Abstract
The identification of eye diseases and their progression often relies on a clear visualization of the anatomy and on different metrics extracted from Optical Coherence Tomography (OCT) B-scans. However, speckle noise hinders the quality of rapid OCT imaging, hampering the extraction and reliability of biomarkers that require time series. By synchronizing the acquisition of OCT images with the timing of the cardiac pulse, we transform a low-quality OCT video into a clear version by phase-wrapping each frame to the heart pulsation and averaging frames that correspond to the same instant in the cardiac cycle. Here, we compare the performance of our one-cycle denoising strategy with a deep-learning architecture, Noise2Noise, as well as classical denoising methods such as BM3D and Non-Local Means (NLM). We systematically analyze different image quality descriptors as well as region-specific metrics to assess the denoising performance based on the anatomy of the eye. The one-cycle method achieves the highest denoising performance, increases image quality and preserves the high-resolution structures within the eye tissues. The proposed workflow can be readily implemented in a clinical setting.
Similar content being viewed by others
Introduction
Technological advances in OCT imaging have allowed unprecedented resolution and signal-to-noise ratio for the visualization of detailed anatomical structures in the eye that are relevant to diagnosis. Single B-scans, 3-dimensional volume reconstructions and even the vasculature are routinely used to extract metrics that allow clinicians to identify oculopathies and monitor their progression. In the case of glaucoma, such quantifiable biomarkers include retinal nerve fiber layer (RNFL) thickness, cup-to-disk ratio, neuroretinal rim area, disk area, and ganglion cell layer thickness, to quantify disease stage and progression1,2. Much of current glaucoma research is focused on the mechanical properties of the eye, which can be extracted from the dynamics of ocular tissue. Hence the need for reliable methods that preserve the high quality of time series images obtained with a short integration time compatible with the clinical setting.
Intraocular pressure (IOP) is the only actionable risk factor for glaucoma patients. Treatments, whether drugs, laser, or surgery, all aim to decrease pressure to limit the progression of the disease2. However, not all glaucoma subjects have elevated IOP and increased IOP is not necessarily an indicator of glaucomatous damage3. Furthermore, the mechanisms and pathogenesis of glaucoma are still unclear2 and research aims to find other biomarkers, straying away from IOP-based descriptors. The characterization of ocular biomechanics, and particularly the calculation of stress and strain in retinal tissues driven by several stimuli was characterized in numerical simulations based on finite element modeling4,5,6, or in non-human primates subjected to acutely elevated IOP7,8,9. Human subjects with high and normal-tension glaucoma were studied using a digital volume correlation technique to map deformations10, a 3D tracking algorithm based on optimizing numerical transformations of OCT volumes was used to quantify strains in optic nerves of glaucoma subjects after IOP reduction by trabeculectomy11, and displacement maps were obtained to characterize the behavior of the retina and optic nerve head in normal and myopia cohorts subjected to optic nerve physiological traction due to eye movement12.
The cardiac and ocular systems are closely intertwined, and many believe that one system can bring information to the other13,14, for example by imaging retinal structures to enhance cardiovascular pathologies detection protocols15,16,17,18. OCT angiography utilizes the variation of the OCT signal taken at the same cross-section of the eye to visualize ocular vasculature19. This visualization helps clinicians in identifying several pathologies, from age related macular degeneration to diabetic retinopathy and more. Doppler OCT enables the visualization and quantification of retinal blood flow, in addition to the structural anatomy given by OCT20. Another study used a pulse oximeter to monitor retinal blood flow using OCT imaging21. A second group, using a methodology similar to the one proposed here, used the cardiac signal of an additional pulse oximeter to improve the sampling of pulsatile blood flow using lower frame-rate systems in Doppler OCT applications22.
Advances in OCT engineering have enabled faster frame rates and larger fields of view, enabling the performance of dynamic imaging of ocular structures, such as the retina, optic nerve head (ONH) and the lamina cribrosa. Dynamic imaging and analysis help to understand physiological changes due to pulsatile blood flow deformations and pulsatile IOP changes23. However, speckle noise significantly hinders the quality of these images. Biomarkers computed on the scans themselves, i.e., RNFL thickness, cup-to-disk ratio, and even new approaches that quantify local strain, are all dependent on signal quality and are severely affected by noise. Averaging successive B-scans, either temporally or spatially, can remove the noise naturally present in OCT images and is the most efficient noise removal technique used to this day in clinical devices24. Yet, this averaging cannot be applied naively to suppress noise in movies, as it erases the small variations due to tissue movement and temporal changes.
