Abstract
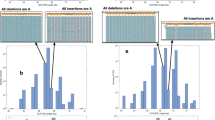
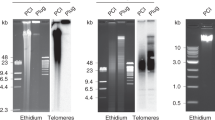
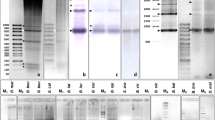
MOST eukayrote genomes contain some highly repeated DNA sequences. DNA hybridisation kinetic studies indicate that the smallest repeating unit of highly repetitious DNA is around 300 base pairs1 but direct sequence analysis of highly repeated DNA has demonstrated a simple basic repeating unit of 4–12 nucleotides2–5. It has been postulated that the repeated sequences have been generated by successive replications of some primary simple sequence1, 6, 7. The discrepancy between sequence data and kinetic analysis may be explained by noting that the rate of reassociation is a function of length8; this data is consistent with a model of internal heterogeneity in the repeated, simple DNA sequence units9, 10. Although the hybridisation and sequence analyses have been successful in establishing certain features of reiteirated DNA, these methods are not well suited to detect and study infrequent but regularly repeated DNA sequences interdigitated within the dominant short repeating units. The existence and nature of such interdigitated repeats are important to our understanding of the mechanism by which repeated DNA is generated. We have used site-specific endonucleases (restriction enzymes) to show that mutational events within simple sequence DNA may have been coupled with cycles of discrete replications.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout
Similar content being viewed by others
References
Britten, R. J., and Kohne, D. E., Science, 161, 529–540 (1968).
Southern, E. M., Nature, 227, 794–798 (1970).
Fry, K., Poon, R., Whitcome, P., Idriss, J., Salser, W., Mazrimas, J., and Hatch, F., Proc. natn. Acad. Sci. U.S.A., 70, 2642–2646 (1973).
Gall, J. G., and Atherton, D. D., J. molec. Biol., 85, 633–664 (1974).
Skinner, D. M., Beattie, W. G., Blattner, F. R., Stark, B. P., and Dahlberg, J. E., Biochemistry, 13, 3930–3937 (1974).
Walker, P. M. B., Prog. Biophys. molec. Biol., 23, 145–190 (1971).
Sutton, W. D., and McCallum, M., J. molec. Biol., 71, 633–656 (1972).
Hutton, J. R., and Wetmur, J. G., Biochem. biophys. Res. Commun., 52, 1148–1155 (1973).
Southern, E. M., Nature new Biol., 232, 82–83 (1971).
Sutton, W. D., and McCallum, M., Nature new Biol., 232, 83–85 (1971).
Kelly, T. J., and Smith, H. O., J. molec. Biol., 51, 393–409 (1970).
Landy, A., Ruidisueli, E., Robinson, L., Foeller, C., and Ross, W., Biochemistry, 13, 2134–2142 (1974).
Mowbray, S. L., and Landy, A., J. Cell Biol., 59, 237a (1973).
Mowbray, S. L., and Landy, A., Proc. natn. Acad. Sci. U.S.A., 71, 1920–1924 (1942).
Philippsen, P., Streeck, R. E., and Zachau, H. G., Eur. J. Biochem., 45, 479–488 (1974).
Botchan, M. R., Nature, 251, 288–292 (1974).
Middleton, J. H., Edgell, M. H., and Hutchison, C. A., J. Virol., 10, 42–50 (1972).
Hedgepeth, J., Goodman, H. M., and Boyer, H. W., Proc. natn. Acad. Sci U.S.A., 69, 3448–3452 (1972).
Roberts, R. J., Arrand, J. R., and Keller, W., Proc. natn. Acad. Sci. U.S.A., 71, 3829–3833 (1974).
Takanami, M., Methods in Molecular Biology (in the press).
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
MOWBRAY, S., GERBI, S. & LANDY, A. Interdigitated repeated sequences in bovine satellite DNA. Nature 253, 367–370 (1975). https://doi.org/10.1038/253367a0
Received:
Revised:
Issue Date:
DOI: https://doi.org/10.1038/253367a0
This article is cited by
-
Analysis of bovidae genomes: Arrangement of repeated and single copy DNA sequences in bovine, goat and sheep
Journal of Biosciences (1982)
-
The sequence organization of bovine DNA
Chromosoma (1980)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.