Abstract
Aim:
The aim of this study was to design and synthesize a series of high activity compounds against aspartyl protease β-secretase (BACE-1) bearing hydroxyethylene (HE) framework.
Methods:
First, we designed the small library based on our previous work and rational analysis. Subsequently, thirteen compounds were selected and synthesized using skilled solid phase synthetic methods to explore the relationship between structure and activity. We then used molecular modeling to explain the possible binding mode.
Results:
Thirteen new compounds (6-18) have been designed, synthesized and bioassayed. Their structures were determined by nuclear magnetic resonance (NMR) spectra, low- and high-resolution mass spectra and optical rotation. Most compounds have shown moderate to excellent activities, and compound 10, which contains fewer amino acids and amide bonds than GRL-7234, was about 5-fold more potent than the control compound 4 discovered by Merck. The molecular modeling results have indicated the possible binding mode and explained the difference between compounds 10 and 16, providing direction for further study.
Conclusion:
This study yielded several high activity compounds bearing fewer amino acids and amide bonds than previous compounds, providing insight into the further development of potent BACE-1 inhibitors for the treatment of Alzheimer's disease.
Similar content being viewed by others
Introduction
Alzheimer's disease (AD), the most common form of dementia in the elderly, is characterized by an impairment of intellectual capacity 1 and brings about a heavy social and financial burden for both society and families. The worldwide AD situation is worsening, with average life expectancy growing 2, 3. To date, there is still no drug that can cure the disease 4, 5. Therefore, the research and development of effective drugs for the prevention and treatment of AD is of significant importance.
Although the actual cause of AD has not been identified with certainty, increasing evidence has linked the disease with β-amyloid peptide (Aβ) 6, 7. Aβ is closely associated with AD and inhibiting Aβ generation may stop or slow AD progression because the polymerization and subsequent aggregation of Aβ cause plaque formation and eventually lead to the loss of neuronal function in AD patients 8, 9. The recently discovered aspartyl protease β-secretase (BACE-1) has received considerable attention because of its key role in the generation of Aβ 10. Moreover, BACE-1 gene knockout mice showed no Aβ production and appeared healthy, which offered further powerful in vivo evidence for the opinion that BACE-1 was an attractive AD target and inhibition of the protease could effectively reduce Aβ formation, thereby halting the progression of AD 11, 12.
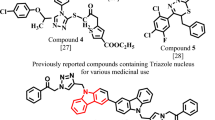
Since first discovered in 1999, numerous BACE-1 inhibitors have been designed, synthesized and evaluated the biological activities, in which some molecules caused an in vivo decrease of amyloids when used in animal models (eg, 1, 2) 13, 14, 15, especially compound GRL-7234 bearing the HE scaffold, which has been shown to induce a reduction of Aβ40 production in transgenic mice after a single intraperitoneal administration (Figure 1) 14. However, the majority of these compounds are peptide-based analogs 16, 17, 18, 19, 20, 21 that, if used as drug agents, will face formidable difficulties due to their vulnerability to degradative enzymes, rapid biliary clearance and poor oral absorption 22. Therefore, the development of therapeutically promising, low molecular weight potent BACE-1 inhibitors is urgently needed.
Merck's group previously found a nonpeptidomimetic BACE-1 inhibitor 3 (Figure 2) through high-throughput screening and lead optimization 23. The group also disclosed the crystal structure of the inhibitor bound to the enzyme, which showed that the molecule occupied the S1-S4 subsites of BACE-1, but lacked direct interaction with the catalytic aspartates of the enzyme; this is presumed to be a main cause of its moderate activity. The authors then synthesized a series of hybrids, such as 4 (Figure 2), by combining 3 and hydoxyethylamine isostere (HEA). These hybrids have been demonstrated to be capable of interacting with catalytic aspartates in previous studies of aspartyl protease inhibitors. Through biological evaluation, the compounds were found to exhibit excellent inhibitory effects and cell-permeability 24. According to these results, many other inhibitors with high potency were also obtained by combining the substituted isopthalamides into the HEA scaffold 25, 26. However, until now, no compound bearing the HEA scaffold has gone to clinical trial.
Hydroxyethylene (HE) is another important isostere of the aspartyl protease inhibitors and the HIV drug, Indina vir, which adopts it as a scaffold, was approved to go on the market several years ago 27. Ghosh and co-workers have proven that compounds that incorporate substituted isopthalamides as P2–P3 ligands in combination with the Leu-Ala HE dipeptide isostere can also form direct hydrogen bonds with the catalytic aspartates and display excellent potencies toward BACE-1; this group discovered the most potent compound, GRL-7234, described above 14. However, this molecule has a large molecular weight and many amide bonds, which may cause problems in further studies. HE isostere has been under study for some time to discover BACE-1 inhibitors in our laboratory 28. Compound 5 was one of the most potent inhibitors found in our previous study. We have also established a general method for the efficacious solid phase synthesis of derivatives based on the HE scaffold through modifying its C-terminus and N-terminus. Based on our previous study, we explored more drug-like inhibitors with fewer amino acids and amide bonds.
Materials and methods
Chemistry
Compound design The structures of the potent HE-based BACE-1 inhibitors, including compound 5 (Figure 3) discovered by our group, share the same HE isostere and similar N-terminal isophthalamide scaffold with GRL-7234 14, 28. These observations promoted us to explore whether potent inhibitors could also be obtained by introducing substituted isopthalamides, discovered by Merck's group and also used by Ghosh and co-workers, into 5. This series may maintain or elevate the inhibitory activities of 5 and possess a lower molecular weight and fewer amide bonds than the compound GRL-7234. Therefore, a small focused library of HE hybrids was designed and constructed. Through biological screening, two highly potent BACE-1 inhibitors, 10 and 11 (IC50=0.010 and 0.031 μmol/L), were identified. In this report, the design, synthesis, and biological evaluation of these inhibitors are described.