Several groups have focused on OCT denoising methods in recent years, most of them using new and improved deep learning architectures. The Noise2Noise architecture25 was first introduced for non-medical images and later applied to OCT scans26. The Noise2void architecture was proposed as a modification of Noise2Noise27, where the unsupervised training is done without the need for paired noisy images, instead using a blind-spot masking scheme with single noisy images. The latter model was applied to OCT scans as well28. Researchers proposed a conditional generative adversarial network to produce contrast-enhanced and speckle-reduced scans29. Another study put forward a convolutional neural network (CNN) trained to learn the difference between noisy OCT scans (with added Gaussian noise) and their corresponding multi-frame OCT scan30. Similarly, one group trained a CNN (a modified deep CNN) to denoise scans based on their multi-frame denoised versions31. This special architecture used a multi-scale structural similarity index as the loss function to obtain denoised images that preserve the structural information present in the images. Although all these methods have been applied to static OCT images or volumes, they could potentially be adapted to dynamic imaging.
Several drawbacks can be found in these methods: a large quantity of data is needed to train these networks without overfitting, they are complex and require significant resources (large amounts of memory, multiple high-performing GPUs), and they lack ground truth images for training supervised architectures. While the trained networks enable fast inference times, the main challenge of using deep learning methods, especially those using a generative workflow, is the lack of reliability of the resulting pixel intensity values. The black-box problem inherent in such models is well known and is especially important in biomedical imaging32,33,34,35. Measurements computed from the scans will influence diagnosis and treatment, and therefore the reliability of the pixel values in denoised scans is paramount.
Using the heart pulsation captured with a simple pulse oximeter synchronized to the OCT video acquisition, we propose instead a video denoising pipeline based on averaging images corresponding to the same structures taken at equivalent instants in the cardiac cycle. Our workflow (one-cycle video denoising) is particularly easy to implement in a clinical setting as it only requires a pulse sensor, and the computational workload is very light. Furthermore, all manipulations to the scans can be interpreted to support reliability of the numerical metrics extracted. The implementation details of our method are presented in the methods below and the code is made available in a public repository. We compare our denoising methods performance with four other denoising techniques: BM3D36,37, Non-Local Means (NLM)38, and a deep-learning architecture, Noise2Noise, trained in unsupervised and supervised ways. This architecture was selected for the comparison due to its well-established denoising capability and straight-forward implementation. The capability of neural networks to develop high complexity algorithms could allow the development of a single scan denoising scheme producing high quality images. Furthermore, we present a set of global and region-specific image quality metrics to estimate the denoising performance in terms of tissue anatomy in the retina and optic nerve head.
Results
We used ten metrics (five global and five local, as described in Sections “Global image quality quantification”, “Region-specific image quality quantification” and “Movement preservation quantification” below) to assess the denoising performance and compare all methods both globally and in specific anatomically defined ROIs. The Wilcoxon test was applied between the one-cycle workflow metrics and every other method (including no denoising, i.e., the original B-scans). Every metric was computed on 100 different images from each video (i.e., each of the 21 scanned eyes). For our method, this was simply all the frames of the one-cycle video. For the other denoising methods, the 100 images per subject were randomly selected from the test sets.
Figure 1 shows examples of the output of the algorithms applied to individual B-scans, while the whole movies for these same subjects can be found in the Supplementary material.
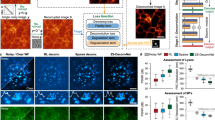
Visual comparison of the denoised scans with all five denoising workflows. Qualitative comparison between the original B-scans (left) and the denoised scans using the one-cycle workflow (second column), the Noise2Noise (N2N) model (third column), Noise2Clean (N2C) model (fourth column), BM3D (fifth column) and NLM (sixth column) on two different eyes (rows). Separate rows show the whole scan after denoising as well as a zoomed portion of the RNFL to RPE layers of the retina. These B-scans were not part of the training sets of the neural networks.
