982).
McHenry, C. S. A. Rev. Biochem. 57, 519–550 (1988).
Scott, J. R. Microbiol. Rev. 48, 1–23 (1984).
Bohr, V. A., Smith, C. A., Okumoto, D. A. & Hanawalt, P. C. Cell 40, 359–369 (1985).
Mellon, I. & Hanawalt, P. C. Nature 342, 95–98 (1989).
Mellon, I., Spivak, G. & Hanawalt, P. C. Cell 51, 241–249 (1988).
Tsurimoto, T. Melendy, T. & Stillman, B. Nature 346, 534–539 (1990).
Roberts, J. D., Thomas, D. C. & Kunkel, T. A. Proc. natn. Acad. Sci. U.S.A. 88, 3465–3469 (1991).
Weaver, D. T. & DePamphilis, M. L. J. molec. Biol. 180, 961–986 (1984).
Duckett, D. R., Murchie, A. I. H. & Lilley, D. M. J. EMBO J. 9, 583–590 (1990).
Duckett, D. R. & Lilley, D. M. J. EMBO J 9, 1659–1664 (1990).
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Trinh, T., Sinden, R. Preferential DNA secondary structure mutagenesis in the lagging strand of replication in E. coli. Nature 352, 544–547 (1991). https://doi.org/10.1038/352544a0
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/352544a0
This article is cited by
-
The association of group IIB intron with integrons in hypersaline environments
Mobile DNA (2021)
-
Spinocerebellar ataxia: an update
Journal of Neurology (2019)
-
Characterization of indels in poxvirus genomes
Virus Genes (2011)
-
Identification of novel intronic BRCA1 variants of uncertain significance in a Thai hereditary breast cancer family
Journal of Genetics (2011)
-
Maintaining genome stability at the replication fork
Nature Reviews Molecular Cell Biology (2010)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.