Abstract
Antimicrobial resistance genes (ARG), such as extended-spectrum β-lactamase (ESBL) and carbapenemase genes, are commonly carried on plasmids. Plasmids can transmit between bacteria, disseminate globally, and cause clinically important resistance. Therefore, targeting plasmids could reduce ARG prevalence, and restore the efficacy of existing antibiotics. Cobalt complexes possess diverse biological activities, including antimicrobial and anticancer properties. However, their effect on plasmid conjugation has not been explored yet. Here, we assessed the effect of four previously characterised bis(N-picolinamido)cobalt(II) complexes lacking antibacterial activity on plasmid conjugation in Escherichia coli and Klebsiella pneumoniae. Antimicrobial susceptibility testing of these cobalt complexes confirmed the lack of antibacterial activity in E. coli and K. pneumoniae. Liquid broth and solid agar conjugation assays were used to screen the activity of the complexes on four archetypical plasmids in E. coli J53. The cobalt complexes significantly reduced the conjugation of RP4, R6K, and R388 plasmids, but not pKM101, on solid agar in E. coli J53. Owing to their promising activity, the impact of cobalt complexes was tested on the conjugation of fluorescently tagged extended-spectrum β-lactamase encoding pCTgfp plasmid in E. coli and carbapenemase encoding pKpQILgfp plasmid in K. pneumoniae, using flow cytometry. The complexes significantly reduced the conjugation of pKpQILgfp in K. pneumoniae but had no impact on pCTgfp conjugation in E. coli. The cobalt complexes did not have plasmid-curing activity, suggesting that they target conjugation rather than plasmid stability. To our knowledge, this is the first study to report reduced conjugation of clinically relevant plasmids with cobalt complexes. These cobalt complexes are not cytotoxic towards mammalian cells and are not antibacterial, therefore they could be optimised and employed as inhibitors of plasmid conjugation.
Similar content being viewed by others
Introduction
Antimicrobial resistance (AMR) is a global public health crisis that jeopardises our ability to treat infectious diseases1,2. The threat of AMR has been further compounded by the dissemination of antimicrobial resistance genes (ARGs) by mobile genetic elements such as plasmids, which can carry multiple genes encoding proteins that confer resistance to a wide range of clinically relevant antibiotics3,4. Worryingly, opportunistic pathogens like multidrug-resistant (MDR) Enterobacteriaceae, such as Escherichia coli and Klebsiella pneumoniae, often harbour plasmids that carry ARGs coding for extended-spectrum β-lactamases (ESBLs e.g., CTX-M) and carbapenemases (e.g., KPC), which confer resistance to β-lactam and carbapenem antibiotics, respectively5,6,7,8. Hence, infections caused by MDR Enterobacteriaceae have become increasingly difficult to treat due to dwindling treatment options9. Consequently, patients with carbapenem-resistant Enterobacteriaceae infections face a significantly greater risk of death compared to patients with carbapenem susceptible Enterobacteriaceae infections10. In 2017, the World Health Organisation (WHO) recognised the threat posed by carbapenem-resistant and ESBL-producing Enterobacteriaceae. It designated this pathogen group as a critical priority for which novel drugs are needed11. Therefore, there is an urgent need to develop new strategies to treat these infections, including making carbapenem-resistant and ESBL-producing Enterobacteriaceae susceptible to existing drugs and preventing the spread of AMR between bacteria.
ARGs are commonly found in conjugative AMR plasmids that can be readily shared between bacteria that occupy the same environmental niche12,13,14. The transmission of AMR plasmids via conjugation between different species of Enterobacteriaceae has been well-documented in both clinical and environmental settings15,16,17. Once Enterobacteriaceae acquire ESBL- or carbapenemase-producing AMR plasmids, they can become MDR and extremely challenging to eradicate when they cause infections18. Several different factors contribute to the prevalence and spread of AMR plasmids, including high conjugation rates, increased plasmid copy number, reduced fitness cost of plasmid carriage due to compensatory mutations, and successful plasmid and clonal group interplay19,20,21,22,23,24. Owing to the potential to transfer multiple ARGs simultaneously, their high mobility and persistence, and the significant impact they have on treatment options, AMR plasmids are a serious threat to both animal and human health.
A potential strategy to overcome the threat of carbapenem-resistant and ESBL-producing Enterobacteriaceae is by reducing the prevalence of AMR plasmids25. This could restore the efficacy of existing well-tolerated drugs and reduce the necessity of using more toxic alternatives. Compounds that target plasmids can work by removing the plasmid from a population by reducing plasmid stability (plasmid curing) and/or interfering with the conjugation process to prevent the transfer of a plasmid to a new host25,26,27. Previous studies have identified compounds with plasmid curing/conjugation inhibiting activity, ranging from phytochemicals to clinically approved drugs28,29,30,31. Except for clinically approved drugs, the majority of the previously described compounds such as biocides, DNA intercalating agents, and detergents are either toxic or have not been tested in clinically relevant AMR plasmids of Enterobacteriaceae25.
Cobalt (Co) is a trace element in the body and is essential for many biological processes, and excess amounts or deficiency of the metal can induce undesired effects32. Cobalt complexes have found usage as anticancer, antiviral, and antimicrobial agents33,34. In particular, Co(II) and Co(III) complexes have been reported to have high activities against Staphylococcus aureus and E. coli35, and fungal strains36. Previously, tris(N-picolinamido)cobalt(III) complexes were reported to have antibacterial activity against Pseudomonas and E. coli35. However, Ghandhi et al.37 reported the ESKAPE screening (CO-ADD; Community for Open Antimicrobial Drug Discovery, The University of Queensland, Australia) of a range of bis(N-picolinamido)cobalt(II) complexes that had antifungal activity but minimal antibacterial activity and importantly, no cytotoxicity against mammalian cell lines37. These results suggest the oxidation state of the cobalt complexes could induce differences in their bacterial mode of action. Although the antimicrobial effects of cobalt complexes have been explored, their impact on plasmid conjugation has not been studied to date. Here, we evaluated the activity of four previously characterised cobalt complexes37 lacking antibacterial activity for their ability to reduce the conjugation of various plasmid types in E. coli J53. The most promising compounds were then tested for their impact on the conjugation of ESBL- or carbapenemase-producing plasmids tagged with gfp by flow cytometric analysis in clinical E. coli and K. pneumoniae isolates, respectively.