Image quality metrics: all five methods show improved contrast and higher similarity between the scans
We measured the contrast and speckle noise in the images by calculating the CNR in the RPE, and the preservation of image information by evaluating the Mattes Mutual Information (MMI) on whole B-scans. Noise reduction was also quantified by analyzing the intensity of the 2-dimensional Fast Fourier Transform (FFT) outside of an ellipse representing the resolution limit of our OCT machine (Fig. 2); no useful information can be contained in the spatial frequency spectrum beyond the resolution of the images. We found that the SNR and CNR computed in the choroid, RNFL and RPE regions yielded quite similar results; thus, only the CNR is discussed further, but both metrics are presented in the Supplementary Fig. A1.
Image quality after denoising characterized by three different global metrics. All metrics were computed in the original B-scans (black violin plots) and for the five denoising methods studied (one-cycle workflow in red, N2N in turquoise, N2C in lavender, BM3D in yellow and NLM in orange). The Wilcoxon test was applied between the one-cycle workflow metrics and every other method (including no denoising, i.e., the original B-scans). Significant differences between methods are indicated by horizontal bars; asterisks indicate statistical strength. (A) CNR computed in the RPE. (B) MMI computed in whole images. (C) Mean intensity of the FFT spectrum beyond theoretical OCT resolution (D). (D) Ellipse (green) constructed using the axial and transverse OCT resolutions for the values of the half axes (pink and blue dots). The intensities of the frequency spectrum are averaged outside this ellipse.
Figure 2 shows the denoising performance as measured by the global image metrics. In Fig. 2A, all five denoising methods show similar CNR. A significant difference was found between the original B-scans and the one-cycle workflow, but none was found between the one-cycle method and any other denoising method. (The Wilcoxon test results are seen at the top the graphs: the horizontal bars and asterisks indicate differences between pairs of methods and their statistical strength). There is a fold-change increase in CNR of 1.57, 1.41, 1.31, 1.56 and 1.55 between the original B-scans and the one-cycle, N2N, N2C, BM3D and NLM denoising methods respectively (fold changes were computed between the median values, with the methods always in the same order: one-cycle, N2N, N2C, BM3D and NLM). The MMI (Fig. 2B) increases by fold changes of 1.27, 1.19, 1.22, 1.24 and 1.25. While the trend is analogous to the CNR, a smaller but significant difference was found between the one-cycle and N2N denoised images. The high frequency components outside the theoretical limits of the OCT resolution display a significant reduction after denoising (fold change decreases of 5.59, 7.46, 9.03, 9.71 and 4.13). All methods showed reduction of signal outside the OCT resolution; a significant decrease was also found between the one-cycle denoised images and each of the denoising methods. Note however that the NLM method doesn’t achieve the same reduction with a fold change of only 4.13 compared to other methods, indicating that a higher quantity of high-frequency noise persists after denoising.
The CNR and MMI both increase significantly using all five denoising techniques, indicating improved contrast and highest similarity between the scans after denoising. The signal present in the FFT spectrum outside the theoretical OCT resolution decreases for every denoising technique, which is consistent with reduced speckle noise. However, there is also a significant decrease in that metric between the one-cycle, N2N, N2C and BM3D denoised images close to the resolution limit. This highlights the blurring of the images by both neural networks and BM3D method (visible also in Fig. 1). Thus, while the speckle noise is reduced, denoising produced by those strategies induces a loss of real spatial information, which can also be seen in the retina-specific metrics (Fig. 3G,H). These three global image quality metrics assess the overall quality improvement of the denoised images by all five methods but are not able to discriminate between them or explain certain effects remarked on specific regions of the scans.
Region-specific assessment of denoising using five different metrics. All metrics were computed in the original B-scans (black violin plots) and the five denoising methods studied (one-cycle workflow in red, N2N in turquoise, N2C in lavender, BM3D in yellow and NLM in orange). The Wilcoxon test was applied between the one-cycle workflow metrics and every other method (including no denoising, i.e., the original B-scans). Significant differences between methods are indicated by horizontal bars; asterisks indicate statistical strength. Horizontal dotted lines show the expected values of the metrics when applicable. (A,C,F) Region considered to compute the metrics and an example. (A,B) Variation of pixel intensity in the sclera (expected value = 0); (C–E) metrics in the vitreous humor (expected mean and variation of 0); (F–H) number of retinal layers detected and average thickness (ideally 11 detected layers with a mean thickness of 50 µm).
