Abstract
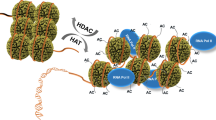
CREB-binding protein (CBP) and p300 are highly conserved and functionally related transcription coactivators and histone/protein acetyltransferases. They are tumor suppressors, participate in a wide variety of physiological events, and serve as integrators among different signal transduction pathways. In this study, 11 distinct proteins that have a high degree of homology with the amino acid sequence of p300 have been identified in current protein databases. All of these 11 proteins belong to either animal or plant multicellular organisms (higher eucaryotes). Conservation of p300/CBP domains among these proteins was examined further by sequence alignment and pattern search. The domains of p300/CBP that are required for the HAT function, including PHD, putative CoA-binding, and ZZ domains, are conserved in all of these 11 proteins. This observation is consistent with the previous functional assays and indicates that they are a family of acetyltransferases, i.e. p300/CBP acetyltransferases (PCAT). TAZ domains (TAZ1 and/or TAZ2) of PCAT proteins may allow them to participate in transcription regulation by either directly recruiting transcription factors, acetylating them subsequently, or directing targeted acetylation of nucleosomal histones.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Aasland R, Gibson TJ, Stewart AF . 1995 Trends Biochem. Sci. 20: 56–59
Abraham SE, Lobo S, Yaciuk P, Wang HG, Moran E . 1993 Oncogene 8: 1639–1647
Adams MD, Celniker SE, Holt RA, Evans CA, Gocayne JD, Amanatides PG, Scherer SE, Li PW, Hoskins RA, Galle RF, George RA, Lewis SE, Richards S, Ashburner M, Henderson SN, Sutton GG, Wortman JR, Yandell MD, Zhang Q, Chen LX, Brandon RC, Rogers YH, Blazej RG, Champe M, Pfeiffer BD, Wan KH, Doyle C, Baxter EG, Helt G, Nelson CR, Gabor GL, Abril JF, Agbayani A, An HJ, Andrews-Pfannkoch C, Baldwin D, Ballew RM, Basu A, Baxendale J, Bayraktaroglu L, Beasley EM, Beeson KY, Benos PV, Berman BP, Bhandari D, Bolshakov S, Borkova D, Botchan MR, Bouck J, Brokstein P, Brottier P, Burtis KC, Busam DA, Butler H, Cadieu E, Center A, Chandra I, Cherry JM, Cawley S, Dahlke C, Davenport LB, Davies P, de Pablos B, Delcher A, Deng Z, Mays AD, Dew I, Dietz SM, Dodson K, Doup LE, Downes M, Dugan-Rocha S, Dunkov BC, Dunn P, Durbin KJ, Evangelista CC, Ferraz C, Ferriera S, Fleischmann W, Fosler C, Gabrielian AE, Garg NS, Gelbart WM, Glasser K, Glodek A, Gong F, Gorrell JH, Gu Z, Guan P, Harris M, Harris NL, Harvey D, Helman TJ, Hernandez JR, Houck J, Hostin D, Houston KA, Howland TJ, Wei MH, Ibegwam C, Jalali M, Kalush F, Karpen GH, Ke Z, Kennison JA, Ketchum KA, Kimmel BE, Kodira CD, Kraft C, Kravitz S, Kulp D, Lai Z, Lasko P, Lei Y, Levitsky AA, Li J, Li Z, Liang Y, Lin X, Liu X, Mattei B, McIntosh TC, McLeod MP, McPherson D, Merkulov G, Milshina NV, Mobarry C, Morris J, Moshrefi A, Mount SM, Moy M, Murphy B, Murphy L, Muzny DM, Nelson DL, Nelson DR, Nelson KA, Nixon K, Nusskern DR, Pacleb JM, Palazzolo M, Pittman GS, Pau S, Pollard J, Puri V, Reese MG, Reinert K, Remington K, Saunders RD, Scheeler F, Shen H, Shue BC, Siden-Kiamos I, Simpson M, Skupski MP, Smith T, Spier E, Spradling AC, Stapleton M, Strong R, Sun E, Svirskas R, Tector C, Turner R, Venter E, Wang AH, Wang X, Wang ZY, Wassarman DA, Weinstock GM, Weissenbach J, Williams SM, Woodage T, Worley KC, Wu D, Yang S, Yao QA, Ye J, Yeh RF, Zaveri JS, Zhan M, Zhang G, Zhao Q, Zheng L, Zheng XH, Zhong FN, Zhong W, Zhou X, Zhu S, Zhu X, Smith HO, Gibbs RA, Myers EW, Rubin GM, Venter JC . 2000 Science 287: 2185–2195
