Abstract
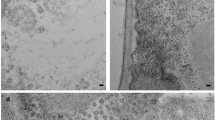
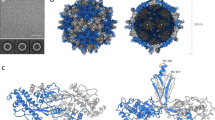
SMALL quasi-isometric particles, mostly occurring in pairs, have been found in, and purified from extracts of plants infected by maize streak1, beet curly top2, tomato golden mosaic3, euphorbia mosaic3, bean golden (yellow) mosaic3–5, cassava latent6 and cassava brown streak viruses6. Individual particles are 15–20 nm in diameter—unusually small for a virus—and in electron micrographs many of the individual particles in the pairs have a five-sided outline in which the contiguous sides seem longer than the others. The pairs of particles of maize streak and cassava latent viruses have sedimentation coefficients of about 76S (refs 1 and 7) and preparations of each yield a single polypeptide species, estimated at about 28,000 and 34,000 daltons respectively7. Their nucleic acid can be resolved into two components by electrophoresis7, and in maize streak virus it has been identified tentatively as RNA on the basis of sensitivity to ribonuclease1. We now report, however, that the nucleic acids of cassava latent and maize streak viruses consist of single-stranded, predominantly circular DNA of molecular weight less than 106.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout
Similar content being viewed by others
References
Bock, K. R., Guthrie, E. J. & Woods, R. D. Ann. appl. Biol. 77, 289–296 (1974).
Mumford, D. L. Phytopathology 64, 136–139 (1974).
Matyis, J. C., Silva, D. M., Oliveira, A. R. & Costa, A. S. Summa Phytopathologica 1, 267–274 (1975).
Galvez, G. E. & Castano, M. Turrialba 26, 205–207 (1976).
Goodman, R. M., Bird, J. & Thongmeearkom, P. Phytopathology 67, 37–42 (1977).
Bock, K. R. & Guthrie, E. J. in African Cassava Mosaic (ed. Nestel, B. L.) 11–16 (International Development Research Centre, Ottawa, 1976).
Bock, K. R., Guthrie, E. J., Meredith, G. & Barker, H. Ann. appl. Biol. 85, 305–308 (1977).
Goodman, R. M. Nature 266, 54–55 (1977).
Rose, J. A., Berns, K. I., Hoggan, M. D. & Koczot, F. J. Proc. natn. Acad. Sci. U.S.A. 64, 863–869 (1969).
Espejo, R. T., Canelo, E. S. & Sinsheimer, R. L. Proc. natn. Acad. Sci. U.S.A. 63, 1164–1168 (1969).
Zuccarelli, A. J., Benbow, R. M. & Sinsheimer, R. L. Proc. natn. Acad. Sci. U.S.A. 69, 1905–1910 (1972).
Davis, R. W., Simon, M. & Davidson, N. Methods in Enzymology (eds Grossman, L. & Moldave, K.) 21 D, 413–428 (Academic Press, New York and London, 1971).
Sinsheimer, R. L. J. mol. Biol. 1, 43–53 (1959).
Dingman, C. W., Fisher, M. P. & Kakefuda, T. Biochemistry 11, 1242–1250 (1972).
Storey, H. H. Ann. appl. Biol. 12, 422–439 (1925).
Goodman, R. M. Virology 83, 171–179 (1977).
Murant, A. F., Mayo, M. A., Harrison, B. D. & Goold, R. A. J. gen. Virol. 16, 327–338 (1972).
Biddlecombe, W. H. et al. J. Physiol. (in the press).
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
HARRISON, B., BARKER, H., BOCK, K. et al. Plant viruses with circular single-stranded DNA. Nature 270, 760–762 (1977). https://doi.org/10.1038/270760a0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/270760a0
This article is cited by
-
Molecular Epidemiology of Begomoviruses Infecting Mungbean from Yellow Mosaic Disease Hotspot Regions of India
Applied Biochemistry and Biotechnology (2023)
-
Infectivity and genetic recombination of yellow mosaic viruses associated with blackgram (Vigna mungo L. Hepper) in South India
Australasian Plant Pathology (2023)
-
Viral metagenomics revealed diverse CRESS-DNA virus genomes in faeces of forest musk deer
Virology Journal (2020)
-
Genome wide molecular evolution analysis of begomoviruses reveals unique diversification pattern in coat protein gene of Old World and New World viruses
VirusDisease (2019)
-
ssDNA viruses: key players in global virome
VirusDisease (2019)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.