Abstract
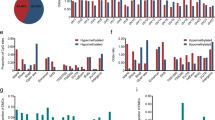
Restriction landmark genomic scanning (RLGS) was utilized to identify novel genomic alterations in hepatocellular carcinoma (HCC). Thirty-one HCC samples were examined by RLGS. Two high intensity spots were common to several RLGS profiles of different HCCs. Nucleotide sequencing and homology search analysis showed that these spots represented repetitive sequences, Human tandem repeat sequence (Genbank, L09552) and centromeric NotI cluster (Genbank, Y10752). These intensified signals were attributable to the occurrence of demethylated areas in the recognition sequence of the NotI site of the corresponding fragments. The intensity of these spots in the RLGS profile reflects their degree of demethylation, which was significantly correlated with postoperative recurrence, even in patients regarded as belonging to the good prognosis group by conventional prognostic factors. Multivariate analysis showed that the intensities of the two spots retained independent prognostic value. This is a new type of predictive factor for HCC based on epigenetic changes in hepatocarcinogenesis, and in the future it is expected to be of great value in making preoperative diagnosis and selecting postoperative therapy.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Aitschul SF, Gish W, Miller W, Myers EW, Lipman DJ . 1990 J. Mol. Biol. 215: 403–410
Arii S, Tanaka J, Yamazoe Y, Minematsu S, Morino T, Fujita K, Maetani S, Tobe T . 1992 Cancer 69: 913–919
Baylin SB, Herman JG, Graff JR, Vertino PM, Issa JP . 1998 Adv. Cancer Res. 72: 141–196
Bender J . 1998 Trends Biochem. Sci. 23: 252–256
Blin N, Stafford DW . 1976 Nucleic Acids Res. 3: 2303–2308
Cravo M, Pinto R, Fidalgo P, Chaves P, Gloria L, Leitao CN, Mira FC . 1996 Gut 39: 434–438
Hatada I, Hayashizaki Y, Hirotsune S, Komatsubara H, Mukai T . 1991 Proc. Natl. Acad. Sci. USA 88: 9523–9527
Himeno Y, Fukuda Y, Hatanaka M, Imura H . 1988 Liver 8: 208–212
Hsu H-C, Wu T-T, Wu M-Z, Sheu J-C, Lee C-S, Chen D-S . 1988 Cancer 61: 2095–2099
Itano O, Ueda M, Kikuchi K, Shimazu M, Kitagawa Y, Aiura K, Kitajima M . 2000 Oncogene 19: 1676–1683
Kim Y-I, Giuliano A, Hatch KD, Schneider A, Nour MA, Dallel GE, Selhub J, Mason JB . 1994 Cancer 74: 893–899
Kubo S, Nishiguchi S, Hirohashi K, Shuto T, Kuroki T, Minamitani S, Ikebe T, Yamamoto T, Wakasa K, Kinoshita H . 1998 Jpn. J. Cancer Res. 89: 419–426
Laird PW, Jaenisch R . 1996 Annu. Rev. Genet. 30: 441–464
Liew CT, Li H-M, Lo K-W, Leow CK, Chan JYH, Hin LY, Lau WY, Lai PBS, Lim BK, Huang J, Leung WT, Wu S, Lee JCK . 1999 Oncogene 18: 789–795
Lise M, Bacchetti S, Da Pian P, Nitti D, Pilati PL, Pigato P . 1998 Cancer 82: 1028–1036
Liver Cancer Study Group of Japan. 1997 Classification of Primary Liver Cancer first English edition Kanehara and Co., Ltd.: Tokyo pp 1–40
Miwa W, Yashima K, Sekine T, Sekiya T . 1995 Electrophoresis 16: 227–232
Nagai H, Kim YS, Yasuda T, Ohmachi Y, Yokouchi H, Monden M, Emi M, Konishi N, Nogami M, Okumura K, Matsubara K . 1999a Gene 237: 15–20
Nagai H, Baba M, Konishi N, Kim YS, Nogami M, Okumura K, Emi M, Matsubara K . 1999b DNA Res. 6: 219–225
Narayan A, Ji W, Zhang XY, Marrogi A, Graff JR, Baylin SB, Ehrlich M . 1998 Int. J. Cancer 776: 833–838
Ohsumi T, Okazaki Y, Hirotsune S, Shibata H, Muramatsu M, Suzuki H, Taga C, Watanabe S, Hayashizaki Y . 1995 Electrophresis 16: 203–209
Qu G, Dubeau L, Narayan A, Yu MC, Ehrlich M . 1999 Mutat. Res. 42312: 91–101
Shen L, Fang J, Qiu D, Zhang T, Yang J, Chen S, Xiao S . 1998 Hepato-Gastroenterology 45: 1753–1759
Shiraishi M, Sekiguchi A, Chuu YH, Sekiya T . 1999 Biol. Chem. 3801: 81–84
Sirabe K, Kanematsu T, Matsumata T, Adachi E, Akazawa K, Sugimachi K . 1991 Hepatology 14: 913–919
Soares J, Pinto AE, Cunha CV, Andre S, Barao I, Sousa JM, Cravo M . 1999 Cancer 85: 112–118
Teubner B, Schulz WA . 1994 Exp. Cell Res. 2102: 192–200
Thoraval D, Asakawa J, Wimmer K, Kuick R, Lamb B, Richardson B, Ambros P, Glover T, Hanash S . 1996 Gene, Chrom. Cancer 17: 234–244
Zhang X-K, Huang D-P, Chiu D-K, Chiu J-F . 1987 Biochem. Biophys. Res. Commun. 142: 932–938
Zrihan-Licht S, Weiss M, Keydar I, Wreschner DH . 1995 Int. J. Cancer 28623: 245–251
Acknowledgements
The authors would like to thank T Muramatsu and Y Inaba for their expert technical assistance and N Sugimoto (Sanwa Kagaku Kenkyusho Co Ltd) for his valuable advice on statistical analysis. This work was supported by the a Grant-in-Aid from the Ministry of Education, Science and Culture and the Ministry of Health and Welfare of Japan and by a National Grant-in-Aid for the Establishment of a High-Tech Research Center in a Private Japanese University.
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Itano, O., Ueda, M., Kikuchi, K. et al. Correlation of postoperative recurrence in hepatocellular carcinoma with demethylation of repetitive sequences. Oncogene 21, 789–797 (2002). https://doi.org/10.1038/sj.onc.1205124
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.onc.1205124
Keywords
This article is cited by
-
Foetal haemoglobin-blood cells (F-cells) as a feature of embryonic tumours (blastomas)
British Journal of Cancer (2007)
-
Epigenetic diagnostics of cancer — the application of DNA methylation markers
Journal of Applied Genetics (2006)
-
Distinction of acute lymphoblastic leukemia from acute myeloid leukemia through microarray-based DNA methylation analysis
Annals of Hematology (2005)