Materials and methods
Strains and plasmids
All bacterial strains and plasmids used in this study are listed in Tables 1 and 2, respectively. Unless stated otherwise, all strains were grown in Luria–Bertani (LB) broth/broth with agar (Merck, Germany) at 37 °C with aeration (liquid cultures).
Determination of antibacterial susceptibility
The broth microdilution method was used to determine the minimum inhibitory concentrations (MICs) of ampicillin and the cobalt complexes Co4, Co5, Co6, and Co8 ranging from 1 to 512 µg/mL according to Clinical and Laboratory Standards Institute guidance38. Ampicillin was included as a control antibiotic for the antimicrobial susceptibility testing as the MIC values for the E. coli NCTC 10418 and S. aureus NCTC 12981 (Table 1) quality control strains are known38. The MIC values were recorded as the lowest concentration at which no bacterial growth was detected. All MICs were carried out using three biological replicates.
Growth kinetic assays
The impact of the cobalt complexes on bacterial growth was determined as previously described39. Briefly, overnight cultures (~ 109 CFU/mL) of the strains used for the conjugation assays were diluted to a starting inoculum of 106 CFU/mL in a 96-well flat bottom plate (Corning, USA). Where appropriate, the test strains were diluted in LB broth supplemented with the cobalt complexes or DMSO vehicle control to a final concentration of 100 µg/mL. Growth was monitored at OD600 at 30-min intervals for 12 h using the FLUOstar OMEGA plate reader (BMG Labtech, Germany). Three independent experiments were carried out, each consisting of three biological replicates.
Construction of the hygromycin-resistant Escherichia coli J53 recipient strain
To obtain a hygromycin-resistant E. coli J53 strain to be used as a recipient for conjugation assays, the hph gene encoding hygromycin B phosphotransferase was inserted into the phenotypically neutral attTn7 site40 using the arabinose inducible recombineering plasmid pSLTS as described previously41. Firstly, the hph gene was amplified from the pSIM18 plasmid using primers that have flanking 40 bp homology to the attTn7 site in E. coli (Supplementary Table S1). The arabinose inducible recombineering plasmid pSLTS was electroporated into E. coli J53 with subsequent electroporation of the PCR-amplified hygromycin resistance cassette. Successful recombinants were selected on LB agar supplemented with 150 µg/mL hygromycin. PCR and Sanger sequencing (Eurofins Genomics, UK) using primers that bind upstream and downstream of the recombination site (Supplementary Table S1), were used to verify the successful insertion of the hph gene at the desired genomic locus. Antimicrobial susceptibility testing was also used to verify the hygromycin-resistant phenotype (MIC > 512 µg/mL).
Liquid broth conjugation assay
The donor E. coli J53 strain with R388, pKM101, RP4 or R6K was paired with the hygromycin-resistant recipient strain E. coli J53 attTn7::hph. The liquid broth conjugation assays were performed as previously described with minor modifications42. Donor and recipient cultures were grown overnight, sub-cultures were prepared in 5 mL LB broth (1% inoculum) and grown to an OD600 of ~ 0.5. A 1 mL culture volume was pelleted, and media were replaced with LB broth to normalise the OD600 to 0.5. Equal volumes of donor and recipient strains were mixed to give a donor-to-recipient ratio of 1:1. Cultures were diluted 1:5 in LB broth containing a final concentration of 100 µg/mL of cobalt complexes or 100 µg/mL DMSO as vehicle control. These were incubated statically at 37 °C for 4 h. Cells were serially diluted in sterile phosphate-buffered saline (PBS) (10–1 to 10–6) and plated on selective media and incubated at 37 °C overnight. Transconjugant colonies carrying RP4, R6K or pKM101 were selected on LB agar supplemented with 150 µg/mL hygromycin B (PhytoTech Labs, USA) and 100 µg/mL carbenicillin (Merck, Germany). Transconjugant colonies carrying R388 were selected on LB agar supplemented with 150 µg/mL hygromycin B and 10 µg/mL trimethoprim (Merck, Germany). Conjugation frequencies (CF) were calculated using the following formula:
Three independent experiments were carried out, each one consisting of four biological replicates.
Solid agar conjugation assay
The donor E. coli J53 strains with R388, pKM101, RP4 or R6K were paired with the hygromycin-resistant recipient strain E. coli J53 attTn7::hph. The solid agar conjugation assay was performed as previously described with minor modifications43. Briefly, a 1 mL volume of overnight cultures of donor and recipient cells was pelleted, washed with LB broth, and the OD600 was adjusted to 0.5. Equal volumes of donor and recipient cells were mixed to give a donor-to-recipient ratio of 1:1. Then, 5 µL of this mixture, which contained bacteria at an OD600 of 0.5, was placed on top of 96-well round bottom plates (Corning, USA) containing 150 µL LB agar supplemented with 100 µg/mL of cobalt complexes or 100 µg/mL DMSO as vehicle control. Conjugation was carried out for 4 h at 37 °C without agitation. Bacteria were resuspended in 150 µL sterile PBS and diluted cells (10–1 to 10–6) were plated on selective media as described above and incubated at 37 °C overnight. Conjugation frequencies were calculated the same way as for the liquid conjugation assay. Three independent experiments were carried out, each one consisting of four biological replicates.