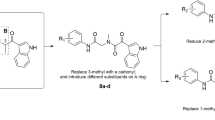
To explore whether substituted isopthalamide fragments fit the S2–S3 pocket of BACE-1 appropriately and helped to improve inhibitory activity when they were introduced into compound 5 and to further investigate the structure-activity relationships (SAR) of these hybrids, we designed and synthesized a focused, small library containing 13 members (Table 1). Isobutyl and cyclopropyl groups were selected as the C-terminal residues of R1 of HE because the isobutyl moiety was found to be a good C-terminus of HE in our previous study 28 and the cyclopropyl group was the C-terminal substituent of the potent HEA-based inhibitors developed by Merck's group 24. N-terminal residues are a series of isophthalamide derivatives with various substituents at the 3- and 5-positions. R3 at the 3-position was investigated using (R)-1-(4-fluorophenyl)ethyl, (S)-1-(4-fluorophenyl)ethyl, (R)-1-phenylethyl, (S)-1-phenylethyl, 4-fluorobenzyl and benzyl groups. An N-methyl(methylsulfonyl)amino, nitro or hydrogen group was incorporated at the 5-position as R2. All designed groups of R3 and R2 were selected according to our previous studies and those by Merck's group 25, 26, 28 and combined several analogues, such as 4-fluorobenzyl and benzyl moieties at the 3- position, for systematically investigating the SAR of these types of inhibitors.
Preparation of the small library by solid-phase synthesis The HE-containing compounds 6–18 (Scheme 1) were synthesized using a solid-phase strategy, which was shown to be effective in the preparation of HE-based compounds in our previous study 28. The route is demonstrated in Scheme 1. First, hydrolysis of the known γ-lactone AA 16, employing lithium hydroxide as base, followed by reaction with allyl bromide, provided allyl ester HE analogue, BB. The esterificaton of BB with TentaGel S COOH resin, using 1-(3-dimethylamino)propyl-3-ethyl-carbodiimide hydrochloride (EDCI) and N,N-dimethylaminopyridine (DMAP), provided the solid supported product, CC. Subsequently, the allyl group of the C-terminus was removed and coupled with the corresponding isobutylamine or cyclopropylamine in the presence of EDCI and 1-hydroxy-benzotriazole (HOBt) to give EE. After the N-Boc group of EE was removed in the presence of 30% CF3COOH in CH2Cl2, the resulting amines were reacted with the corresponding monoallyl isophthalic ester derivatives to yield GG, employing benzotriazole-1-yl-oxy-trispyrrolidino-phosphonium hexafluorophosphate (PyBOP) and HOBt as condensation agents. Finally, removal of the allyl group of GG and reaction with various amines afforded II, which was cleaved with 10% triethylamine in methanol to produce designed compounds 6–18. All compounds were obtained in >60% total yields and in >85% purity before further purification.
Syntheses of compounds 6–18. Reagents and conditions: (a) i) LiOH, H2O, CH3OH, rt; ii) NaHCO3, allyl bromide, DMF, 30 °C; (b) TentaGel S COOH resin, EDCI, DMAP, DMF/CH2Cl2, rt; (c) Pd(PPh3)4, DMBA, CH2Cl2, rt; (d) R1NH2, EDCI, HOBt, DIPEA,DMF, rt; (e) 30% TFA/CH2Cl2, rt; (f) allyl mono-isophthalic ester, PyBOP, HOBt, DIPEA, rt; (g) Pd(PPh3)4, DMBA, CH2Cl2, rt; (h) R3NH2, HBTU, HOBt, DIPEA, DMF, rt; (i) 10% Et3N/CH3OH, 55 °C.
Enzyme-based assay of BACE-1
Recombinant human β-secretase ectodomain (amino acid residues 1–460) was expressed as a secreted protein with a C-terminal His tag in insect cells using baculovirus infection. The BACE activity was determined at room temperature by monitoring the hydrolysis of FRET substrate DABCYL-Ser-Glu-Val-Asn-Leu-Asp-Ala-Glu-Phe-EDANS (SynPep Corp, USA). In a typical 100 μL assay, in a mixture containing 100 mmol/L ammonium acetate, pH 4.0, 20 μmol/L substrate, and 50 nmol/L purified recombinant human BACE-1/Fc, the enzyme activity was continuously monitored with excitation at 355 nm and emission at 460 nm for 20 min and the initial rate of the hydrolysis was determined using the early linear region of the enzymatic reaction kinetic curve.
Molecular docking
AutoDock Tool software was utilized to prepare the proteins and ligands for docking studies. The hydrogens were added to the protein structure and the protein atomic partial charges were assigned with an Amber force field. The partial charges of ligands in the docking study were calculated using the Gasteiger method. The software AutoDock4 29 was adopted to dock ligands 10 and 16 into the binding site of BACE-1. The Lamarckian genetic algorithm (LGA) 30 was applied to deal with the protein-ligand interactions. A Solis and Wets local search was performed for energy minimization on a user-specified proportion of the population. To explore the conformational space of ligands, the overall translation steps were set to 0.2 Å, and the overall rotation and torsion rotation steps were set to 5 degrees in the docking studies. The number of GA generations, energy evaluations, and docking runs was set to 370 000, 8 000 000, and 50, respectively. The 3-dimensional crystal structure of BACE-1 was retrieved from the PDB database (access PDB code 1TQF).
Results
Library design and synthesis
On the basis of the framework of compound 5, compounds 6-18 were designed and synthesized; their chemical structures are shown in Table 1. These compounds were synthesized through the route outlined in Scheme 1. All compounds were obtained in >60% total yields and in >85% purity before further purification. To meet the assay requirements, some key compounds were further purified by preparative thin-layer chromatography (>98%). Details for synthetic procedures and structural characterizations have been described above.
Inhibitory activity toward BACE-1
We also evaluated the BACE-1 inhibitory activities of designed compounds (Table 1). The wavelengths of some compounds were the same as that used to monitor activity in the assay, which led to serious interference on the fluorescence-based bio-assaying; therefore we could not carry out enzyme-level evaluations for those compounds. Encouragingly, most compounds displayed high inhibition against BACE-1, especially compound 10, which was about 5-fold more potent than the control compound 4 discovered by Merck.
