Abstract
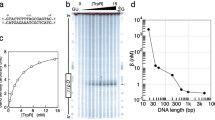
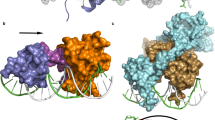
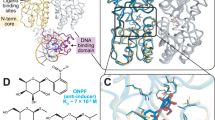
The represser encoded by bacteriophage 434 binds to its operators by inserting a 'recognition' α-helix into the major groove of the DNA1,2. We have identified an amino acid–base pair contact that determines (in part) the DNA-binding specificity of 434 represser. The identification is based on the properties of a 'new-specificity' mutant, named Represser[Ala 28], which bears the substitution of Ala for Gln at the first residue of its recognition α-helix. Represser[Ala 28] binds with high affinity to a particular doubly mutant operator bearing the same substitution at position 1 in each half-site, but does not bind to either the wild-type operator or to other mutant operators. We describe molecular models of residue 28–base pair 1 interactions that account for the binding specificities of both the mutant and wild-type proteins.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout
Similar content being viewed by others
References
Anderson, J. E., Ptashne, M. & Harrison, S. C. Nature 316, 596–601 (1985).
Wharton, R. P. & Ptashne, M. Nature 316, 601–605 (1985).
Miller, J. H. Experiments in Molecular Genetics (Cold Spring Harbor Laboratory, New York, 1972).
Anderson, J. E., Ptashne, M. & Harrison, S. C. Nature 326, 846–852 (1987).
Seeman, N. C., Rosenberg, J. M. & Rich, A. Proc. natn. Acad. Sci. U.S.A. 73, 804–808 (1976).
Fersht, A. R., Shindler, J. S. & Tsui, W.-C. Biochemistry 19, 5520–5524 (1980).
Goeddel, D. V., Yansura, D. G. & Caruthers, M. H. Proc. natn. Acad. Sci. U.S.A. 75, 3578–3582 (1978).
Fersht, A. R. et al. Nature 314, 235–238 (1985).
Schevitz, R. W. et al. Nature 317, 782–786 (1985).
Lewis, M. et al. Cold Spring Harb. Symp. quant. Biol. 47, 435–440 (1983).
Ohlendorf, D. H., Anderson, W. F., Fisher, R. G., Takeda, Y. & Matthews, B. W. Nature 298, 718–723 (1982).
Hochschild, A., Douhan, J. III & Ptashne, M. Cell 47, 807–816 (1986).
Hochschild, A. & Ptashne, M. Cell 44, 925–933 (1986).
McClarin, J. A. et al. Science 234, 1526–1541 (1986).
Koudelka, G. B., Harrison, S. C. & Ptashne, M. Nature 326, 886–881 (1987).
Wharton, R. P. thesis, Harvard Univ. (1985).
Youderian, P., Vershon, A., Bouvier, S., Sauer, R. T. & Susskind, M. M. Cell 35, 777–783 (1983).
Yu, X.-M. & Reznikoff, W. S. Nucleic Acids Res. 12, 5449–5464 (1984).
Benson, N., Sugiono, P., Bass, S., Mendelman, L. V. & Youderian, P. Genetics 114, 1–14 (1986).
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Wharton, R., Ptashne, M. A new-specificity mutant of 434 repressor that defines an amino acid–base pair contact. Nature 326, 888–891 (1987). https://doi.org/10.1038/326888a0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/326888a0
This article is cited by
-
Selection and design of high affinity DNA ligands for mutant single-chain derivatives of the bacteriophage 434 repressor
Science in China Series C: Life Sciences (2001)
-
Dramatic changes in DNA-binding specificity caused by single residue substitutions in an Arc/Mnt hybrid repressor
Nature Structural & Molecular Biology (1995)
-
The possible roles of residues 79 and 80 of the Trp repressor from Escherichia coli K-12 in trp operator recognition
Molecular and General Genetics MGG (1995)
-
Identification of an amino acid–base contact in the GCN4–DNA complex by bromouracil-mediated photocrosslinking
Nature (1992)
-
DNA twisting and the effects of non-contacted bases on affinity of 434 operator for 434 represser
Nature (1992)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.