Region-specific metrics: retinal biomarkers are blurred by CNN-based denoising strategies
Figure 3 shows the results of comparing the denoising methods based on region-specific metrics. The standard variation computed in the sclera, expected to be close to zero (Fig. 3B), clearly decreases between the original B-scans and all denoising strategies. While the trend is similar between the N2C and one-cycle denoised images, a significant difference is found between the one-cycle images and all other denoised images. The fold change decreases for this metric are of 1.83, 0.96 (increase of 1.04), 1.10, 1.50 and 1.55.
Both the mean and standard deviation of the pixel intensities in the vitreous humor should be close to zero (Fig. 3D,E). All denoising methods cause an increase in the mean pixel intensity values, with fold changes of 1.15, 1.16, 1.95, 1.04 and 1.02. The standard deviation is more discriminative between the denoising methods, with fold change decreases of 4.63, 2.20, 1.89, 3.61 and 4.06.
We assumed the retina is composed of 11 layers with an average thickness of approximately 50 µm. This metric is highly discriminative between denoising methods, with median values of 7, 5, 3, 8 and 9 layers detected (fold change decrease of 2.46, 3.37, 7.04, 2.38 and 1.93) and 51, 77, 151, 45 and 35 µm (fold change increase of 2.77, 4.20, 8.28, 2.44 and 1.92) for their respective average thickness (Fig. 3G,H).
Ocular structures of homogenous tissue characteristics (sclera and vitreous humor) are denoised by all strategies. The one-cycle workflow produces images with small standard deviations, nevertheless increasing the average pixel intensity in the vitreous humor compared to raw data (Fig. 3D,E). The performance of the N2N network is relatively close to the one-cycle workflow, however this network shows significant increases in the mean pixel intensity in the sclera as well as in standard variation in the vitreous humor compared to the proposed method. While the variation in the sclera is not significantly different between the one-cycle and N2C-denoised images, the performance in the vitreous humor, both for mean intensity and standard deviation, is significantly worse for the supervised network. The BM3D and NLM methods achieve slightly better performance in terms of standard deviation of pixel intensities in the sclera and vitreous humor.
Distinguishing the layered structure of the retina, a key clinical landmark, is best achieved with the non-AI denoising strategies. A lower-than-expected number of layers is found for the N2N model, and an abnormally low number of layers for the N2C model. While the extracted number of layers for the one-cycle denoised images is higher, it still falls short of the expected value. This is mainly due to the low contrast between certain layers (e.g., the ganglion cell and inner plexiform layers) that are often detected as one homogeneous layer, even in highly contrasted denoised images. This phenomenon is noticed for the BM3D denoising method, while the layer detection in the NLM-denoised scans is closest to the expected values. We observe the same trend for the mean distance detected between layers. While the one-cycle method yields an average of 51 µm between layers, BM3D of 45 µm, NLM of 35 µm (lower than expected) and the N2N method a value of 77 µm (higher than expected), the N2C method produces an abnormal mean distance of 151 µm. This is caused by the excessive blurring induced by the AI strategies, resulting in fewer peaks with larger distances between them; in the noisy original scans, the opposite situation yields many peaks and smaller distances.
Movement preservation: AI, BM3D and NLM denoising strategies hide physiologically induced movement in scans
Denoising strategies must preserve true intensities of the scans, the anatomy of tissues and, in temporal acquisitions, allow quantification of movement of these tissues. In order to benchmark this capacity, we calculated the tissue displacement fields from artificially deformed images, and using consecutive frames of a movie acquired only 10 ms apart. Figure 4A shows the average median relative error after inducing a Gaussian expansion of the choroid of 5 pixels of amplitude and 20 pixels of standard deviation on consecutive pairs of images after denoising. This deformation metric illustrates fold changes between every metric and the original B-scans cohort of 3.69, 1.13 and 1.24 decreases for the one-cycle, N2N and N2C methods respectively, and of 1.35 and 1.35 increases for BM3D and NLM. The one-cycle method outshines the other denoising methods in preserving an induced deformation in pairs of images. The neural networks perform slightly better than the noisy scans, while the traditional methods seem to modify the induced deformation (if they erase, amplify, or misdirect the deformation was not studied). Figure 4B assesses the displacement present between pairs of consecutive frames after denoising by all strategies. Consecutive scans are obtained at 100 Hz and, as such, should contain small but non-null tissue deformation. Figure 4B shows an average median displacement in the order of a pixel for every denoising strategy.