Molecular modeling
With the aim of obtaining information about the possible interactions of the new molecules with BACE-1, which, in turn, could be useful in the design of more potent BACE-1 inhibitors, a docking study was performed using the advanced AutoDock program (Figure 4). Compounds 10 and 16 were docked into the protein to compare the relationship between structure and activity. Their interactions have revealed their binding modes and explained the binding differences between the substituted groups at the 5-position of isophthalamide.
The interactions between compounds 10 and 16 with BACE predicted by docking studies were illustrated by Pymol program. (A) It shows the overall conformation of compound 10 situated in the binding site of BACE, the binding site was depicted in surface model; (B) Close-by view of the interaction between compound 10 and BACE. Three important hydrogen bonds between methyl(methylsulfonyl)amino group and residues Asn233, Arg235 and Ser325 of the enzyme were highlighted; (C) Close-by view of the interaction between compound 16 and BACE, which the distances between the nitro group and residues Asn 233 and Arg235 were obviously larger and beyond the normal hydrogen bonding distance (3.5Å).
Discussion
According to the bioassay results shown in Table 1, compounds 10 and 11, bearing a (R)-1-(4-fluorophenyl)ethyl group at the 3-position of the N-terminal isophthalamide and methyl(methylsulfonyl)amino at the 5-position, showed highly potent activities, comparable to that of inhibitor 4 developed by Merck's group (IC50: 0.010 and 0.031 μmol/L vs 0.049 μmol/L). This suggested that like GRL-7234, substituted isopthalamides could also fit the S2–S3 pocket effectively by combination with 5. Interestingly, our compounds may have good properties, with lower molecular weights and fewer amide bonds than GRL-7234. Similar to the observations by Merck's group, we also found that HE-based inhibitors (10 and 11) with (R)-1-(4-fluorophenyl)ethyl groups at the 3-position of isophthalamide were much more potent than those (12 and 13) with the (S)-isomer as a substituent at the site. By comparing 10 and 11 with 14 and 15, it could be observed that the methyl substituent at the benzyl group of R3 is important for potency and its removal would lead to an immense decrease in the activities of the inhibitors. From the results in Table 1, we could also see that isobutyl and cyclopropyl groups displayed no obvious differences at the C-terminus of HE (10-11, 12-13, 16-17). Moreover, according to the results demonstrated by 16-18, compared with N-methyl(methylsulfonyl)amino, nitro and hydrogen groups are clearly not suitable at the 5-position of N-terminal isophthalamide. The results were unexpected because they were in contradiction with our previous study, in which we found the nitro substituent was as good as the methyl(methylsulfonyl)amino moiety at the 5-position. Furthermore, based on previous SAR studies 28, the (4-fluorophenyl)ethyl group does not seem to be an good substituent at the 3-position of N-terminal isophthalamide in HE-based inhibitors because the ligand containing the aromatic group is relatively large. Compounds 10 and 11, which contain the substituent, did show nanomolar activities. The unexpected results pushed us to carefully study the complex crystal structures of 3 and 4 14, 15, 23, 24. Coburn provided a possible explanation for the unexpected phenomena 23 that the α-methyl of (4-fluorophenyl)ethyl of 3 at the 3-position packs firmly against Ile110 of BACE-1, which orients the 4-fluorophenyl ring toward S3 and creates a novel S3 subpocket (S3SP). The subpocket is unique, different from other reported BACE-1 complexed structures, which may make (4-fluorophenyl)ethyl as a suitable substituent as R3, even for HE-based inhibitors 14, 15, although it is relatively large for the S3 pocket. The special orientation of the molecules in the active pockets of BACE-1 is closely associated with the groups of the 5-position of N-terminal isophthalamide, which may result in N-methyl(methylsulfonyl)amino being more preferable than the nitro group at this site when (4-fluorophenyl)ethyl group is selected as R3.
To support this assumption, docking studies have been performed to gain insight into specific ligand-binding site interactions. The modeled structures of 10 and 16 bound to BACE-1 active sites were constructed (PDB entry 1TQF). As shown in Figure 4, compound 10 was accommodated in the active site, which may explain the high potency toward BACE-1. As we expected, at the N-terminal isophthalamide, 10 and 16 demonstrated certain differences. Although the (4-fluorophenyl)ethyl group at 3-position of two compounds fills the S3 subpocket, the compounds show different interactions with BACE-1 at the 5-position. The two oxygens of the methylsulfonyl substituent in 10 form hydrogen bonds with Asn233, Arg235, and Ser325. On the contrary, the oxygens of the nitro group in 16 cannot interact with the ambient residues effectively, which may result in the dramatic loss of inhibitory activity. These observations are consistent with the results of biological evaluations. The analysis from modeling improved the rational design of new BACE-1 inhibitors. Further explorations are actively being pursued and will be reported in due course.
Conclusion
To develop more drug-like BACE-1 potent inhibitors, a series of isophthalamide-HE hybrids was designed and synthesized. During the study, it was found that both isobutyl and cyclopropyl amines were the optimal C-caps of the HE scaffold, and isopthalamide fragments used in the HEA scaffold by Merck' group introduced at the N-terminus as P2–P3 ligands helped to improve potency. Notably, compounds 10 and 11, which have lower molecular weights and fewer amide bonds than GRL-7234, exhibited excellent enzyme-inhibiting potency. With the aid of molecular modeling, valuable information regarding enzyme-inhibitor interactions was obtained, which may help in the rational design of new potent inhibitors. Further studies are in progress and will be reported in due course.
Appendix
The reagents (chemicals) were purchased from Lancaster (Morecambe, England), Acros (Geel, Belgium) and Shanghai Chemical Reagent Company (Shanghai, China) and were used without further purification. Analytical thin-layer chromatography was performed on HSGF 254 (150–200 μm thickness; Yantai Huiyou Company, Yantai, Shandong, China). The 1H NMR (300 MHz or 400 MHz) spectra were recorded on a Varian Mercury-300 or 400 High Performance Digital FT-NMR with TMS as internal standard, and the 13C NMR (100 MHz) spectra were determined using a Varian Mercury-400 High Performance Digital FT-NMR. Chemical shifts were reported in parts per million (ppm, d) downfield from tetramethylsilane. Proton coupling patterns were described as singlet (s), doublet (d), triplet (t), quartet (q), multiplet (m), and broad (br). LC-MS was carried out on Thermo Finnigan LCQDECAXP and HRMS was performed with Finnigan MAT 95, EI: 70 eV, R: 10000, or Micromass Q-TOF Ultima. Purity was determined by Gilson high-performance liquid chromatography (HPLC) (306 pump, uv/vis-156 Detector, 215 liquid handle). The optical rotation value was determined by a Perkin Elmer-341 (589 nm). TentaGel S COOH resin was purchased from Acros.