Measurement of plasmid conjugation by flow cytometry
The conjugation of pCTgfp in E. coli ST131 EC958c and pKpQILgfp in K. pneumoniae Ecl8 was measured by flow cytometry as previously described30. In our experience, bacteria grown on solid agar surfaces are less suited to flow cytometry than liquid cultures as the bacteria tend to form clumps and doublets. Therefore, the conjugation of pCTgfp in E. coli ST131 EC958c and pKpQILgfp in K. pneumoniae Ecl8 was determined in liquid broth. Briefly, 1 mL overnight cultures of the donor (E. coli with pCTgfp or K. pneumoniae with pKpQILgfp) and the recipient (E. coli or K. pneumoniae with chromosomal mCherry) strains were pelleted, washed in sterile PBS, and diluted to an OD600 of 0.5. Equal volumes of donor and recipient strains were mixed to give a donor-to-recipient ratio of 1:1. A 20 µL volume of the donor-recipient mix was inoculated into 180 µL of LB broth supplemented with a final concentration of 100 µg/mL of cobalt complexes or 100 µg/mL DMSO as vehicle control in a 96-well round bottom plate (Corning, USA). The plate was incubated at 37 °C with gentle agitation (∼100 rpm) for 4 h. Following incubation, 20 µL was removed and serially diluted 1:1000 in filter-sterilised Dulbecco’s PBS (Merck, Germany). Samples were analysed on the Attune NxT acoustic focusing flow cytometer with Autosampler (Thermo Scientific, USA). GFP emission was collected using the BL1-H channel and the mCherry emission was collected using the YL2-H channel. Plasmid conjugation was measured by quantifying the number of green fluorescent protein (GFP)-positive bacteria (donor), mCherry-positive bacteria (recipient), and GFP-positive/mCherry-positive bacteria (transconjugants). Gating strategies were exactly as previously described30. Plasmid conjugation was calculated as the number of dual fluorescent bacterial events divided by the total bacterial events relative to the DMSO control. Three independent experiments were carried out, each one consisting of four biological replicates.
Assessment of plasmid-curing activity
Overnight cultures of E. coli J53 carrying RP4, R6K, R388, or pKM101, E. coli ST131 EC958c carrying pCTgfp, and K. pneumoniae Ecl8 carrying pKpQILgfp were grown. Sub-cultures were prepared in 5 mL LB broth (5% inoculum) and grown to an OD600 of ~ 0.6. A 1 mL volume of culture was pelleted, and media were replaced with LB broth to normalise the OD600 to 0.5. A 10 µL volume of culture was inoculated into 190 µL of LB broth supplemented with a final concentration of 100 µg/mL cobalt complexes or 100 µg/mL DMSO as vehicle control in a 96-well round bottom plate (Corning, USA). The plate was then incubated at 37 °C for 24 h without agitation. Following 24 h incubation, each well was serially diluted to 10–6 in sterile PBS. A 10 µL volume of the 10–6 diluted culture was then used to passage the cells in 190 µL LB broth supplemented with a final concentration of 100 µg/mL cobalt complexes or 100 µg/mL DMSO as vehicle control in a 96-well round bottom plate for a further 24 h incubation. This dilution factor was used to impose a bottleneck on the population during passage experiments. Cells were serially diluted in sterile PBS (10–1 to 10–6) and the first (24 h) and second passage (48 h) post-inoculation were plated onto both selective media and non-selective media and incubated at 37 °C overnight. The plasmids RP4, R6K, and R388 were selected on 100 µg/mL carbenicillin, R388 on 10 µg/mL trimethoprim, and pCTgfp and pKpQILgfp on 50 µg/mL kanamycin. The percentage of plasmid persistence was calculated as
Three independent experiments were carried out, each one consisting of four biological replicates.
Statistical analysis
Unpaired t-tests were used for statistical analysis with GraphPad Prism version 10 for MacOS, San Diego, California USA, http://www.graphpad.com. Only P-values less than or equal to 0.05 were considered statistically significant.
Results
Cobalt complexes are not antibacterial
Firstly, the susceptibility of the quality control and the strains used for the plasmid conjugation assays (Table 1) to the cobalt complexes Co4, Co5, Co6, and Co8 (Supplementary Fig. S1) was determined to detect any antibacterial activity and to identify a suitable concentration to evaluate their effect on plasmid conjugation. In agreement with the previous study37, none of the tested cobalt complexes exhibited antibacterial activity against the strains tested up to a concentration of 512 µg/mL (Supplementary Table S2). Preliminary work showed that 100 µg/mL was the lowest concentration which showed activity in plasmid conjugation assays. Therefore, the effect of the cobalt complexes on plasmid conjugation was tested at 100 µg/mL. Before the plasmid conjugation assays, and to ensure 100 µg/mL had no impact on growth, bacterial growth kinetics in the presence of 100 µg/mL cobalt complexes were compared to 100 µg/mL DMSO vehicle control. These experiments showed that none of the cobalt complexes had any significant adverse effects on bacterial growth over 12 h (Supplementary Fig. S2).
Cobalt complexes affected plasmid conjugation differently in liquid broth and solid agar mating
The cobalt complexes were first screened using a panel of E. coli J53 strains carrying different plasmid types (Table 1). Plasmid conjugation frequencies (CF) are also known to differ in liquid broth and on solid surfaces44. Therefore, the effect of cobalt complexes on plasmid conjugation was tested using both liquid broth and agar mating experiments. In agreement with previous studies44,45, the IncP plasmid RP4, the IncX2 plasmid R6K, the IncW plasmid R388 and the IncN plasmid pKM101 displayed higher CFs on solid agar compared to liquid broth mating (Fig. 1 and Table 3).
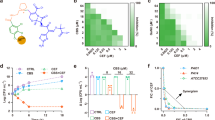
The effect of cobalt complexes on the conjugation frequencies of plasmids with different incompatibility groups in liquid LB broth and on LB agar. Conjugation frequencies of (a) the IncP plasmid RP4, (b) the IncX2 plasmid R6K, (c) the IncW R388, and (d) the IncN plasmid pKM101, from E. coli J53 to hygromycin resistant E. coli J53 attTn7::hph in the presence of 100 µg/mL DMSO vehicle control or 100 µg/mL cobalt compound after four-hour incubation. Data shown are the mean ± standard deviation of three independent experiments, each carried out with four biological replicates. Cobalt complexes that significantly affected conjugation frequency compared to DMSO control are indicated with * (p ≤ 0.05), ** (p ≤ 0.01) or *** (p ≤ 0.001). ns, not significant.