Movement preservation after denoising with the five different strategies. (A) The average absolute error between an induced and a computed displacement field was calculated for 100 pairs of images. The metric was computed in the original B-scans (black violin plots) and the five denoising methods studied (one-cycle workflow in red, N2N in turquoise, N2C in lavender, BM3D in yellow and NLM in orange). The Wilcoxon test was applied between the one-cycle workflow metrics and every other method (including no denoising, i.e., the original B-scans). Significant differences between methods are indicated by horizontal bars; asterisks indicate statistical strength. (B) The average absolute median displacement was computed between consecutive frames after denoising, for 100 pairs of denoised images by each strategy.
Discussion
All five methods achieve a significant reduction of speckle noise, but a closer look at the results reveals important differences. Overall, the one-cycle workflow produces the highest quality scans, in terms of speckle noise reduction, definition of ocular structures and preservation of movement. Figure 1 shows that the N2C model increases the overall intensities of the scans, a trend corroborated in Fig. 3D, while the N2N model enhances the contrast of different structures. Figure 4 shows that BM3D and NLM strategies, while obtaining good results in terms of region-specific metrics (Fig. 3), are not reliable regarding the accuracy of movement quantification. Whether this phenomenon comes from induced artefacts in the denoised images needs to be further investigated.
Overall, the one-cycle denoising presents several advantages compared to the other alternatives. First, our method is simple and effective in the context of dynamic OCT acquisition. It is easy to implement and requires only the addition of a pulse sensor connected to the subject’s finger during the video acquisition. The computational workload is quite low, and the code needed to synchronize the pulse signal and OCT timestamps can be adapted to clinical OCT equipment. Most importantly, the rationale for the algorithm is rooted in basic physiology and hence clear and understandable. The result follows a well-known method, and the output can be easily comprehended, and even debugged, for clinical decisions.
Denoising methods requiring pairs of images, such as our one-cycle method and both neural networks approaches, depend heavily on the accuracy of the eye tracking system of our OCT machine. This system’s precision is an ongoing challenge in both commercial and research available OCT apparatus. Averaging images corresponding to different ocular structures, or trying to transform one ocular plane into another, would result in overall bad behavior of those three methods. Those same approaches depend on the accuracy of the image registration as well. The global registration removes large changes due to breathing, head and eye movement, while the finer A-scan registration corrects tilt motion artefacts that occur during a B-scan acquisition and fixation issues. These tilting artefacts are typically not corrected by common rigid registration techniques, nor by the hardware-based motion artefact correction modules of commercial OCT machines. For videos spanning long periods (30 s in our case), A-scan tilts can accumulate over time and severely hinder the registration and matching of structures between pairs of images. Left uncorrected, they lead to blurring artefacts in ocular structures when averaging several images; they can also bias the reconstruction of denoised scans in CNN approaches. The cross-correlation A-scan registration technique we use is an efficient and simple method to correct these tilts.
It is important to note that what renders the one-cycle method efficient can also be considered a shortcoming: as it assumes temporal redundancy between cardiac cycles, any non-periodic information in the video will be lost. An approach not relying on averaging between cycles could, in principle, detect abnormalities manifesting a non-periodic behavior. Alternatively, averaging to several cycles rather than a single one is possible, but the quality of the denoised images would be affected. In order to reach the same level of image enhancement as one-cycle, the OCT acquisition time would need to be increased proportionally. This could lead to difficulty in registering the images, for example due to increased subject movement caused by fatigue.
In comparison, the N2N workflow can train a network in an unsupervised way that still gives high quality results. Once trained, the model only needs the current image to output a denoised version, and temporal information from multiple heart cycles is preserved. However, the reliability of the denoised scans at the pixel level is not assured, even if in the present case, only simple convolution and pooling operations are used. In order to use the AI-denoised images for biomarker quantification (pixel-wise), more work would need to be done in understanding how the network’s operations affect its reliability. This is true for BM3D and NLM as well, being single scans denoising methods which obtain high image quality metrics, but where the trustworthiness of the pixel intensities is not guaranteed. In contrast, the one-cycle method establishes a reliability that is lacking in these other automated approaches.
Limitations of the study
The Noise2Noise (and Noise2Clean) workflows were used in this work with a simple U-net architecture. This training scheme and architecture could potentially be improved, as has been tried in the past (e.g., GANs or fully connected convolutional networks) in the OCT denoising field. A more complete comparison between image-processing-based and neural network denoising methods could be carried out with more complex models as well (e.g., vision transformers). However, these architectures come with increased complexity of data acquisition and training. Moreover, the reliability of denoised pixel intensities would need to be extensively assessed, especially in the case of generative methods.