General procedures for the preparations of 6–18
BB (64 mg, 0.188 mmol), EDCI (36 mg, 0.188 mmol), and DMAP (9 mg, 0.075 mmol) were added to 0.15 g TentaGel S COOH resin (loading: ∼0.25 mmol/g) in 3 mL of a mixture of CH2Cl2 and DMF (4:1) and the mixture was reacted overnight. The mixture was filtered and about 46 mg BB was recovered from the filtrate by chromatography. The resin was washed three times each with DMF, 2-propanol, and CH2Cl2. CC was mixed with Pd(PPh3)4 (9 mg, 0.0075 mmol) in 0.25 mol/L DMBA in 3 mL of CH2Cl2 under an Ar atmosphere for six hours and washed three times each with 20% acetic acid in DMF, DMF, 2-isopropanol, and CH2Cl2. Then the resulting resin was treated with a mixture of isobutylamine (0.038 mL, 0.375 mmol) or cyclopropylamine (0.375 mmol), EDCI (108 mg, 0.563 mmol), HOBt (76 mg, 0.563 mmol), and DIPEA (0.066 mL, 0.375 mmol) in 3 mL anhydrous DMF for 1 day. The resin, EE, was washed three times each with DMF, 2-propanol, and CH2Cl2 and treated with 3 mL of 30% trifluoroacetate in CH2Cl2 for 1 h. The resin was then quickly washed three times each with CH2Cl2, 10% triethylamine in CH2Cl2, CH2Cl2, and DMF. The resulting resin, FF, was reacted overnight with a 25 mL mixture of 0.15 mol/L of the corresponding 3-(allyloxycarbonyl)-benzoic acid derivative, 0.15 mol/L PyBOP, 0.15 mol/L HOBt and 0.45 mol/L DIPEA in 3 mL DMF. The resulting GG were washed three times each with DMF, 2-propanol, and CH2Cl2. Subsequently, GG was treated with Pd(PPh3)4 in 0.25 mol/L DMBA under an Ar atmosphere for 6 h and washed three times each with 20% acetic acid in DMF, DMF, 2-propanol, and CH2Cl2. Then, HH was treated with a mixture of the corresponding amine (0.094 mmol), HBTU (0.036 g, 0.094 mmol), HOBt (0.019 g, 0.14 mmol), and DIPEA (0.033 mL, 0.188 mmol) in 3 mL anhydrous DMF. Then, the product was washed three times each with DMF, 2-propanol, and CH2Cl2. II was reacted with 5 mL 10% triethylamine in methanol at 55 °C for 18 h, and the resin was filtered and washed three times with a mixture of methanol and dichloromethane (1:1). The combined filtrate was concentrated to yield the product. Compounds 6-18 were obtained in 61.6%–74.3% yield (based on theoretical loading value of resin), showed 86.12%–94.56% purity, and were further purified by preparative TLC and showed >98% purity, as determined by HPLC, before biological evaluation.
N1-((4S,5S,7R)-7-(Isobutylcarbamoyl)-5-hydroxy-2-methyloctan-4-yl)-N3-((R)-1-phenylethyl)-5-(methyl(methylsulfonyl)amino)benzene-1,3-diamide (6)
1H NMR (300 MHz, CD3OD): δ 8.25 (t, J=1.4 Hz, 1H), 8.02 (m, 2H), 7.40 (m, 2H), 7.33 (m, 2H), 7.25 (m, 1H), 5.25 (q, J=7.2 Hz, 1H), 4.18 (m, 1H), 3.62 (m, 1H), 3.52 (dt, J=10.4, 3.2 Hz, 1H), 3.35 (s, 3H), 2.95 (s, 3H), 2.92 (d, J=7.2 Hz, 1H), 2.88 (d, J=6.6 Hz, 1H), 2.63 (m, 1H), 1.92 (m, 1H), 1.62 (m, 1H), 1.58 (d, J=7.0 Hz, 3H), 1.40 (m, 2H), 1.30 (m, 2H), 1.20 (d, J=7.2 Hz, 3H), 0.94 (d, J=6.4 Hz, 6H), 0.83 (d, J=6.8 Hz, 3H), 0.81 (d, J=6.4 Hz, 3H); LC-MS: m/z 617.3 [M+H]+; HRMS: calcd for C32H48N4O6SNa [M+Na]+ 639.3192, found 639.3181.
N1-((4S,5S,7R)-7-(Cyclopropylcarbamoyl)-5-hydroxy-2-methyloctan-4-yl)-N3-((R)-1-phenylethyl)-5-(methyl(methylsulfonyl)amino)benzene-1,3-diamide (7)
1H NMR (300 MHz, CD3OD): δ 8.20 (t, J=1.8 Hz, 1H), 7.96 (m, 2H), 7.35 (m, 2H), 7.25 (m, 2H), 7.18 (m, 1H), 5.20 (q, J=7.0 Hz, 1H), 4.20 (dt, J=10.6, 3.1 Hz, 1H), 3.56 (m, 1H), 3.48 (dt, J=10.6, 3.0 Hz, 1H), 3.30 (s, 3H), 2.90 (s, 3H), 2.50 (m, 2H), 1.80 (m, 1H), 1.60 (m, 1H), 1.54 (d, J=6.8 Hz, 3H), 1.34 (m, 1H), 1.24 (m, 2H), 1.06 (d, J=6.9 Hz, 3H), 0.90 (d, J=6.1 Hz, 6H), 0.60 (m, 2H), 0.39 (m, 2H); LC-MS: m/z 601.2 [M+H]+; HRMS: calcd for C31H44N4O6SNa [M+Na]+ 623.2879, found 623.2910.
