Abstract
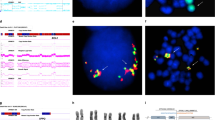
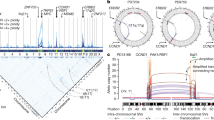
Fusion of TLS/FUS and CHOP gene by reciprocal translocation t(12;16)(q32;q16) is a common genetic event found in myxoid and round-cell liposarcomas. Characterization of this genetic event was performed by three methods, Southern blot, RT – PCR, and genomic long-distance PCR in nine myxoid and three round-cell liposarcomas. All but one tumors showed genetic alternations indicating the fusion of TLS/FUS and CHOP gene. Two novel types of fusion transcripts were found, of which one lacked exon 2 sequence of CHOP gene, and the other lacked 3′ half of exon 5 of TLS gene. The latter case was caused by a cryptic splicing site which was created by the genomic fusion. Detailed analyses genomic fusion points revealed several sequence characteristics surrounding the fusion points. Homology analyses of breakpoint sequences with known sequence motifs possibly involve in the process of translocation uncovered Translin binding sequences at both of TLS/FUS and CHOP breakpoints in two cases. Translocations were always associated with other genetic alterations, such as deletions, duplications, or insertions. Short direct repeats were almost always found at both ends of deleted or duplicated fragments some of which had apparently been created by joining of sequences that flank the rearrangement. Finally, consensus topoisomerase II cleavage sites were found at breakpoints in all cases analysed, suggesting a role of this enzyme in creating staggered ends at the breakpoint. These data suggested that sequence characteristics may play an important role to recruit several factors such as Translin and topoisomerase II in the process of chromosomal translation in liposarcomas.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 50 print issues and online access
$259.00 per year
only $5.18 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
Aoki K, Suzuki K, Sugano T, Tasaka T, Nakahara K, Kuge O, Omori A and Kasai M. . 1994 Oncogene 9: 1109–1115.
Aoki K, Suzuki K, Sugano T, Tasaka T, Nakahara K, Kuge O, Omori A and Kasai M. . 1995 Nat. Genet. 10: 167–173.
Canning S and Dryja TP. . 1989 Proc, Natl. Acad. Sci. USA 86: 5044–5048.
Chalk JG, Barr FG and Mitchell CD. . 1997 Oncogene 15: 1199–1205.
Cheng SJ, Chen Z, D'Auriol L, Le Coniat M, Grausz D and Berger R. . 1989 Oncogene 4: 195–202.
Cheng S, Fockler C, Barnes WM and Higuchi R. . 1994 Proc. Natl. Acad. Sci. USA 91: 5695–5699.
Crowther PJ, Doherty JP, Linsenmeyer ME, Williamson MR and Woodcock DM. . 1991 Nucleic Acids Res. 19: 2395–2401.
Crozat A, Aman P, Mandahl N and Ron D. . 1993 Nature 363: 640–644.
De Klein A, Van Kessek AG . 1982 Nature 300: 765–767.
Efstratiadis A, Posakony JW, Maniatis T, Lawn RM, O'Connell C, Spritz RA, Deriel JK, Forget BG, Weissman SM, Slighton JL, Blechl AE, Smithies O, Baralle FE, Shoulders CC and Proudfoot NJ. . 1980 Cell 21: 653–668.
Heistelkamp N, Stam K, Groffern J, de Klein A and Grosveld G. . 1985 Nature 315: 758–761.
Kariya Y, Kato K, Hayashizaki Y, Himeno S, Tarui S and Matsubara K. . 1987 Gene 53: 1–10.
Kas E and Laemmli UK. . 1992 EMBO J. 11: 705–716.
Kasai M, Aoki K, Matsuo Y Minowada J, Maziarz RT and Strominger JL. . 1994 Int. Immun. 6: 1017–1025.
Knight JC, Renwick PJ, Cin PD, Van den Berghe H and Fletcher CDM. . 1995 Cancer Res. 55: 24–27.
Kovalchuk AL, Miller J and Janz S. . 1997 Oncogene 15: 2369–2377.
Krowczynska AM, Rudders RA and Krontiris TG. . 1990 Nucleic Acids Res. 18: 1121–1127.
Kuroda M, Ishida T, Horiuchi H, Kida N, Uozaki H, Takeuchi H, Tsuji K, Imamura T, Mori S, Machinami R and Watanabe T. . 1995 Am. J. Pathol. 147: 1221–1227.
Kuroda M, Ishida T, Takahashi M, Satoh M, Machinami R and Watanabe T. . 1997 Am. J. Pathol. 151: 735–744.
Nilbert M, Mandahl N, Aman P, Rydholm A and Mitelman F. . 1995 Cancer Genet. Cytogenet. 72: 155–156.
