Abstract
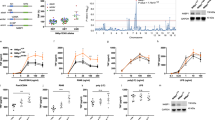
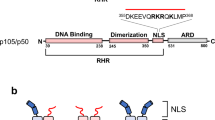
RelB is an unusual member of the Rel/NF-κB family of transcription factors which are involved in oncogenic processes. Due to a relaxed control by the IκBs, the cytosolic NF-κB inhibitors, RelB is constitutively expressed in the nuclei of lymphoid cells. We show here that RelB is inducibly degraded upon activation of T cells in a fashion similar to the IκBs. However, RelB degradation differs from that of IκBs since it is not induced by TNFα but only by T cell receptor or TPA/ionomycin stimulation. Moreover, RelB degradation occurs in three steps: (i) after stimulation RelB is rapidly phosphorylated at amino acids Thr84 and Ser552 followed by (ii) an N-terminal cut and, finally, (iii) the complete degradation in the proteasomes. Since mutation of the two phosphoacceptor sites to non-acceptor sites abolished RelB phosphorylation in vivo and led to the stabilization of the mutated RelBDM, site-specific phosphorylation appears to be a necessary prerequisite for RelB degradation. RelB is a crucial regulator of NF-κB-dependent gene expression. Thus, the signal-induced degradation of RelB should be an important control mechanism of NF-κB activity.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Baldwin Jr AS . 1996 Annu. Rev. Immunol. 14: 649–683
Barton D, HogenEsch H, Weih F . 2000 Eur. J. Immunol. 30: 2323–2332
Brown K, Gerstberger S, Carlson L, Franzoso G, Siebenlist U . 1995 Science 267: 1485–1488
Burkly L, Hession C, Ogata L, Reilly C, Marconi LA, Olson D, Tizard R, Cate R, Lo D . 1995 Nature 373: 531–536
Chen Z, Hagler J, Palombella VJ, Melandri F, Scherer D, Ballard D, Maniatis T . 1995 Genes Dev. 9: 1586–1597
DiDonato J, Mercurio F, Rosette C, Wu-Li J, Suyang H, Ghosh S, Karin M . 1996 Mol. Cell. Biol. 16: 1295–1304
Dobrzanski P, Ryseck RP, Bravo R . 1993 Mol. Cell. Biol. 13: 1572–1582
Dobrzanski P, Ryseck RP, Bravo R . 1994 EMBO J. 13: 4608–4616
Ghosh S, May MJ, Kopp EB . 1998 Annu. Rev. Immunol. 16: 225–260
Gilmore TD, Koedood M, Piffat KA, White DW . 1996 Oncogene 13: 1367–1378
Hibi M, Lin A, Smeal T, Minden A, Karin M . 1993 Genes Dev. 7: 2135–2148
Karin M, Ben-Neriah Y . 2000 Annu. Rev. Immunol. 18: 621–663
Lernbecher T, Muller U, Wirth T . 1993 Nature 365: 767–770
Marienfeld R, Neumann M, Chuvpilo S, Escher C, Kneitz B, Avots A, Schimpl A, Serfling E . 1997 Eur. J. Immunol. 27: 1601–1609
Neumann M, Grieshammer T, Chuvpilo S, Kneitz B, Lohoff M, Schimpl A, Franza Jr BR, Serfling E . 1995 EMBO J. 14: 1991–2004
Neumann M, Marienfeld R, Serfling E . 1997 Int. J. Oncol. 11: 1335–1347
Regnier CH, Song HY, Gao X, Goeddel DV, Cao Z, Rothe M . 1997 Cell 90: 373–383
Ryseck RP, Bull P, Takamiya M, Bours V, Siebenlist U, Dobrzanski P, Bravo R . 1992 Mol. Cell. Biol. 12: 674–684
Suhasini M, Reddy CD, Reddy EP, DiDonato JA, Pilz RB . 1997 Oncogene 15: 1859–1870
Tojima Y, Fujimoto A, Delhase M, Chen Y, Hatakeyama S, Nakayama K, Kaneko Y, Nimura Y, Motoyama N, Ikeda K, Karin M, Nakanishi M . 2000 Nature 404: 778–782
Weih F, Carrasco D, Bravo R . 1994 Oncogene 9: 3289–3297
Weih F, Carrasco D, Durham SK, Barton DS, Rizzo CA, Ryseck RP, Lira SA, Bravo R . 1995 Cell 80: 331–340
Xia Y, Chen S, Wang Y, Mackman N, Ku G, Lo D, Feng L . 1999 Mol. Cell. Biol. 19: 7688–7696
Acknowledgements
We thank I Pietrowski and H Wecklein for excellent technical assistance. For critical reading of the manuscript we are indebted to Drs A Avots, S Klein-Hessling, A Schimpl, A Wilisch-Neumann and T Wirth. This work was supported by the Deutsche Forschungsgemeinschaft (DFG) (NE 608/1-3 and 608/2-2 to M Neumann) and the Wilhelm-Sander Stiftung.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Marienfeld, R., Berberich-Siebelt, F., Berberich, I. et al. Signal-specific and phosphorylation-dependent RelB degradation: a potential mechanism of NF-κB control. Oncogene 20, 8142–8147 (2001). https://doi.org/10.1038/sj.onc.1204884
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.onc.1204884
Keywords
This article is cited by
-
Biological characteristics of transcription factor RelB in different immune cell types: implications for the treatment of multiple sclerosis
Molecular Brain (2019)
-
GSK3β modulates NF-κB activation and RelB degradation through site-specific phosphorylation of BCL10
Scientific Reports (2018)
-
The paracaspase MALT1: biological function and potential for therapeutic inhibition
Cellular and Molecular Life Sciences (2016)
-
Decreased expression of the NF-κB family member RelB in lung fibroblasts from Smokers with and without COPD potentiates cigarette smoke-induced COX-2 expression
Respiratory Research (2015)
-
Transcriptional repression by the HDAC4–RelB–p52 complex regulates multiple myeloma survival and growth
Nature Communications (2015)