Interestingly, the cobalt complexes affected plasmid CFs differently depending on the conjugation condition. Co4 and Co5 significantly increased the CF of RP4 in liquid broth mating, whilst in solid agar mating they both significantly reduced the CF of RP4 (Fig. 1a) from 1.42 × 10–1 in DMSO control to 6.79 × 10–2 in Co4 (p = 0.0186) and 4.49 × 10–2 in Co5 (p = 0.0197) (Table 3). Whilst Co6 had no impact on the CF of RP4 in liquid broth (Fig. 1a), it significantly reduced CF in solid agar mating from 1.42 × 10–1 in DMSO control to 1.61 × 10–2 in Co6 (p = 0.0029) (Table 3). Co8 did not affect RP4 CF in liquid broth and solid agar mating (Fig. 1a). For R6K and R388, Co4 significantly reduced their CF in both conditions, but the reduction in CF was more pronounced in solid agar mating (R6K, p = 0.0062 and R388, p = 0.0048) (Fig. 1b and c). Co6 had no significant impact on R6K CF in solid agar mating (Fig. 1b) but significantly increased CF from 6.26 × 10–5 in DMSO control to 1.63 × 10–4 in liquid broth (p = 0.0146) (Table 3). Additionally, Co6 and Co8 had no significant impact on the CF of R388 in liquid broth (Fig. 1c), but significantly reduced its CF in solid agar mating from 4.19 × 10–2 in DMSO control to 9.71 × 10–3 (p = 0.0004) and 6.49 × 10–3 (p = 0.0003) in Co6 and Co8, respectively (Table 3). None of the cobalt complexes had a significant impact on the CF of pKM101 in both liquid broth and solid agar mating (Fig. 1d), indicating that IncN plasmids are possibly not targeted by cobalt complexes.
Impact of cobalt complexes on the conjugation of plasmids carrying extended-spectrum β-lactamase and carbapenemase genes
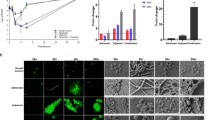
After confirming anti-plasmid activity in the J53 E. coli isolates carrying three of the four plasmids (RP4, R6K, and R388), we wanted to assess their activity in clinical bacterial isolates with plasmids conferring resistance to ESBLs and carbapenems. To test the impact of the four cobalt complexes on the conjugation of clinically relevant strains/plasmids using a previously developed liquid-conjugation assay that uses flow cytometry to quantify transconjugants30. The recipient E. coli and K. pneumoniae strains expressed a chromosomal mCherry gene whilst the donor E. coli strain carrying the IncK type plasmid pCT or the K. pneumoniae strain carrying IncFII type plasmid pKpQIL were tagged with constitutively active gfp. In this setup, transconjugant bacteria were measured based on their dual fluorescence of GFP and mCherry proteins. None of the four cobalt complexes had a significant effect on the percentage of transconjugant E. coli carrying pCTgfp compared to DMSO control (Fig. 2a), suggesting this IncK plasmid was not the target of the tested cobalt complexes in this study. On the other hand, all four cobalt complexes significantly reduced the percentage of transconjugant K. pneumoniae carrying pKpQILgfp compared to DMSO control (Fig. 2b).
The impact of cobalt complexes on the conjugation of plasmids carrying extended spectrum β-lactamase and carbapenemase genes measured by flow cytometry. The percentage of transconjugant (a) Escherichia coli ST131 EC958c carrying pCTgfp and (b) Klebsiella pneumoniae Ecl8 carrying pKpQILgfp, following incubation with 100 µg/mL cobalt compound compared to 100 µg/mL DMSO vehicle control after 4 h incubation. The data shown are the mean ± standard deviation from three independent experiments, each carried out with four biological replicates. Cobalt complexes that significantly reduced the percentage of transconjugant bacteria compared to DMSO control are indicated with ** (p ≤ 0.01) or *** (p ≤ 0.001). ns, not significant.
Cobalt complexes do not have plasmid-curing activity
To determine whether the cobalt complexes affected plasmid stability and maintenance, the impact of the cobalt complexes as plasmid curing agents was measured over 48 h. This time frame was chosen to reflect the change in transconjugant production observed over the conjugation assays (4- and 24-h), as a compound which cures a donor strain of the plasmid would generate fewer transconjugants in a conjugation assay. The DMSO (100 µg/mL) control did not affect plasmid persistence after 24 and 48 h (Fig. 3). The cobalt complexes did not display plasmid-curing activity against any of the six plasmids compared to the DMSO control after 24 and 48 h (Fig. 3). This suggested that the cobalt complexes affected the conjugation process rather than plasmid persistence in the donor bacteria.
The effect of cobalt complexes on plasmid persistence. The persistence of (a) the IncP plasmid RP4, (b) the IncX2 plasmid R6K, (c) the IncW R388, (d) the IncN plasmid pKM101, (e) the IncK plasmid pCT with tagged with a gfp gene, and (f) the IncFII plasmid pKpQIL tagged with a gfp gene, in the presence of 100 µg/mL of cobalt complexes after 24 and 48 h compared to 100 µg/mL DMSO control. The data shown are the mean ± standard deviation from three independent experiments, each conducted with four biological replicates. ns, not significant.
Discussion
The rise in AMR combined with the dwindling pipeline of new antibiotics in development warrants novel strategies to combat the AMR crisis2. Plasmids play a key role in the global dissemination of AMR genes in MDR Gram-negative bacteria3. Targeting plasmids is a novel strategy to combat AMR by reducing the prevalence of AMR genes and sensitising bacteria to existing antibiotics25. In addition, such complexes could be used in a One-Health setting by removing or reducing AMR genes in animals and/or the environment3.
Metal ion complexes represent an increasing trend in the development of antimicrobial agents46. Cobalt complexes have essential biochemical functions and have been reported to possess antibacterial, antifungal, and antiviral properties37,47. However, the impact of cobalt complexes on plasmid conjugation has never been explored. In this study, four previously characterised bis(N-picolinamido)cobalt(II) complexes (Co4, Co5, Co6, and Co8) were assessed for their ability to reduce conjugation of different plasmids and to see whether they exhibited plasmid-curing activity. The results showed that the cobalt complexes did not have a plasmid-curing effect on the tested plasmid types after 48 h. Previous studies that have investigated plasmid-curing agents reported significant plasmid elimination after 18 – 48 hr48,49,50. Hence, the cobalt complexes were likely to be affecting the conjugation process rather than ridding bacterial cells of their residing plasmid.