N1-((4S,5S,7R)-7-(Isobutylcarbamoyl)-5-hydroxy-2-methyloctan-4-yl)-N3-((S)-1-phenylethyl)-5-(methyl(methylsulfonyl)amino)benzene-1,3-diamide (8)
1H NMR (300 MHz, CD3OD): δ 8.25 (m, 1H), 8.06 (m, 2H), 7.43 (m, 2H), 7.33 (m, 2H), 7.25 (m, 1H), 5.25 (q, J=7.0 Hz, 1H), 4.20 (dt, J=6.8, 3.5 Hz, 1H), 3.55 (dt, J=6.8, 4.0 Hz, 1H), 3.38 (s, 3H), 2.95 (s, 3H), 2.88 (m, 2H), 2.65 (m, 1H), 1.90 (m, 1H), 1.60 (m, 1H), 1.58 (d, J=7.0 Hz, 3H), 1.40 (m, 2H), 1.30 (m, 2H), 1.13 (d, J=6.9 Hz, 3H), 0.94 (d, J=6.4 Hz, 6H), 0.85 (d, J=6.7 Hz, 3H), 0.83 (d, J=6.6 Hz, 3H); LC-MS: m/z 617.3 [M+H]+; HRMS: calcd for C32H48N4O6SNa [M+Na]+639.3192, found 639.3179.
N1-((4S,5S,7R)-7-(Cyclopropylcarbamoyl)-5-hydroxy-2-methyloctan-4-yl)-N3-((S)-1-phenylethyl)-5-(methyl(methylsulfonyl)amino)benzene-1,3-diamide (9)
1H NMR (300 MHz, CD3OD): δ 8.20 (t, J=1.4 Hz, 1H), 7.96 (m, 2H), 7.38 (m, 2H), 7.28 (m, 2H), 7.20 (m, 1H), 5.20 (q, J=7.1 Hz, 1H), 4.10 (dt, J=10.4, 3.8 Hz, 1H), 3.46 (dt, J=10.4, 2.9 Hz, 1H), 3.30 (s, 3H), 2.90 (s, 3H), 2.55 (m, 2H), 1.80 (m, 1H), 1.60 (m, 1H), 1.53 (d, J=7.1 Hz, 3H), 1.40 (m, 1H), 1.20 (m, 2H), 1.06 (d, J=6.9 Hz, 3H), 0.90 (d, J=6.6 Hz, 6H), 0.60 (m, 2H), 0.40 (m, 2H); LC-MS: m/z 601.1 [M+H]+.
N1-((4S,5S,7R)-7-(Isobutylcarbamoyl)-5-hydroxy-2-methyloctan-4-yl)-N3-((R)-1-(4-fluorophenyl)ethyl)-5-(methyl(methylsulfonyl)amino)benzene-1,3-diamide (10)
1H NMR (300 MHz, CD3OD): δ 8.20 (t, J=1.8 Hz, 1H), 7.98 (dd, J=1.9, 1.8 Hz, 1H), 7.90 (dd, J=2.0, 1.8 Hz, 1H), 7.38 (m, 2H), 7.00 (m, 2H), 5.20 (q, J=6.7 Hz, 1H), 4.15 (dt, J=10.4, 3.6 Hz, 1H), 3.50 (dt, J=10.0, 3.0 Hz, 1H), 3.30 (s, 3H), 2.90 (s, 3H), 2.95 (d, J=6.8 Hz, 1H), 2.93 (d, J=6.8 Hz, 1H), 2.60 (m, 1H), 1.80 (m, 1H), 1.60 (m, 2H), 1.52 (d, J=7.2 Hz, 3H), 1.35 (m, 2H), 1.24 (m, 1H), 1.09 (d, J=7.0 Hz, 3H), 0.90 (d, J=6.5 Hz, 6H), 0.78 (d, J=6.5 Hz, 3H), 0.76 (d, J=7.0 Hz, 3H); 13C NMR (100MHz, CDCl3): 177.155, 165.952, 164.745, 163.260, 160.815, 142.325, 138.827, 135.985, 135.803, 128.016, 127.938, 127.802, 123.944, 115.583, 115.373, 70.602, 52.603, 49.170, 46.897, 41.045, 38.240, 38.135, 38.003, 35.835, 29.673, 28.403, 24.973, 28.403, 24.973, 23.238, 22.136, 21.776, 20.023, 17.714; LC-MS: m/z 635.3 [M+H]+; HRMS: calcd for C32H47N4O6FS, 634.3200, found 634.3193; [α]D20=-37° (c 0.1050, Acetone).
N1-((4S,5S,7R)-7-(Cyclopropylcarbamoyl)-5-hydroxy-2-methyloctan-4-yl)-N3-((R)-1-(4-fluorophenyl)ethyl)-5-(methyl(methylsulfonyl)amino)benzene-1,3-diamide (11)
1H NMR (300 MHz, CD3OD): δ 8.20 (t, J=1.4Hz, 1H), 8.00 (d, J=1.5 Hz, 2H), 7.40 (m, 2H), 7.00 (m, 2H), 5.20 (q, J=7.0 Hz, 1H), 4.10 (dt, J=10.5, 3.5 Hz, 1H), 3.48 (dt J=10.8, 2.8 Hz, 1H), 3.30 (s, 3H), 2.90 (s, 3H), 2.50 (m, 2H), 1.80 (m, 1H), 1.60 (m, 1H), 1.58 (d, J=7.2 Hz, 3H), 1.35 (m, 1H), 1.24 (m, 1H), 1.06 (d, J=7.0 Hz, 3H), 0.90 (d, J=6.7 Hz, 6H); 13C NMR (100 MHz, CDCl3): 178.703, 166.129, 164.822, 163.160, 160.719, 142.203, 138.879, 135.768, 128.204, 127.995, 127.913, 123.983, 115.494, 115.285, 70.807, 52.391, 49.153, 40.619, 38.210, 37.968, 37.504, 35.655, 29.667, 24.931, 23.264, 22.581, 22.016, 21.729, 17.499, 6.410; LC-MS: m/z 619.1 [M+H]+; HRMS: calcd for C31H43N4O6FS, 618.2887, found 618.2902; [α]D20=-34° (c 0.1250, Acetone).