Akimaru H, Chen Y, Dai P, Hou DX, Nonaka M, Smolik SM, Armstrong S, Goodman RH, Ishii S . 1997a Nature 386: 735–738
Akimaru H, Hou DX, Ishii S . 1997b Nat. Genet. 17: 211–214
Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ . 1990 J. Mol. Biol. 215: 403–410
Arany Z, Sellers WR, Livingston DM, Eckner R . 1994 Cell 77: 799–800
Bannister AJ, Kouzarides T . 1996 Nature 384: 641–643
Bordoli L, Netsch M, Luthi U, Lutz W, Eckner R . 2001 Nucleic Acids Res. 29: 589–597
Boyes J, Byfield P, Nakatani Y, Ogryzko V . 1998 Nature 396: 594–598
Chakravarti D, LaMorte VJ, Nelson MC, Nakajima T, Schulman IG, Juguilon H, Montminy M, Evans RM . 1996 Nature 383: 99–103
Chrivia JC, Kwok RP, Lamb N, Hagiwara M, Montminy MR, Goodman RH . 1993 Nature 365: 855–859
Clements A, Rojas JR, Trievel RC, Wang L, Berger SL, Marmorstein R . 1999 EMBO J. 18: 3521–3532
Combet C, Blanchet C, Geourjon C, Deleage G . 2000 Trends Biochem. Sci. 25: 147–150
Corpet F . 1988 Nucleic Acids Res. 16: 10881–10890
Cress WD, Seto E . 2000 J. Cell. Physiol. 184: 1–16
Deng L, de la Fuente C, Fu P, Wang L, Donnelly R, Wade JD, Lambert P, Li H, Lee CG, Kashanchi F . 2000 Virology 277: 278–295
Dhalluin C, Carlson JE, Zeng L, He C, Aggarwal AK, Zhou MM . 1999 Nature 399: 491–496
Eckner R, Arany Z, Ewen M, Sellers W, Livingston DM . 1994a Cold Spring Harb. Symp. Quant. Biol. 59: 85–95
Eckner R, Ewen ME, Newsome D, Gerdes M, DeCaprio JA, Lawrence JB, Livingston DM . 1994b Genes Dev. 8: 869–884
Fontes JD, Kanazawa S, Jean D, Peterlin BM . 1999 Mol. Cell. Biol. 19: 941–947
Giles RH, Petrij F, Dauwerse HG, den Hollander AI, Lushnikova T, van Ommen GJ, Goodman RH, Deaven LL, Doggett NA, Peters DJ, Breuning MH . 1997 Genomics 42: 96–114
Giordano A, Avantaggiati ML . 1999 J. Cell. Physiol. 181: 218–230
Goodman RH, Smolik S . 2000 Genes Dev. 14: 1553–1577
Goodrich JA, Tjian R . 1994 Curr. Opin. Cell. Biol. 6: 403–409
Gu W, Roeder RG . 1997 Cell 90: 595–606
Harlow E, Whyte P, Franza BR, Schley C . 1986 Mol. Cell. Biol. 6: 1579–1589
Imhof A, Yang XJ, Ogryzko VV, Nakatani Y, Wolffe AP, Ge H . 1997 Curr. Biol. 7: 689–692
Kamei Y, Xu L, Heinzel T, Torchia J, Kurokawa R, Gloss B, Lin SC, Heyman RA, Rose DW, Glass CK, Rosenfeld MG . 1996 Cell 85: 403–414
Karplus K, Barrett C, Hughey R . 1998 Bioinformatics 14: 846–856
Koken MH, Saib A, de The H . 1995 C R Acad. Sci. III 318: 733–739
Kraus WL, Kadonaga JT . 1998 Genes Dev. 12: 331–342
Kundu TK, Palhan VB, Wang Z, An W, Cole PA, Roeder RG . 2000 Mol. Cell 6: 551–561
Kwok RP, Lundblad JR, Chrivia JC, Richards JP, Bachinger HP, Brennan RG, Roberts SG, Green MR, Goodman RH . 1994 Nature 370: 223–226
Lundblad JR, Kwok RP, Laurance ME, Harter ML, Goodman RH . 1995 Nature 374: 85–88
Naar AM, Beaurang PA, Robinson KM, Oliner JD, Avizonis D, Scheek S, Zwicker J, Kadonaga JT, Tjian R . 1998 Genes Dev. 12: 3020–3031