Plasmids persist in cells through different mechanisms including toxin-antitoxin, restriction-modification, and entry-exclusion systems51. These ensure plasmids are stably maintained in bacterial cells during cell division (e.g. through toxin-mediated killing of plasmid-free daughter cells) and conjugative plasmid transfer51. Therefore, the lack of activity of the cobalt complexes on plasmid stability could be attributed to the diverse persistence mechanisms that act to maintain plasmids during cell division. It is plausible that such mechanisms were responsible for the high level of plasmid stability seen in our assays, but effective curing compounds must be able to overcome these mechanisms.
The conjugation frequencies of RP4, R6K, R388, and pKM101 plasmids were higher on solid agar compared to liquid broth mating. This could be partly due to the differences in the dilution rates used during the experimental setup between liquid broth and solid agar mating, as well as the changes in the lifestyle of bacteria (growing in a biofilm on solid agar versus planktonic culture in liquid broth), which is known to impact conjugation rates42. The cobalt complexes were most effective at reducing plasmid conjugation on solid agar, rather than in liquid mating assays. Complexes Co4 and Co5 significantly increased the conjugation frequency of RP4 and Co6 increased the conjugation frequency of R6K in liquid broth (Fig. 1). On the other hand, complexes Co4, Co5, and Co6 significantly reduced the conjugation frequency of RP4 on solid agar (Fig. 1). Similarly, Co4, Co6, and Co8 significantly reduced R388 conjugation on solid agar but only Co4 reduced R388 conjugation in liquid broth as well (Fig. 1). The IncP plasmid RP4 and the IncW plasmid R388 have been previously shown to have constitutive rigid pilus synthesis that plays an important role in conjugation on solid surfaces44. Cobalt complexes that reduced conjugation specifically on solid agar may target plasmid-specific pilus formation/assembly to impede donor-recipient contact and transfer of single-stranded plasmid DNA52. For the IncX2 plasmid R6K, only Co4 significantly reduced its conjugation frequency in both liquid broth and on solid agar. Indeed, Co4 reduced solid agar conjugations of 3/4 plasmids, with only pKM101 (which was not affected by any compound in solid or liquid assays) showing no impact. Hence, Co4 may target a shared component of the type 4 secretion systems (T4SSs) that mediates the conjugative transfer of plasmid DNA from the donor to the recipient cell.
Co4 possesses diisothiocyanato ligands that could explain its effect on plasmid conjugation. Isothiocyanates have been shown to affect cell membrane potential and bacterial redox systems53. The morphogenesis of T4SSs requires both the proton motive force and ATP energy54. Therefore, the diisothiocyanato ligands of Co4 may potentially interfere with the assembly and function of T4SSs by disrupting the membrane potential. Cobalt has multiple effects on bacterial physiology and metabolism55. Some plasmids encode proteins with metal-binding domains that could potentially be targeted by cobalt. The single-stranded DNA-binding protein ArdC, encoded by the R388 plasmid, is important for conjugation and depends on manganese binding for its activity56. However, cobalt(II) ions also bind to the active site of ArdC and inhibit its activity56. Conjugative plasmids encode T4SS primases like TraC of RP4 that contain metallopeptidase domains and relaxases like TraI of RP4 that contain magnesium-binding sites. Cobalt(II) can occupy the same binding site as magnesium or manganese to form different coordination bonds and alter the properties of an active centre57. Hence, cobalt complexes may target metal-binding T4SS proteins and interfere with their function.
All four cobalt complexes significantly reduced the conjugation of the IncFII plasmid pKpQIL in K. pneumoniae as measured by flow cytometry (Fig. 2b). Therefore, it is plausible that they have a common target that is necessary for successful conjugative plasmid transfer, such as the type 4 secretion system58, or a common effect on K. pneumoniae. The cobalt complexes did not impact the growth kinetics of K. pneumoniae strains. The cobalt complexes had no significant effect on the pKM101 conjugation frequency in both solid agar and liquid broth (Fig. 1d). They also did not affect the conjugation of the IncK plasmid pCT in E. coli as measured by flow cytometry (Fig. 2a). These results suggested that the cobalt complexes did not target these IncK and IncN plasmids. This is possibly due to diverse elements in the conjugation apparatus between different plasmid incompatibility groups43,59,60,61.
None of the cobalt complexes exhibited antibacterial activity (Supplementary Table S2), which corroborates with the previously reported data37. Moreover, the previous study demonstrated that these cobalt complexes have no cytotoxicity towards mammalian cells37. To date, this is the first description of cobalt complexes that reduced plasmid conjugation. The efficacy of the cobalt complexes on plasmid conjugation on solid agar as opposed to in liquid broth suggests that they could be candidates for inhibitors of plasmid conjugation in solid or semi-solid environments where bacteria reside, such as biofilms on hospital surfaces, plumbing, and indwelling surfaces62,63,64. Further work involving structural modification and mechanism of action studies on bis(N-picolinamido)cobalt(II) complexes could potentially lead to the development of broad-range conjugation inhibitors.
Data availability
The authors confirm that the data supporting the findings of this study are available within the article and its supplementary materials.
References
O’Neill, J. Tackling Drug-Resistant Infections Globally: Final Report and Recommendations (Wellcome Trust, HM Government, 2016).
Murray, C. J. L. et al. Global burden of bacterial antimicrobial resistance in 2019: A systematic analysis. Lancet 399, 629–655. https://doi.org/10.1016/S0140-6736(21)02724-0 (2022).
Rozwandowicz, M. et al. Plasmids carrying antimicrobial resistance genes in Enterobacteriaceae. J. Antimicrob. Chemother. 73, 1121–1137. https://doi.org/10.1093/jac/dkx488 (2018).
Dimitriu, T. Evolution of horizontal transmission in antimicrobial resistance plasmids. Microbiology https://doi.org/10.1099/mic.0.001214 (2022).
Cottell, J. L. et al. Complete sequence and molecular epidemiology of IncK epidemic plasmid encoding blaCTX-M-14. Emerg. Infect. Dis. 17, 645–652. https://doi.org/10.3201/eid1704.101009 (2011).
Dhanji, H. et al. Dissemination of pCT-like IncK plasmids harboring CTX-M-14 extended-spectrum beta-lactamase among clinical Escherichia coli isolates in the United Kingdom. Antimicrob. Agents Chemother. 56, 3376–3377. https://doi.org/10.1128/AAC.00313-12 (2012).
