Abstract
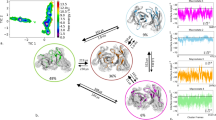
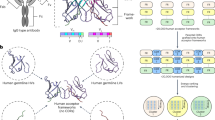
The antigen-combining site of antibody molecules consists of six separate loops supported by a conserved β-sheet framework; antibody specificity arises from length and sequence variation of these 'hypervariable' loops1 and can be manipulated by transferring sets of loops between different frameworks2. Irregular loops are the most difficult parts of protein structure to understand and to model correctly3–6. Here, we describe two computer experiments where all the hypervariable loops were deleted from X-ray structures of mouse immunoglobulins and reconstructed using the conformational search program CONGEN7. A protocol was developed for reconstruction of the hypervariable loops in McPC 603 antibody. Calculated loop conformations were generated and a model of the combining site was built from selected low-energy conformations. We then modelled hypervariable loops in another antibody molecule, HyHEL-5. Both models agreed well with the known crystal structures. Our results hold out promise for the success of future modelling studies of complete antigen-combining sites from amino acid sequences.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout
Similar content being viewed by others
References
Wu, T. T. & Kabat, E. A. J. exp. Med 132, 211–250 (1970).
Jones, P. T., Dear, P. H., Foote, J., Neuberger, M. S. & Winter, G. Nature 321, 522–525 (1986).
Richardson, J. Adv. Protein Chem. 34, 167–339 (1981).
Sibanda, B. L. & Thornton, J. Nature 316, 170–174 (1985).
Rose, G. D., Gierasch, I. M. & Smith, J. A. Adv. Protein Chem. 37, 1–50 (1985).
Leszczynski, J. F. & Rose, G. D. Science 234, 849–855 (1986).
Bruccoleri, R. E. & Karplus, M. Biopolymers 26, 137–168 (1987).
Kabat, E. A. & Wu, T. T. Proc. natn. Acad. Sci. U.S.A. 69, 960–964 (1972).
Chothia, C. et al. Science 233, 755–758 (1986).
de la Paz, P., Sutton, B. J., Darsley, M. J. & Rees, A. R. EMBO J. 5, 415–425 (1986).
Fine, R. M., Wang, H., Shenkin, P. S., Yarmush, D. L. & Levinthal, C. Proteins 1, 342–362 (1986).
Smith-Gill, S. J. et al. J. molec. Biol. 194, 713–724 (1987).
Chothia, C. & Lesk, A. M. J. molec. Biol. 196, 901–917 (1987).
Shih, H. L., Brady, J. & Karplus, M. Proc. natn. Acad. Sci. U.S.A. 82, 1697–1700 (1985).
Snow, M. E. & Amzel, M. L. Proteins 1, 276–279 (1986).
Moult, J. & James, M. N. G. Proteins 1, 146–163 (1986).
Satow, Y., Cohen, G. H., Padlan, E. A. & Davies, D. R. J. molec. Biol. 190, 593–604 (1986).
Gō, N. & Scheraga, H. A. Macromolecules 3, 178–187 (1970).
Bruccoleri, R. E. & Karplus, M. Macromolecules 18, 2767–2773 (1985).
Brooks, B. et al. J. comput. Chem. 4, 187–217 (1983).
Chothia, C., Novotny, J., Bruccoleri, R. E. & Karplus, M. J. molec. Biol. 186, 651–663 (1985).
Novotny, J. & Haber, E. Proc. natn. Acad. Sci. U.S.A. 82, 4592–4596 (1985).
Lee, B. K. & Richards, F. M. J. molec. Biol. 55, 379–400 (1971).
Rashin, A. A. Biopolymers 23, 1605–1620 (1984).
Novotny, J., Bruccoleri, E. R. & Karplus, M. J. molec. Biol. 177, 787–818 (1984).
Eisenberg, D. & McLachlan, A. Nature 319, 199–203 (1986).
Ooi, T., Oobatake, M., Nemethy, G. & Scheraga, H. A. Proc. natn. Acad. Sci. U.S.A. 84, 3086–3090 (1987).
Chou, P. Y. & Fasman, G. Biochemistry 16, 222–244 (1974).
Dyson, H. J. et al. Nature 318, 480–483 (1985).
Pincus, M. R., Klausner, R. D. & Scheraga, H. A. Proc. natn. Acad. Sci. U.S.A. 79, 5107–5110 (1982).
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Bruccoleri, R., Haber, E. & Novotný, J. Structure of antibody hypervariable loops reproduced by a conformational search algorithm. Nature 335, 564–568 (1988). https://doi.org/10.1038/335564a0
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/335564a0
This article is cited by
-
The biology of immunoglobulin free light chains and kidney injury
Kidney International (2011)
-
Identification of a LNCaP-Specific Binding Peptide Using Phage Display
Pharmaceutical Research (2011)
-
Prediction of a common neutralizing epitope of H5N1 avian influenza virus by in silico molecular docking
Science Bulletin (2008)
-
ProRegIn: A regularity index for the selection of native-like tertiary structures of proteins
Journal of Biosciences (2007)
-
Cellular and Behavioral Effects of D2 Dopamine Receptor Hydrophobic Eigenmode-Targeted Peptide Ligands
Neuropsychopharmacology (2003)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.