N1-((4S,5S,7R)-7-(Isobutylcarbamoyl)-5-hydroxy-2-methyloctan-4-yl)-N3-((S)-1-(4-fluorophenyl)ethyl)-5-(methyl(methylsulfonyl)amino)benzene-1,3-diamide (12)
1H NMR (300 MHz, CD3OD): δ 8.20 (t, J=1.5 Hz, 1H), 7.98 (dd, J=2.0, 1.6 Hz, 1H), 7.96 (dd, J=1.9, 1.6 Hz, 1H), 7.40 (m, 2H), 7.00 (m, 2H), 5.20 (q, J=7.1 Hz, 1H), 4.15 (dt, J=10.0, 3.0 Hz, 1H), 3.50 (dt, J=10.0, 3.0 Hz, 1H), 3.30 (s, 3H), 2.93 (s, 3H), 2.90 (m, 2H), 2.60 (m, 1H), 1.80 (m, 1H), 1.45 (m, 2H), 1.52 (d, J=6.9 Hz, 3H), 1.35 (m, 1H), 1.24 (m, 2H), 1.09 (d, J=7.5 Hz, 3H), 0.90 (d, J=6.4 Hz, 6H), 0.80 (d, J=6.3 Hz, 3H), 0.78 (d, J=7.2 Hz, 3H); 13C NMR (100 MHz, CDCl3): 177.178, 165.956, 164.663, 163.256, 160.819, 142.320, 138.827, 135.981, 135.780, 128.052, 127.975, 127.788, 123.917, 115.592, 115.378, 70.601, 52.603, 49.138, 46.920, 41.072, 38.130, 37.998, 35.821, 29.669, 28.435, 24.960, 23.238, 22.660, 22.113, 21.808, 20.041, 17.669; LC-MS: m/z 635.3 [M+H]+; HRMS: calcd for C32H47N4O6FS, 634.3200, found 634.3210; [α]D20=-2° (c 0.3250, Acetone).
N1-((4S,5S,7R)-7-(Cyclopropylcarbamoyl)-5-hydroxy-2-methyloctan-4-yl)-N3-((S)-1-(4-fluorophenyl)ethyl)-5-(methyl(methylsulfonyl)amino)benzene-1,3-diamide (13)
1H NMR (300 MHz, CD3OD): δ 8.20 (t, J=1.5 Hz, 1H), 7.95 (dd, J=1.8, 1.5 Hz, 1H), 7.90 (dd, J=1.8, 1.5 Hz, 1H), 7.40 (m, 2H), 7.00 (m, 2H), 5.20 (q, J=7.3 Hz, 1H), 4.10 (dt, J=10.6, 3.5 Hz, 1H), 3.48 (dt, J=10.0, 2.7 Hz, 1H), 3.40 (s, 3H), 2.90 (s, 3H), 2.50 (m, 2H), 1.80 (m, 1H), 1.60 (m, 1H), 1.50 (d, J=7.0 Hz, 3H), 1.40 (m, 2H), 1.20 (m, 1H), 1.05 (d, J=6.9 Hz, 3H), 0.90 (d, J=6.5 Hz, 6H), 0.60 (m, 2H), 0.38 (m, 2H). 13C NMR (100 MHz, CDCl3): 178.694, 166.129, 164.713, 163.256, 160.815, 142.311, 138.918, 135.912, 135.826, 128.057, 127.975, 127.829, 123.899, 115.565, 115.351, 70.784, 52.499, 49.179, 40.776, 38.203, 37.975, 37.607, 35.753, 29.673, 24.978, 23.256, 22.687, 22.054, 21.795, 17.427, 6.479; LC-MS: m/z 619.2 [M+H]+; HRMS: calcd for C31H43N4O6FS 618.2888, found 618.2898; [α]D20=-1° (c 0.3000, Acetone).
N1-((4S,5S,7R)-7-(Isobutylcarbamoyl)-5-hydroxy-2-methyloctan-4-yl)-N3-(phenylmethyl)-5-(methyl(methylsulfonyl)amino)benzene-1,3-diamide (14)
1H NMR (300 MHz, CD3OD): δ 8.20 (dd, J=1.6, 1.5 Hz, 1H), 8.02 (dd, J=2.2, 1.5 Hz, 1H), 7.96 (dd, J=2.2, 1.6 Hz, 1H), 7.33–7.20 (m, 5H), 4.55 (s, 2H), 4.15 (dt, J=10.3, 3.6 Hz, 1H), 3.50 (dt, J=10.0, 3.0 Hz, 1H), 3.35 (s, 3H), 2.92 (s, 3H), 2.90 (d, J=6.6 Hz, 2H), 2.60 (m, 1H), 1.80 (m, 1H), 1.68–1.58 (m, 2H), 1.40 (m, 2H), 1.20 (m, 1H), 1.05 (d, J=7.0 Hz, 3H), 0.89 (d, J=6.5 Hz, 6H), 0.80 (d, J=6.8 Hz, 3H), 0.78 (d, J=6.5 Hz, 3H); LC-MS: m/z 603.2 [M+H]+.
N1-((4S,5S,7R)-7-(Isobutylcarbamoyl)-5-hydroxy-2-methyloctan-4-yl)-N3-((4-fluorophenyl)methyl)-5-(methyl(methylsulfonyl)amino)benzene-1,3-diamide (15)
1H NMR (300 MHz, CD3OD): δ 8.24 (s, 1H), 8.03 (s, 1H), 8.01 (s, 1H), 7.90 (t, J=5.9 Hz, 1H), 7.40 (m, 2H), 7.05 (m, 2H), 4.55 (s, 2H), 4.20 (m, 1H), 3.55 (m, 1H), 3.35 (s, 3H), 2.95 (s, 3H), 2.90 (m, 2H), 2.62 (m, 1H), 1.80 (m, 1H), 1.60 (m, 3H), 1.40 (m, 2H), 1.12 (d, J=7.2 Hz, 3H), 0.94 (d, J=6.5 Hz, 6H), 0.85 (d, J=6.4 Hz, 3H), 0.83 (d, J=6.8 Hz, 3H); LC-MS: m/z 621.4 [M+H]+; HRMS: calcd for C31H45N4O6FSNa [M+Na]+ 643.2942, found 643.2883; [α]D20=-20.7° (c 1.0300; Acetone).