Leavitt, A., Chmelnitsky, I., Carmeli, Y. & Navon-Venezia, S. Complete nucleotide sequence of KPC-3-encoding plasmid pKpQIL in the epidemic Klebsiella pneumoniae sequence type 258. Antimicrob. Agents Chemother. 54, 4493–4496. https://doi.org/10.1128/AAC.00175-10 (2010).
Doumith, M. et al. Major role of pKpQIL-like plasmids in the early dissemination of KPC-type carbapenemases in the UK. J. Antimicrob. Chemother. 72, 2241–2248. https://doi.org/10.1093/jac/dkx141 (2017).
Bassetti, M., Peghin, M., Vena, A. & Giacobbe, D. R. Treatment of infections due to MDR Gram-negative bacteria. Front. Med. 6, 74. https://doi.org/10.3389/fmed.2019.00074 (2019).
Zhou, R. et al. Impact of carbapenem resistance on mortality in patients infected with Enterobacteriaceae: A systematic review and meta-analysis. BMJ Open 11, e054971. https://doi.org/10.1136/bmjopen-2021-054971 (2021).
Tacconelli, E. et al. Discovery, research, and development of new antibiotics: The WHO priority list of antibiotic-resistant bacteria and tuberculosis. Lancet Infect. Dis. 18, 318–327. https://doi.org/10.1016/S1473-3099(17)30753-3 (2018).
Holmes, A. H. et al. Understanding the mechanisms and drivers of antimicrobial resistance. Lancet 387, 176–187. https://doi.org/10.1016/S0140-6736(15)00473-0 (2016).
Bottery, M. J. Ecological dynamics of plasmid transfer and persistence in microbial communities. Curr. Opin. Microbiol. 68, 102152. https://doi.org/10.1016/j.mib.2022.102152 (2022).
McInnes, R. S., McCallum, G. E., Lamberte, L. E. & van Schaik, W. Horizontal transfer of antibiotic resistance genes in the human gut microbiome. Curr. Opin. Microbiol. 53, 35–43. https://doi.org/10.1016/j.mib.2020.02.002 (2020).
Weingarten, R. A. et al. Genomic analysis of hospital plumbing reveals diverse reservoir of bacterial plasmids conferring carbapenem resistance. mBio https://doi.org/10.1128/mBio.02011-17 (2018).
Stecher, B. et al. Gut inflammation can boost horizontal gene transfer between pathogenic and commensal Enterobacteriaceae. Proc. Natl. Acad. Sci. USA 109, 1269–1274. https://doi.org/10.1073/pnas.1113246109 (2012).
Gona, F. et al. In vivo multiclonal transfer of bla(KPC-3) from Klebsiella pneumoniae to Escherichia coli in surgery patients. Clin. Microbiol. Infect. 20, O633-635. https://doi.org/10.1111/1469-0691.12577 (2014).
Bassetti, M., Peghin, M. & Pecori, D. The management of multidrug-resistant Enterobacteriaceae. Curr. Opin. Infect. Dis. 29, 583–594. https://doi.org/10.1097/QCO.0000000000000314 (2016).
Buckner, M. M. C. et al. Clinically relevant plasmid-host interactions indicate that transcriptional and not genomic modifications ameliorate fitness costs of Klebsiella pneumoniae carbapenemase-carrying plasmids. mBio https://doi.org/10.1128/mBio.02303-17 (2018).
Cottell, J. L., Webber, M. A. & Piddock, L. J. Persistence of transferable extended-spectrum-beta-lactamase resistance in the absence of antibiotic pressure. Antimicrob. Agents Chemother. 56, 4703–4706. https://doi.org/10.1128/AAC.00848-12 (2012).
Coque, T. M. et al. Dissemination of clonally related Escherichia coli strains expressing extended-spectrum beta-lactamase CTX-M-15. Emerg. Infect. Dis. 14, 195–200. https://doi.org/10.3201/eid1402.070350 (2008).
Peirano, G. & Pitout, J. D. Molecular epidemiology of Escherichia coli producing CTX-M beta-lactamases: The worldwide emergence of clone ST131 O25:H4. Int. J. Antimicrob. Agents 35, 316–321. https://doi.org/10.1016/j.ijantimicag.2009.11.003 (2010).
Whitmer, G. R., Moorthy, G. & Arshad, M. The pandemic Escherichia coli sequence type 131 strain is acquired even in the absence of antibiotic exposure. PLoS Pathog. https://doi.org/10.1371/journal.ppat.1008162 (2019).
Dimitriu, T., Matthews, A. C. & Buckling, A. Increased copy number couples the evolution of plasmid horizontal transmission and plasmid-encoded antibiotic resistance. Proc. Natl. Acad. Sci. USA https://doi.org/10.1073/pnas.2107818118 (2021).
Buckner, M. M. C., Ciusa, M. L. & Piddock, L. J. V. Strategies to combat antimicrobial resistance: Anti-plasmid and plasmid curing. FEMS Microbiol. Rev. 42, 781–804. https://doi.org/10.1093/femsre/fuy031 (2018).
Vrancianu, C. O., Popa, L. I., Bleotu, C. & Chifiriuc, M. C. Targeting plasmids to limit acquisition and transmission of antimicrobial resistance. Front. Microbiol. 11, 761. https://doi.org/10.3389/fmicb.2020.00761 (2020).
Getino, M. & de la Cruz, F. Natural and artificial strategies to control the conjugative transmission of plasmids. Microbiol. Spectr. https://doi.org/10.1128/microbiolspec.MTBP-0015-2016 (2018).
Oyedemi, B. O., Kotsia, E. M., Stapleton, P. D. & Gibbons, S. Capsaicin and gingerol analogues inhibit the growth of efflux-multidrug resistant bacteria and R-plasmids conjugal transfer. J. Ethnopharmacol. 245, 111871. https://doi.org/10.1016/j.jep.2019.111871 (2019).
Patwardhan, R. B., Dhakephalkar, P. K., Chopade, B. A., Dhavale, D. D. & Bhonde, R. R. Purification and characterization of an active principle, lawsone, responsible for the plasmid curing activity of Plumbago zeylanica root extracts. Front. Microbiol. 9, 2618. https://doi.org/10.3389/fmicb.2018.02618 (2018).