Panagopoulos I, Mandahl N, Ron D, Hoglund M, Nilbert M, Mertens F, Mitelman F and Aman P. . 1994 Cancer Res. 54: 6500–6503.
Panagopoulos I, Mandahl N, Mitelman F and Aman P. . 1995 Oncogene 11: 1133–1137.
Panagopoulos I, Aman P, Mertens F, Mandahl N, Rydholm A, Bauer HFC and Mitelman F. . 1996a Genes Chromosom. Cancer 17: 102–107.
Panagopoulos I, Hoglund M, Merterns F, Mandahl N, Mitelman F and Aman P. . 1996b Oncogene 12: 489–494.
Panagopoulos I, Lassen C. Isaksson M, Mitelman F, Mandahl N and Aman P. . 1997 Oncogene 15: 1357–1362.
Park JS, Luethy JD, Wang MG, Fargnoli J, Fornance Jr AJ, McBride OW and Holbrook MJ. . 1992 Gene 116: 259–267.
Rabbitts TH, Forster A, Larson R and Nathan P. . 1993 Nat. Genet. 4: 175–180.
Rabbitts TH. . 1994 Nature 372: 143–149.
Ron D and Habener JF. . 1992 Genes Develop. 6: 439–453.
Roth DB, Porter TN and Wilson JH. . 1985 Mol. Cell. Biol. 5: 2599–2607.
Sambrook J. Fritish EF and Maniatis T. . 1989 Molecular cloning: a laboratory manual (2nd ed.), Cold Spring Harbor Laboratory, Cold Spring Harbor, NY.
Sanchez-Garcia I and Rabbitts TH. . 1994 Proc. Natl. Acad. Sci. USA 91: 7869–7873.
Sander M and Hsieh T. . 1985 Nucleic Acids Res. 13: 1057–1072.
Sowerby SJ, Kennedy MA, Fitzgerald PH and Morris CM. . 1993 Oncogene 8: 1679–1683.
Sperry AO, Blasquez VC and Garrard WT. . 1989 Proc. Natl, Acad. Sci. USA 86: 5497–5501.
Sptizner JR and Muller MT. . 1988 Nucleic Acids Res. 16: 5533–5556.
Taub R, Kirsch C, Morton G, Lenoir D, Swan S, Tronick S, Aaronson S and Leder P. . 1982 Proc. Natl. Acad. Sci. USA 79: 7837–7841.
Ubeda M, Wang XZ, Zinszner H, Wu I, Habener JF and Ron D. . 1996 Mol. Cell. Biol. 16: 1479–1489.
Zhang JG, Goldman JM and Cross NCP. . 1995 Br. J. Haematol. 90: 138–146.
Zinszner H, Albalat R and Ron D. (1994). . Genes Develop. 8: 2513–2526.
Acknowledgements
We thank Drs K Kusuzaki, Y Ueda, K Hyakuna and H Murakami for kindly providing materials for this study, and Drs Y Kotoura and M Oka for helpful comments and supports for this study. This work was supported in part by Grant-in-Aid for Scientific Research from the Ministry of Education, Science, and Culture (A2-08557085), a Grant from the Princess Takamatsu Cancer Research Found, a Grant from Sagawa Foundation for Promotion of Cancer Research (to JT), and by Grant CA60945 from NIH (to DR).
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Kanoe, H., Nakayama, T., Hosaka, T. et al. Characteristics of genomic breakpoints in TLS-CHOP translocations in liposarcomas suggest the involvement of Translin and topoisomerase II in the process of translocation. Oncogene 18, 721–729 (1999). https://doi.org/10.1038/sj.onc.1202364
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.onc.1202364
Keywords
This article is cited by
-
Phase II study of amrubicin (SM-5887), a synthetic 9-aminoanthracycline, as first line treatment in patients with metastatic or unresectable soft tissue sarcoma: durable response in myxoid liposarcoma with TLS-CHOP translocation
Investigational New Drugs (2016)
-
Involvement of a citrus meiotic recombination TTC-repeat motif in the formation of gross deletions generated by ionizing radiation and MULE activation
BMC Genomics (2015)
-
Synovial sarcoma is a gateway to the role of chromatin remodeling in cancer
Cancer and Metastasis Reviews (2015)
-
Genomic profiling of bone and soft tissue tumors with supernumerary ring chromosomes using tiling resolution bacterial artificial chromosome microarrays
Oncogene (2006)
-
Identification of sequence motifs at the breakpoint junctions in three t(1;9)(p36.3;q34) and delineation of mechanisms involved in generating balanced translocations
Human Genetics (2006)