Abstract
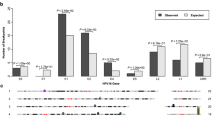
Persistent high risk type human papillomavirus (HR–HPVs) infections induce dysplasia or cancer of the anogenital tract, most notably of the uterine cervix. The viral genome usually persists and replicates as an episomal molecule in early dysplasia, whereas in advanced dysplasia or cervical cancer HPV genomes are frequently integrated into the chromosomal DNA of the host cell. Previous studies suggested that modification of critical cellular sequences by integration of HPV genomes might significantly contribute to the neoplastic transformation of anogenital epithelia (insertional mutagenesis). This prompted us to characterize the integration loci of high risk HPV genomes in a large set of genital lesions. We amplified E6/E7 oncogene transcripts derived from integrated HPV16 and HPV18 genomes and characterized in detail the co-transcribed cellular sequences of 64 primary genital lesions and five cervical cancer cell lines. Database analyses of the cellular parts of these fusion transcripts revealed 51 different integration loci, including 26 transcribed genes (14 known genes, 12 EST sequences with unknown gene function). Seventeen sequences showed similarity to repetitive elements, and 26 sequences did not show any database match other than genomic sequence. Chromosomal integration loci were distributed over almost all human chromosomes. Although we found HPV sequences integrated into cancer related genes and close to fragile sites, no preferential site or integration motif could be identified. These data demonstrate that target directed insertional mutagenesis might occur in few HPV-induced anogenital lesions, however, it is rather the exception than the rule.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ . 1990 J. Mol. Biol. 215: 403–410
Amati B, Frank SR, Donjerkovic D, Taubert S . 2001 Biochem. Biophys. Acta. 1471: M135–M145
Baker CC, Phelps WC, Lindgren V, Braun MJ, Gonda MA, Howley PM . 1987 J. Virol. 61: 962–971
Baron BW, Nucifora G, McCabe N, Espinosa III R, Le Beau MM, McKeithan TW . 1993 Proc. Natl. Acad. Sci. USA 90: 5262–5266
Cannizzaro LA, Durst M, Mendez MJ, Hecht BK, Hecht F . 1988 Cancer Genet. Cytogenet. 33: 93–98
Carmody MW, Jones M, Tarraza H, Vary CP . 1996 Mol. Cell Probes 10: 107–116
Chen M, Tomkins DJ, Auerbach W, McKerlie C, Youssoufian H, Liu L, Gan O, Carreau M, Auerbach A, Groves T, Guidos CJ, Freedman MH, Cross J, Percy DH, Dick JE, Joyner AL, Buchwald M . 1996 Nat. Genet. 12: 448–451
Couturier J, Sastre-Garau X, Schneider-Maunoury S, Labib A, Orth G . 1991 J. Virol. 65: 4534–4538
Cullen AP, Reid R, Campion M, Lorincz AT . 1991 J. Virol. 65: 606–612
Duensing S, Duensing A, Crum CP, Munger K . 2001 Cancer Res. 61: 2356–2360
Duensing S, Lee LY, Duensing A, Basile J, Piboonniyom S, Gonzalez S, Crum CP, Munger K . 2000 Proc. Natl. Acad. Sci. USA 97: 10002–10007
Durst M, Croce CM, Gissmann L, Schwarz E, Huebner K . 1987 Proc. Natl. Acad. Sci. USA 84: 1070–1074
Durst M, Glitz D, Schneider A, zur Hausen H . 1992 Virology 189: 132–140
el Awady MK, Kaplan JB, O'Brien SJ, Burk RD . 1987 Virology 159: 389–398
Frohman MA . 1993 Meth. Enzymol. 218: 340–356
Haskill S, Peace A, Morris J, Sporn SA, Anisowicz A, Lee SW, Smith T, Martin G, Ralph P, Sager R . 1990 Proc. Natl. Acad. Sci. USA 87: 7732–7736
Hibi K, Trink B, Patturajan M, Westra WH, Caballero OL, Hill DE, Ratovitski EA, Jen J, Sidransky D . 2000 Proc. Natl. Acad. Sci. USA 97: 5462–5467