Buckner, M. M. C. et al. HIV drugs inhibit transfer of plasmids carrying extended-spectrum beta-lactamase and carbapenemase genes. mBio https://doi.org/10.1128/mBio.03355-19 (2020).
Shriram, V. et al. A potential plasmid-curing agent, 8-epidiosbulbin E acetate, from Dioscorea bulbifera L. against multidrug-resistant bacteria. Int. J. Antimicrob. Agents 32, 405–410. https://doi.org/10.1016/j.ijantimicag.2008.05.013 (2008).
Czarnek, K., Terpilowska, S. & Siwicki, A. K. Selected aspects of the action of cobalt ions in the human body. Cent. Eur. J. Immunol. 40, 236–242. https://doi.org/10.5114/ceji.2015.52837 (2015).
Munteanu, C. R. & Suntharalingam, K. Advances in cobalt complexes as anticancer agents. Dalton Trans. 44, 13796–13808. https://doi.org/10.1039/C5DT02101D (2015).
Chang, E. L., Simmers, C. & Knight, D. A. Cobalt complexes as antiviral and antibacterial agents. Pharmaceuticals 3, 1711–1728. https://doi.org/10.3390/ph3061711 (2010).
Mishra, A., Kaushik, N. K., Verma, A. K. & Gupta, R. Synthesis, characterization and antibacterial activity of cobalt(III) complexes with pyridine-amide ligands. Eur. J. Med. Chem. 43, 2189–2196. https://doi.org/10.1016/j.ejmech.2007.08.015 (2008).
Gaëlle, D. S. Y., Yufanyi, D. M., Jagan, R. & Agwara, M. O. Synthesis, characterization and antimicrobial properties of cobalt(II) and cobalt(III) complexes derived from 1,10-phenanthroline with nitrate and azide co-ligands. Cogent Chem. 2, 1253201. https://doi.org/10.1080/23312009.2016.1253201 (2016).
Ghandhi, L. H. D., Bidula, S., Pask, C. M., Lord, R. M. & McGowan, P. C. Bis(N-picolinamido)cobalt(II) complexes display antifungal activity toward Candida albicans and Aspergillus fumigatus. ChemMedChem 16, 3210–3221. https://doi.org/10.1002/cmdc.202100159 (2021).
CLSI. Performance Standards for Antimicrobial Susceptibility Testing. 32 edn, vol. 42 (Clinical and Laboratory Standards Institute, 2022).
Alav, I., Bavro, V. N. & Blair, J. M. A. A role for the periplasmic adaptor protein AcrA in vetting substrate access to the RND efflux transporter AcrB. Sci. Rep. 12, 4752. https://doi.org/10.1038/s41598-022-08903-9 (2022).
Holden, E. R., Wickham, G. J., Webber, M. A., Thomson, N. M. & Trampari, E. Donor plasmids for phenotypically neutral chromosomal gene insertions in Enterobacteriaceae. Microbiology 166, 1115–1120. https://doi.org/10.1099/mic.0.000994 (2020).
Kim, J., Webb, A. M., Kershner, J. P., Blaskowski, S. & Copley, S. D. A versatile and highly efficient method for scarless genome editing in Escherichia coli and Salmonella enterica. BMC Biotechnol. 14, 84. https://doi.org/10.1186/1472-6750-14-84 (2014).
Element, S. J. et al. Growth in a biofilm promotes conjugation of a bla (NDM-1)-bearing plasmid between Klebsiella pneumoniae strains. mSphere 8, e0017023. https://doi.org/10.1128/msphere.00170-23 (2023).
Getino, M. et al. Synthetic fatty acids prevent plasmid-mediated horizontal gene transfer. mBio 6, e01032-01015. https://doi.org/10.1128/mBio.01032-15 (2015).
Bradley, D. E., Taylor, D. E. & Cohen, D. R. Specification of surface mating systems among conjugative drug resistance plasmids in Escherichia coli K-12. J. Bacteriol. 143, 1466–1470. https://doi.org/10.1128/jb.143.3.1466-1470.1980 (1980).
del Campo, I. et al. Determination of conjugation rates on solid surfaces. Plasmid 67, 174–182. https://doi.org/10.1016/j.plasmid.2012.01.008 (2012).
Frei, A. et al. Metal complexes as a promising source for new antibiotics. Chem. Sci. 11, 2627–2639. https://doi.org/10.1039/c9sc06460e (2020).
Heffern, M. C., Yamamoto, N., Holbrook, R. J., Eckermann, A. L. & Meade, T. J. Cobalt derivatives as promising therapeutic agents. Curr. Opin. Chem. Biol. 17, 189–196. https://doi.org/10.1016/j.cbpa.2012.11.019 (2013).
Kwapong, A. A., Stapleton, P. & Gibbons, S. Inhibiting plasmid mobility: The effect of isothiocyanates on bacterial conjugation. Int. J. Antimicrob. Agents. 53, 629–636. https://doi.org/10.1016/j.ijantimicag.2019.01.011 (2019).
Riva, S., Fietta, A., Berti, M., Silvestri, L. G. & Romero, E. Relationships between curing of the F episome by rifampin and by acridine orange in Escherichia coli. Antimicrob. Agents Chemother. 3, 456–462. https://doi.org/10.1128/aac.3.4.456 (1973).
Bouanchaud, D. H. & Chabbert, Y. A. Practical effectiveness of agents curing r factors and plasmids. Ann. N. Y. Acad. Sci. 182, 305–311. https://doi.org/10.1111/j.1749-6632.1971.tb30666.x (1971).
Bahl, M. I., Hansen, L. H. & Sørensen, S. J. In Horizontal Gene Transfer: Genomes in Flux (eds Gogarten, M. B., Gogarten, J. P. & Olendzenski, L. C.) 73–102 (Humana Press, 2009).
Ou, J. T. & Anderson, T. F. Role of pili in bacterial conjugation. J. Bacteriol. 102, 648–654. https://doi.org/10.1128/jb.102.3.648-654.1970 (1970).
Sofrata, A. et al. Benzyl isothiocyanate, a major component from the roots of Salvadora persica is highly active against gram-negative bacteria. PLoS ONE 6, e23045. https://doi.org/10.1371/journal.pone.0023045 (2011).