Nakajima T, Uchida C, Anderson SF, Lee CG, Hurwitz J, Parvin JD, Montminy M . 1997 Cell 90: 1107–1112
Neuwald AF, Landsman D . 1997 Trends Biochem. Sci. 22: 154–155
Ogryzko VV, Schiltz RL, Russanova V, Howard BH, Nakatani Y . 1996 Cell 87: 953–959
Ponting CP, Blake DJ, Davies KE, Kendrick-Jones J, Winder SJ . 1996 Trends Biochem. Sci. 21: 11–13
Puri PL, Sartorelli V . 2000 J. Cell Physiol. 185: 155–173
Radhakrishnan I, Perez-Alvarado GC, Parker D, Dyson HJ, Montminy MR, Wright PE . 1999 J. Mol. Biol. 287: 859–865
Rojas JR, Trievel RC, Zhou J, Mo Y, Li X, Berger SL, Allis CD, Marmorstein R . 1999 Nature 401: 93–98
Rost B, Sander C . 1993 J. Mol. Biol. 232: 584–599
Shi Y, Mello C . 1998 Genes Dev. 12: 943–955
Shikama N, Lyon J, La Thangue NB . 1997 Trends Cell Biol. 7: 230–236
Thompson JD, Higgins DG, Gibson TJ . 1994 Nucleic Acids Res. 22: 4673–4680
Utley RT, Ikeda K, Grant PA, Cote J, Steger DJ, Eberharter A, John S, Workman JL . 1998 Nature 394: 498–502
Wilson R, Ainscough R, Anderson K, Baynes C, Berks M, Bonfield J, Burton J, Connell M, Copsey T, Cooper J, Coulson A, Craxton H, Dear S, Du Z, Durbin R, Favello A, Frazer A, Fulton L, Gardner A, Green P, Hawkins T, Hillier L, Jior M, Johnston L, Jones M, Kershaw J, Kirsten J, Laisster N, Latreille P, Lightning J, Lloyd C, Mortimer B, O'Callaghan M, Parsons J, Percy C, Rifken L, Roopra A, Saunders D, Shownkeen R, Simms M, Smaldon N, Smith A, Smith M, Sonnhammer E, Staden R, Sulston J, Thierry-Mieg J, Thomas K, Vaudin M, Vaughan K, Waterston R, Watson A, Weinstock L, Wilkinson-Sproat J, Wohldman P . 1994 Nature 368: 32–38
Yao TP, Oh SP, Fuchs M, Zhou ND, Ch'ng LE, Newsome D, Bronson RT, Li E, Livingston DM, Eckner R . 1998 Cell 93: 361–372
Yuan LW, Gambee JE . 2000 J. Biol. Chem. 275: 40946–40951
Yuan W, Condorelli G, Caruso M, Felsani A, Giordano A . 1996 J. Biol. Chem. 271: 9009–9013
Yuan YP, Eulenstein O, Vingron M, Bork P . 1998 Bioinformatics 14: 285–289
Zhang W, Bieker JJ . 1998 Proc. Natl. Acad. Sci. USA 95: 9855–9860
Acknowledgements
We thank Ms Marie L Basso for her editorial assistance during manuscript preparation. This work was supported by NIH grants to A Giordano.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Yuan, L., Giordano, A. Acetyltransferase machinery conserved in p300/CBP-family proteins. Oncogene 21, 2253–2260 (2002). https://doi.org/10.1038/sj.onc.1205283
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.onc.1205283
Keywords
This article is cited by
-
KRAS mutation in secondary malignant histiocytosis arising from low grade follicular lymphoma
Diagnostic Pathology (2018)
-
CREBBP and p300 lysine acetyl transferases in the DNA damage response
Cellular and Molecular Life Sciences (2018)
-
GPS-PAIL: prediction of lysine acetyltransferase-specific modification sites from protein sequences
Scientific Reports (2016)
-
The structural basis of protein acetylation by the p300/CBP transcriptional coactivator
Nature (2008)
-
Pro-apoptotic activity of oncogenic H-Ras for histone deacetylase inhibitor to induce apoptosis of human cancer HT29 cells
Journal of Cancer Research and Clinical Oncology (2007)