N1-((4S,5S,7R)-7-(Isobutylcarbamoyl)-5-hydroxy-2-methyloctan-4-yl)-N3-((R)-1-(4-fluorophenyl)ethyl)-5-nitrobenzene-1,3-diamide (16)
1H NMR (400 MHz, CD3OD): δ 8.82 (s, 2H), 8.69 (s, 1H), 7.43 (m, 2H), 7.02 (m, 2H), 5.28 (q, J=7.0 Hz, 1H), 4.20 (m, 1H), 3.55 (m, 1H), 2.90 (m, 2H), 2.63 (m, 1H), 1.90 (m, 1H), 1.68 (m, 2H), 1.60 (d, J=6.8 Hz, 3H), 1.40 (m, 2H), 1.28 (m, 1H), 1.13 (d, J=7.0 Hz, 3H), 0.95 (d, J=6.0 Hz, 6H), 0.84 (d, J=7.0 Hz, 3H), 0.82 (d, J=7.0 Hz, 3H); LC-MS: m/z 573.6 [M+H]+; HRMS: calcd for C30H41N4O6F 572.3010, found 572.2990; [α]D 20 = -51° (c 0.3650, Acetone).
N1-((4S,5S,7R)-7-(Iyclopropylcarbamoyl)-5-hydroxy-2-methyloctan-4-yl)-N3-((R)-1-(4-fluorophenyl) ethyl)-5-nitrobenzene-1,3-diamide (17)
1H NMR (400 MHz, CD3OD): δ 8.83 (s, 1H), 8.81 (s, 1H), 8.71 (s, 1H), 7.42 (m, 2H), 7.05 (m, 2H), 5.26 (q, J=7.0 Hz, 1H), 4.18 (m, 1H), 3.52 (m, 1H), 2.55 (m, 2H), 1.90 (m, 1H), 1.68 (m, 1H), 1.58 (d, J=6.8 Hz, 3H), 1.40 (m, 2H), 1.22 (m, 2H), 1.12 (d, J=7.4 Hz, 3H), 0.96 (d, J=6.6 Hz, 3H), 0.94 (d, J=6.6 Hz, 3H), 0.62 (m, 2H), 0.40 (m, 2H); LC-MS: m/z 557.2 [M+H]+; HRMS: calcd for C29H37N4O6FNa [M+Na]+ 579.2595, found 579.2628; [α]D20=-39° (c 0.4100, Acetone).
N1-((4S,5S,7R)-7-(Isobutylcarbamoyl)-5-hydroxy-2-methyloctan-4-yl)-N3-((R)-1-(4-fluorophenyl)ethyl)-1,3-diamide (18)
1H NMR (400 MHz, CD3OD): δ 8.29 (s, 1H), 7.97 (m, 2H), 7.56 (m, 1H), 7.42 (m, 2H), 7.05 (m, 2H), 5.24 (q, J=7.5 Hz, 1H), 4.18 (m, 1H), 3.55 (m, 1H), 2.92 (d, J=3.8 Hz, 1H), 2.89 (d, J=3.9 Hz, 1H), 2.63 (m, 1H), 1.88 (m, 1H), 1.68 (m, 3H), 1.57 (d, J=6.9 Hz, 3H), 1.40 (m, 2H), 1.13 (d, J=6.9 Hz, 3H), 0.94 (d, J=6.4 Hz, 6H), 0.83 (d, J=7.1 Hz, 3H), 0.81 (d, J=7.2 Hz, 3H); LC-MS: m/z 528.5 [M+H]+; HRMS: calcd for C30H42N3O4F 527.3160, found 527.3163; [α]D20=-56° (c 0.2050, Acetone).
Author contribution
Jing-kang SHEN and Jia LI designed research; Yi-ping ZHU, Kun XIAO, Hai-ping YU, Lan-ping MA, Bing XIONG, Hai-yan ZHANG, and Jing-ya LI performed research; Xin WANG contributed new analytical tools and reagents; Yi-ping ZHU, Kun XIAO, and Lan-ping MA analyzed data; Yi-ping ZHU wrote the paper.
Accession codes
References
McGeer PL, McGeer EG . Alzheimer's disease: arthritis of the brain?. Drugs News Prospect 1995; 8: 80–3.
Hendriksen JV, Nottet HS, Smits HA . Secretases as targets for drug design in Alzheimer's disease. Eur J Clin Invest 2002; 32: 60–8.
Citron M . Beta-secretase inhibition for the treatment of Alzheimer's disease-promise and challenge. Trends Pharmacol Sci 2004; 25: 92–7.
Arendt T . Alzheimer's disease as a disorder of mechanisms underlying structural brain self-organization. Neuroscience 2001; 102: 723–65.
Enz A, Amstutz R, Boddeke H, Gmelin G, Malanowski J . Brain selective inhibition of acetylcholinesterase: a novel approach to therapy for Alzheimer's disease. Prog Brain Res 1993; 98: 431–8.
Monczor M . Diagnosis and treatment of Alzheimer's disease. Curr Med Chem-Central Nervous System Agents 2005; 5: 5–13.
Selkoe DJ . Translating cell biology into therapeutic advances in Alzheimer's disease. Nature 1999; 399 (6738 Suppl): A23–31.
Citron M . β-Secretase inhibition for the treatment of Alzheimer's diseasepromise and challenge. Trends Pharmacol Sci 2004; 25: 92–7.
Hardy JA, Higgins GA . Alzheimer's disease: the amyloid cascade hypothesis. Science 1992; 25: 184–5.
Ghosh AK, Hong L, Tang J . β-Secretase as a therapeutic target for inhibitor drugs. Curr Med Chem 2002; 9: 1135–44.