Christie, P. J. & Cascales, E. Structural and dynamic properties of bacterial type IV secretion systems (review). Mol. Membr. Biol. 22, 51–61. https://doi.org/10.1080/09687860500063316 (2005).
Okamoto, S. & Eltis, L. D. The biological occurrence and trafficking of cobalt. Metallomics 3, 963–970. https://doi.org/10.1039/c1mt00056j (2011).
Gonzalez-Montes, L., Del Campo, I., Garcillan-Barcia, M. P., de la Cruz, F. & Moncalian, G. ArdC, a ssDNA-binding protein with a metalloprotease domain, overpasses the recipient hsdRMS restriction system broadening conjugation host range. PLoS Genet. 16, e1008750. https://doi.org/10.1371/journal.pgen.1008750 (2020).
Khrustalev, V. V. et al. Cobalt(II) cation binding by proteins. Metallomics 11, 1743–1752. https://doi.org/10.1039/c9mt00205g (2019).
Boudaher, E. & Shaffer, C. L. Inhibiting bacterial secretion systems in the fight against antibiotic resistance. MedChemComm 10, 682–692. https://doi.org/10.1039/c9md00076c (2019).
Alvarez-Martinez, C. E. & Christie, P. J. Biological diversity of prokaryotic type IV secretion systems. Microbiol. Mol. Biol. Rev. 73, 775–808. https://doi.org/10.1128/MMBR.00023-09 (2009).
Smillie, C., Garcillan-Barcia, M. P., Francia, M. V., Rocha, E. P. & de la Cruz, F. Mobility of plasmids. Microbiol. Mol. Biol. Rev. 74, 434–452. https://doi.org/10.1128/MMBR.00020-10 (2010).
Getino, M. et al. Tanzawaic acids, a chemically novel set of bacterial conjugation inhibitors. PLoS ONE 11, e0148098. https://doi.org/10.1371/journal.pone.0148098 (2016).
Soto-Giron, M. J. et al. Biofilms on hospital shower hoses: Characterization and implications for nosocomial infections. Appl. Environ. Microbiol. 82, 2872–2883. https://doi.org/10.1128/AEM.03529-15 (2016).
Pelling, H. et al. Bacterial biofilm formation on indwelling urethral catheters. Lett. Appl. Microbiol. 68, 277–293. https://doi.org/10.1111/lam.13144 (2019).
Lindsay, D. & von Holy, A. Bacterial biofilms within the clinical setting: What healthcare professionals should know. J. Hosp. Infect. 64, 313–325. https://doi.org/10.1016/j.jhin.2006.06.028 (2006).
Mortelmans, K. E. & Stocker, B. A. Ultraviolet light protection, enhancement of ultraviolet light mutagenesis, and mutator effect of plasmid R46 in Salmonella Typhimurium. J. Bacteriol. 128, 271–282. https://doi.org/10.1128/jb.128.1.271-282.1976 (1976).
Garcia-Fernandez, A. et al. Multilocus sequence typing of IncN plasmids. J. Antimicrob. Chemother. 66, 1987–1991. https://doi.org/10.1093/jac/dkr225 (2011).
Datta, N. & Hedges, R. W. Trimethoprim resistance conferred by W plasmids in Enterobacteriaceae. J. Gen. Microbiol. 72, 349–355. https://doi.org/10.1099/00221287-72-2-349 (1972).
Revilla, C. et al. Different pathways to acquiring resistance genes illustrated by the recent evolution of IncW plasmids. Antimicrob. Agents Chemother. 52, 1472–1480. https://doi.org/10.1128/AAC.00982-07 (2008).
Kontomichalou, P., Mitani, M. & Clowes, R. C. Circular R-factor molecules controlling penicillinase synthesis, replicating in Escherichia coli under either relaxed or stringent control. J. Bacteriol. 104, 34–44. https://doi.org/10.1128/jb.104.1.34-44.1970 (1970).
Dobiasova, H. & Dolejska, M. Prevalence and diversity of IncX plasmids carrying fluoroquinolone and beta-lactam resistance genes in Escherichia coli originating from diverse sources and geographical areas. J. Antimicrob. Chemother. 71, 2118–2124. https://doi.org/10.1093/jac/dkw144 (2016).
Lowbury, E. J. L., Lilly, H. A., Kidson, A., Ayliffe, G. A. J. & Jones, R. J. Sensitivity of Pseudomonas aeruginosa to antibiotics: Emergence of strains highly resistant to carbenicillin. Lancet 294, 448–452. https://doi.org/10.1016/S0140-6736(69)90163-9 (1969).
Popowska, M. & Krawczyk-Balska, A. Broad-host-range IncP-1 plasmids and their resistance potential. Front. Microbiol. 4, 44. https://doi.org/10.3389/fmicb.2013.00044 (2013).
Chan, W. et al. A recombineering based approach for high-throughput conditional knockout targeting vector construction. Nucleic Acids Res. 35, e64. https://doi.org/10.1093/nar/gkm163 (2007).
Acknowledgements
I.A. and M.M.C.B. were funded by the MRC grant MR/V009885/1 (New Investigator Research Grant to M.M.C.B.). H.P. was funded by a Frank Kerr undergraduate research award. R.M.L. was funded by the University of East Anglia start-up and the UKRI FLF MR/T041315/1.
Author information
Authors and Affiliations
Contributions
R.L. provided cobalt complexes. I.A. carried out liquid broth, solid agar, and flow cytometry conjugation assays. P.P. carried out liquid broth conjugation and plasmid persistence assays. P.E.d.R carried out the antimicrobial susceptibility testing and H.P carried out the growth kinetic assays. M.M.C.B. conceived the project and oversaw the work. S.G. provided experimental input. I.A. and M.M.C.B drafted the manuscript. All authors contributed feedback on the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Alav, I., Pordelkhaki, P., de Resende, P.E. et al. Cobalt complexes modulate plasmid conjugation in Escherichia coli and Klebsiella pneumoniae. Sci Rep 14, 8103 (2024). https://doi.org/10.1038/s41598-024-58895-x
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-024-58895-x
Keywords
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.