Abstract
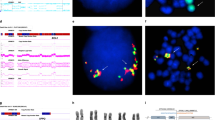
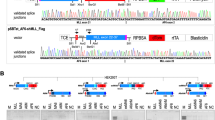
A region on chromosome 14q32.1 is often involved in chromosomal translocations and inversions with one of the T-cell receptor loci in T-cell lymphoproliferative diseases. The breakpoints of the different rearrangements segregate into two clusters; a cluster due to inversion on the centromeric side and a cluster due to simple balanced translocations on the telomeric side. If the target gene activated by these different types of chromosomal rearrangements is the same, the gene must be localized between the two clusters of breakpoints in a region of around 160 kb. Within this breakpoint cluster region, we isolated two genes; namely, TCL1 and TML1/TCL1b genes. In the course of characterizing the TML1 gene, we further identified a third novel gene, which we named TCL6 (T-cell leukemia/lymphoma 6), from a region 7 kb upstream of the TML1 locus. The TCL6 gene expressed at least 11 isoforms through very complex alternative-splicing, including splicing with the TML1 gene. Those isoforms encode at least five open reading frames (ORFs) with no homology to known sequences. The localization of the proteins corresponding to these ORF was determined by fusing green fluorescence protein at the carboxyl terminal of each ORF. ORF141 and ORF72 were observed in the cytoplasmic region, while ORF105, ORF119, and ORF163 were predominantly localized in the nuclear region. Since the TCL6 gene was expressed in T-cell leukemia carrying a t(14;14)(q11;q32.1) chromosome translocation and was not expressed in normal T-cells (just like the TML1 and TCL1 genes), it is also a candidate gene potentially involved in leukemogenesis.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Brito-Babapulle V and Catovsky D . 1991 Cancer Genet Cytogenet 55: 1–9
Croce CM . 1987 Cell 49: 155–156
Davey MP, Bertness V, Nakahara K, Johnson JP, McBride OW, Waldmann TA and Kirsch IR . 1988 Proc Natl Acad Sci USA 85: 9287–9291
Goswami S, Gupta A and Sharma SK . 1998 J Neurochem 71: 1837–1845
Hallas C, Pekarsky Y, Itoyama T, Varnum J, Bichi R, Rothstein JL and Croce CM . 1999 Proc Natl Acad Sci USA 96: 14418–14423
Isobe M, Erikson J, Emanuel BS, Nowell PC and Croce CM . 1985 Science 228: 580–582
Isobe M, Russo G, Haluska FG and Croce CM . 1988 Proc Natl Acad Sci USA 85: 3933–3937
Kanzaki T, Kubonishi I, Eguchi T, Yano S, Sonobe H, Ohyashiki JH, Ohyashiki K, Toyama K, Ohtsuki Y and Miyoshi I . 1992 Cancer 69: 1034–1041
Kubonishi I, Daibata M, Yano S, Isobe M, Kurosawa N, Nagumo H, Ogita Z, Ohyashiki JH, Toyama K and Miyoshi I . 1991 Am J Hematol 37: 179–185
Mengle-Gaw L, Albertson DG, Sherrington PD and Rabbitts TH . 1988 Proc Natl Acad Sci USA 85: 9171–9175
Mengle-Gaw L, Willard HF, Smith CI, Hammarstrom L, Fischer P, Sherrington P, Lucas G, Thompson PW, Baer R and Rabbits TH . 1987 EMBO J 6: 2273–2280
Mount SM . 1982 Nucleic Acids Res 10: 459–472
Murre C, Waldmann RA, Morton CC, Bongiovanni KF, Waldmann TA, Shows TB and Seidman JG . 1985 Nature 316: 549–552
Narducci MG, Virgilio L, Isobe M, Stoppacciaro A, Elli R, Fiorilli M, Carbonari M, Antonelli A, Chessa L, Croce CM and Russo G . 1995 Blood 86: 2358–2364
Russo G, Isobe M, Gatti R, Finan J, Batuman O, Huebner K, Nowell PC and Croce CM . 1989 Proc Natl Acad Sci USA 86: 602–606
Smith SD, McFall P, Morgan R, Link M, Hecht F, Cleary M and Sklar J . 1989 Blood 73: 2182–2187
Stem MH . 1996 Pathol Biol Paris 44: 689–693
Sugimoto J, Hatakeyama T, Narducci MG, Russo G and Isobe M . 1999 Cancer Res 59: 2313–2317
Virgilio L, Isobe M, Narducci MG, Carotenuto P, Camerini B, Kurosawa N, Abbas ar R, Croce CM and Russo G . 1993 Proc Natl Acad Sci USA 90: 9275–9279
Virgilio L, Narducci MG, Isobe M, Billips LG, Cooper MD, Croce CM and Russo G . 1994 Proc Natl Acad Sci USA 91: 12530–12534
Acknowledgements
We thank Dr T Matsuse, Dr R Timms and Dr K Sasaki for critical reading of the manuscripts. This work was supported in part by a Grant-in Aid from the Ministry of Education, Science, Sports and Culture, by a Grant from Joint-Research Project for Regional Intensive from Japan Science and Technology, and by Special Coordination Funds of the Science Technology Agency of the Japanese Government. The sequence accession number of DDBJ corresponding to TCL6 cDNA and genomic sequences are ABO35332, ABO35333, ABO35334, ABO35335, ABO35337, ABO35338, ABO35339, ABO35340, ABO35341, ABO35342, ABO35343, ABO35344, ABO35345 and ABO35346.
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Saitou, M., Sugimoto, J., Hatakeyama, T. et al. Identification of the TCL6 genes within the breakpoint cluster region on chromosome 14q32 in T-cell leukemia. Oncogene 19, 2796–2802 (2000). https://doi.org/10.1038/sj.onc.1203604
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.onc.1203604
Keywords
This article is cited by
-
lncRNA TCL6 correlates with immune cell infiltration and indicates worse survival in breast cancer
Breast Cancer (2020)
-
Long non-coding RNAs defining major subtypes of B cell precursor acute lymphoblastic leukemia
Journal of Hematology & Oncology (2019)
-
Expression of the long non-coding RNA TCL6 is associated with clinical outcome in pediatric B-cell acute lymphoblastic leukemia
Blood Cancer Journal (2019)
-
A Decision Tree Based Classifier to Analyze Human Ovarian Cancer cDNA Microarray Datasets
Journal of Medical Systems (2016)
-
TCL1A gene involvement in T-cell prolymphocytic leukemia in Japanese patients
International Journal of Hematology (2012)