Luo Y, Bolon B, Kahn S, Bennett BD, Babu-Khan S, Denis P, et al. Mice deficient in BACE1, the Alzheimer′s beta-secretase, have normal phenotype and abolished beta-amyloid generation. Nat Neurosci 2001; 4: 231–2.
Cai H, Wang Y, McCarthy D, Wen H, Borchelt DR, Price DL, et al. BACE1 is the major beta secretase for generation of Abeta peptides by neurons. Nat Neurosci 2001; 4: 233–4.
Durham TB, Shepherd TA . Progress toward the discovery and development of efficacious BACE inhibitors. Curr Opin Drug Dis 2006; 9: 776–91.
Ghosh AK, Kumaragurubaran N, Hong L, Kulkarni SS, Xu X, Chang W, et al. Design, synthesis, and X-ray structure of potent memapsin 2 (β-secretase) inhibitors with isophthalamide derivatives as the P2-P3-ligands. J Med Chem 2007; 50: 2399–407.
Ghosh AK, Kumaragurubaran N, Hong L, Kulkarni S, Xu X, Miller HB, et al. Potent memapsin 2 (β-secretase) inhibitors: design, synthesis, protein-ligand X-ray structure, and in vivo evaluation. Bioorg Med Chem Lett 2008; 8: 1031–6.
Ghosh AK, Bilcer G, Harwood C, Kawahama R, Shin D, Hussain KA, et al. Structure-based design: potent inhibitors of human brain memapsin 2 (β-secretase). J Med Chem 2001; 44: 2865–8.
Shuto D, Kasai S, Kimura T, Liu P, Hidaka K, Hamada T, et al. KMI-008, a novel β-secretase inhibitor containing a hydroxymethylcarbonyl isostere as a transition-state mimic: design and synthesis of substrate-based octapeptides. Bioorg Med Chem Lett 2003; 13: 4273–6.
Kimura T, Shuto D, Kasai S, Liu P, Hidaka K, Hamada T, et al. KMI-358 and KMI-370, highly potent and small-sized BACE1 inhibitors containing phenylnorstatine. Bioorg Med Chem Lett 2004; 14: 1527–31.
Hanessian S, Yun H, Hou Y, Yang G, Bayrakdarian M, Therrien E, et al. Structure-based design, synthesis, and memapsin 2 (bace) inhibitory activity of carbocyclic and heterocyclic peptidomimetics. J Med Chem 2005; 48: 5175–90.
Hanessian S, Yang G, Rondeau JM, Neumann U, Betschart C, Tintelnot-Blomley M . Structure-based design and synthesis of macroheterocyclic peptidomimetic inhibitors of the aspartic protease β-site amyloid precursor protein cleaving enzyme (BACE). J Med Chem 2006; 49: 4544–67.
Freskos JN, Fobian YM, Benson TE, Bienkowski MJ, Brown DL, Emmons TL, et al. Design of potent inhibitors of human β-secretase. Bioorg Med Chem Lett 2007; 17: 73–7.
Plattner JJ, Norbeck DW . In: Drug discovery technologies. Clark CR, Moos WH, Eds. Chichester: Ellis Horwood Limited; 1990. p92–126.
Coburn CA, Stachel SJ, Li YM . Rush DM, Steele TG, Chen-Dodson E, et al. Identification of a small molecule nonpeptide active site β-secretase inhibitor that displays a nontraditional binding mode for aspartyl proteases. J Med Chem 2004; 47: 6117–9.
Stachel SJ, Coburn CA, Steele TG, Jones KG, Loutzenhiser EF, Gregro AR, et al. Structure-based design of potent and selective cell-permeable inhibitors of human β-secretase (bace-1). P J Med Chem 2004; 47: 6447–50.
Iserloh U, Wu Y, Cumming JN, Pan J, Wang LY, Stamford AW, et al. Potent pyrrolidine- and piperidine-based BACE-1 inhibitors. Bioorg Med Chem Lett 2008; 18: 414–7.
Maillard MC, Hom RK, Benson TE, Moon JB, Mamo S, Bienkowski M, et al. Design, synthesis, and crystal structure of hydroxyethyl secondary amine-based peptidomimetic inhibitors of human β-secretase. J Med Chem 2007; 50: 776–81.
Abdel-Rahman HM, Al-Karamany GS, El-Koussi NA, Youssef AF, Kiso Y . HIV protease inhibitors: peptidomimetic drugs and future perspectives. Curr Med Chem 2002; 9: 1905–22.
Xiao K, Li X, Li JY, Ma LP, Hu B, Yu HP, et al. Design, synthesis, and evaluation of Leu*Ala hydroxyethylene-based non-peptide β-secretase (BACE) inhibitors. Bioorg Med Chem 2006; 14: 4535–51.
Huey R, Morris GM, Olson AJ, Goodsell DS . A semiempirical free energy force field with charge-based desolvation. J Comput Chem 2007; 28: 1145–52.
Morris GM, Goodsell DS, Halliday RS, Huey R, Hart WE, Belew RK, et al. Automated docking using a Lamarckian genetic algorithm and an empirical binding free energy function. J Comput Chem 1998; 19: 1639–62.
Acknowledgements
Project was supported by the National Natural Science Foundation of China (Grant No 30672538).
Author information
Authors and Affiliations
Corresponding authors
Rights and permissions
About this article
Cite this article
Zhu, Yp., Xiao, K., Yu, Hp. et al. Discovery of potent β-secretase (bace-1) inhibitors by the synthesis of isophthalamide-containing hybrids. Acta Pharmacol Sin 30, 259–269 (2009). https://doi.org/10.1038/aps.2008.26
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/aps.2008.26
Keywords
This article is cited by
-
In silico screening of potential β-secretase (BACE1) inhibitors from VIETHERB database
Journal of Molecular Modeling (2022)
-
Per-Residue Energy Footprints-Based Pharmacophore Modeling as an Enhanced In Silico Approach in Drug Discovery: A Case Study on the Identification of Novel β-Secretase1 (BACE1) Inhibitors as Anti-Alzheimer Agents
Cellular and Molecular Bioengineering (2016)