Hopper JL . 2001 Breast Cancer Res. 3: 154–157
Jeon S, Lambert PF . 1995 Proc. Natl. Acad. Sci. USA 92: 1654–1658
Kessis TD, Connolly DC, Hedrick L, Cho KR . 1996 Oncogene 13: 427–431
Klaes R, Woerner SM, Ridder R, Wentzensen N, Duerst M, Schneider A, Lotz B, Melsheimer P, von Knebel DM . 1999 Cancer Res. 59: 6132–6136
Lazo PA . 1999 Br. J. Cancer 80: 2008–2018
Li Y, Kanki H, Hachiya T, Ohyama T, Irie S, Tang G, Mukai J, Sato T . 2000 Int. J. Cancer 87: 473–479
Luft F, Klaes R, Nees M, Durst M, Heilmann V, Melsheimer P, von Knebel Doeberitz M . 2001 Int. J. Cancer 92: 9–17
Macdonald JS . 1999 Semin. Oncol. 26: 556–560
Meyer T, Arndt R, Stockfleth E, Flammann HT, Wolf H, Reischl U . 1995 Biotechniques 19: 632–639
Mincheva A, Gissmann L, zur Hausen H . 1987 Med. Microbiol. Immunol. (Berl.) 176: 245–256
Popescu NC, DiPaolo JA, Amsbaugh SC . 1987 Cytogenet. Cell Genet. 44: 58–62
Remm M, Remm A, Ustav M . 1999 J. Virol. 73: 3062–3070
Reuter S, Bartelmann M, Vogt M, Geisen C, Napierski I, Kahn T, Delius H, Lichter P, Weitz S, Korn B, Schwarz E . 1998 EMBO J. 17: 215–222
Scheffner M, Romanczuk H, Munger K, Huibregtse JM, Mietz JA, Howley PM . 1994 Curr. Top. Microbiol. Immunol. 186: 83–99
Schneider-Gadicke A, Schwarz E . 1986 EMBO J. 5: 2285–2292
Schwarz E, Freese UK, Gissmann L, Mayer W, Roggenbuck B, Stremlau A, zur Hausen H . 1985 Nature 314: 111–114
Shirasawa H, Tomita Y, Fuse A, Yamamoto T, Tanzawa H, Sekiya S, Takamizawa H, Simizu B . 1989 J. Gen. Virol. 70: 1913–1919
Smith CA, Gruss HJ, Davis T, Anderson D, Farrah T, Baker E, Sutherland GR, Brannan CI, Copeland NG, Jenkins NA . 1993 Cell 73: 1349–1360
Smith PP, Friedman CL, Bryant EM, McDougall JK . 1992 Genes Chromosomes Cancer 5: 150–157
Stoler MH, Rhodes CR, Whitbeck A, Wolinsky SM, Chow LT, Broker TR . 1992 Hum. Pathol. 23: 117–128
Sutherland GR, Richards RI . 1999 Am. J. Hum. Genet. 64: 354–359
Thorland EC, Myers SL, Persing DH, Sarkar G, McGovern RM, Gostout BS, Smith DI . 2000 Cancer Res. 60: 5916–5921
Tu Y, Wu C . 1999 Biochem. Biophys. Acta. 1489: 452–456
von Knebel Doeberitz M, Oltersdorf T, Schwarz E, Gissmann L . 1988 Cancer Res. 48: 3780–3786
von Knebel Doeberitz M, Bauknecht T, Bartsch D, zur Hausen H . 1991 Proc. Natl. Acad. Sci. USA 88: 1411–1415
Walboomers JM, Jacobs MV, Manos MM, Bosch FX, Kummer JA, Shah KV, Snijders PJ, Peto J, Meijer CJ, Munoz N . 1999 J. Pathol. 189: 12–19
zur Hausen H . 1999 Semin. Cancer Biol. 9: 405–411
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Wentzensen, N., Ridder, R., Klaes, R. et al. Characterization of viral-cellular fusion transcripts in a large series of HPV16 and 18 positive anogenital lesions. Oncogene 21, 419–426 (2002). https://doi.org/10.1038/sj.onc.1205104
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.onc.1205104
Keywords
This article is cited by
-
Human papillomavirus integration transforms chromatin to drive oncogenesis
Genome Biology (2023)
-
Recurrent integration of human papillomavirus genomes at transcriptional regulatory hubs
npj Genomic Medicine (2021)
-
Whole genome sequencing reveals complexity in both HPV sequences present and HPV integrations in HPV-positive oropharyngeal squamous cell carcinomas
BMC Cancer (2019)
-
Biological relevance of human papillomaviruses in vulvar cancer
Modern Pathology (2017)
-
Functional variants of human papillomavirus type 16 demonstrate host genome integration and transcriptional alterations corresponding to their unique cancer epidemiology
BMC Genomics (2016)