Abstract
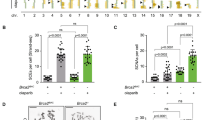
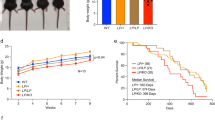
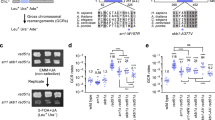
To measure the impact of the RAD51 pathway on Sister-Chromatid Exchanges (SCE), we used hamster cells expressing either the wild-type MmRAD51, which stimulates, or the dominant negative SMRAD51, which inhibits, gene conversion without affecting cell viability of untreated as well as γ-rays irradiated cells. We show that MmRAD51 did not affect the rate of spontaneous SCE while it strongly stimulated spontaneous recombination between tandem repeats. No spontaneous recombinant was detected when expressing SMRAD51 while spontaneous SCE were only slightly diminished. After treatment by an alkylating agent (MNU), MmRAD51 stimulated MNU-induced recombination whereas no recombinant was detected when expressing SMRAD51. MNU induced SCE in all cell lines, even in the SMRAD51 expressing lines, but the induction of SCE was slightly more efficient in lines expressing MmRAD51 and less efficient in lines expressing SMRAD51. Thus, in mammalian cells, the RAD51-dependent gene conversion pathway drastically affects recombination between intrachromosomal tandem repeats, whereas it only partially participates in SCE formation, measured at a chromosomal level. These results show that RAD51-gene conversion can participate in induced SCE but that alternative pathways should exist.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Buerstedde JM, Takeda S . 1991 Cell 67: 179–188
Connell JR, Medcalf AS . 1982 Carcinogenesis 3: 385–390
Dillehay LE, Jacobson-Kram D, Williams JR . 1989 Mutat. Res. 215: 15–23
Dronkert ML, Beverloo HB, Johnson RD, Hoeijmakers JH, Jasin M, Kanaar R . 2000 Mol. Cell. Biol. 20: 3147–3156
Essers J, Hendriks RW, Swagemakers SM, Troelstra C, de Wit J, Bootsma D, Hoeijmakers JH, Kanaar R . 1997 Cell 89: 195–204
Fabre F, Boulet A, Roman H . 1984 Mol. Gen. Genet. 195: 139–143
Fuller LF, Painter RB . 1988 Mut. Res. 193: 109–121
Golub EI, Kovalenko OV, Gupta RC, Ward DC, Radding CM . 1997 Nucleic Acids Res. 25: 4106–4110
Haber JE . 2000 Trends Genet. 16: 259–264
Johnson RD, Jasin M . 2000 EMBO J. 19: 3398–3407
Johnson RD, Liu N, Jasin M . 1999 Nature 401: 397–399
Kadyk LC, Hartwell LH . 1992 Genetics 132: 387–402
Lambert S, Lopez BS . 2000 EMBO J. 19: 3090–3099
Liang F, Han M, Romanienko PJ, Jasin M . 1998 Proc. Natl. Acad. Sci. USA 95: 5172–5177
Pierce AJ, Johnson RD, Thompson LH, Jasin M . 1999 Genes Dev. 13: 2633–2638
Sonoda E, Sasaki MS, Buerstedde JM, Bezzubova O, Shinohara A, Ogawa H, Takata M, Yamaguchi-Iwai Y, Takeda S . 1998 EMBO J. 17: 598–608
Sonoda E, Sasaki MS, Morrison C, Yamaguchi-Iwai Y, Takata M, Takeda S . 1999 Mol. Cell. Biol. 19: 5166–5169
Tsuzuki T, Fujii Y, Sakumi K, Tominaga Y, Nakao K, Sekiguchi M, Matsushiro A, Yoshimura Y, Morita T . 1996 Proc. Natl. Acad. Sci. USA 93: 6236–6240
Zhang H, Tsujimura T, Bhattacharyya NP, Maher VM, McCormick JJ . 1996 Carcinogenesis 17: 2229–2235
Acknowledgements
Thanks are due to Dr M Jasin for providing us CHO-DRA10. We thank Drs B Dutrillaux, L Sabatier and M Ricoul, P Bertrand and Dr D Marsh for helpful discussion and correction of the manuscript, Dr J Brenot for statistical expertise. S Lambert is supported by a fellowship from ‘la Ligue Nationale Française contre le Cancer’. This work was supported by Electricité de France, ANRS and ARC (9822).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Lambert, S., Lopez, B. Role of RAD51 in sister-chromatid exchanges in mammalian cells. Oncogene 20, 6627–6631 (2001). https://doi.org/10.1038/sj.onc.1204813
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.onc.1204813
Keywords
This article is cited by
-
Sister chromatid exchanges induced by perturbed replication can form independently of BRCA1, BRCA2 and RAD51
Nature Communications (2022)
-
Genetic interactions between RAD51 and its paralogues for centrosome fragmentation and ploidy control, independently of the sensitivity to genotoxic stresses
Oncogene (2005)
-
Defective DNA single-strand break repair in spinocerebellar ataxia with axonal neuropathy-1
Nature (2005)
-
Targeting of Rad51-dependent homologous recombination: implications for the radiation sensitivity of human lung cancer cell lines
British Journal of Cancer (2005)
-
The dds20 + Gene Controls a Novel Rad51Sp-Dependent Pathway of Recombinational Repair in Schizosaccharomyces pombe
Russian Journal of Genetics (2005)