Conclusion
The proposed one-cycle workflow is a simple and effective denoising method to facilitate analysis of dynamic OCT data. The temporal redundancy of single-scan video acquisitions allows to average scans without the risk of blurring small movements needed for dynamic analysis of the tissues, as is risky with classical averaging techniques. Compared to traditional denoising strategies such as BM3D and NLM in addition to convolutional neural network (CNN) architectures in both unsupervised and supervised training schemes, our one-cycle method achieves the highest denoising performance, increases image quality globally and in specific areas of the eye, and preserves physiological motion before and after denoising. In comparison, CNNs, particularly in unsupervised training, achieve remarkable performance but blur the images. Hence, they achieve high denoising performance in terms of CNR and MMI, but at the cost of losing high-resolution details of ocular structures. BM3D and NLM strategies are easy to use and quantitatively improve image quality, but do not preserve induced deformation. While the one-cycle method benefits from multiple input images, all other denoising techniques are single-scan methods and thus achieve impressive performance.
Methods
One-cardiac-cycle OCT videos
We present a denoising strategy based on the assumption that ocular tissue images in a temporal sequence will repeat periodically with the heart cycle. Images from the same cardiac phase can thus be averaged without blurring changes due to movement or physiological processes. Rigid registrations applied to the B-scans in a video are sufficient to correct global movement due to eye saccades, lack of gaze fixation and breathing. The resulting video captures changes due only to the blood flow in and out of tissues. As such, the registered frames corresponding to the same instant in a cardiac cycle, i.e., all frames taken at equal delays with respect to systole, should look the same. Averaging only those specific images reduces speckle and other noise sources present in the images while preserving visible pulsation-driven movements.
We acquire OCT videos at a B-scan rate of 100 Hz for a duration of approximately 30 s (see section “Data acquisition”). These videos image a single plane of the retina and optic nerve head throughout multiple heart cycles. The heart cycle is simultaneously measured with a pulse sensor and digitized. The peaks of the cardiac signal are detected, and the period is divided into 100 intervals, referred hereafter as bins. Images are assigned to bins using the timestamps that indicate the moment of acquisition of every frame of the OCT movie. The 3000 frames of our movies are thereby split in 100 bins; each bin receives approximately 30 frames representing identical instants of different cycles. For every bin, the registration steps described in section “Image registration” are applied, using the first frame of that bin as the reference frame. After registration, all images in every bin are averaged together, yielding a single denoised image. To align the averaged frames together and recreate a spatially coherent movie, a second registration is applied, using the same workflow as in section “Image registration”. This transforms a 30-s-long OCT video of 3000 frames into a one-cycle, approximately 1-s-long video, where every frame is the average of approximately 30 images. See Fig. 5 for a visual representation of this workflow.
Data acquisition
Time series are obtained using the Zeiss PlexElite 9000, a swept-source OCT machine. We use a custom mode to acquire 30-s-long videos at a B-scan frame rate of approximately 100 Hz and an A-scan rate of approximately 100 kHz. An eye tracker ensures that the same region of the eye is being imaged throughout the whole 30 s. This set-up produces 3000-frames B-scan movies mapping a single plane of the optic nerve head (ONH) in less than a minute. The output images have a size of 1536 × 1024 pixels, with pixel size of 1.95 µm axially and approximately 5.9 µm transversally. The OCT resolution is of 6.3 µm axially and 20 µm transversally. A more detailed description of the data acquisition protocol was previously published12.
The subject’s heart beats are measured simultaneously with the OCT video using a pulse sensor (PulseSensor, World Famous Electronics llc.) in contact with the subject’s finger. This sensor shines light (λ = 515 ± 15 nm) on the finger and a photodiode measures the backscattering intensity, which is proportional to oxygenated hemoglobin. Signal digitization is achieved with an analog-to-digital-converter (ADS1263, Waveshare) connected to a Raspberry Pi (Model 4B). This set-up was first described elsewhere39. 21 videos of different eyes were analyzed in total, corresponding to 15 different subjects (some were imaged in both eyes).
Image registration
We use a two-step registration workflow: a global translation using an arbitrary reference frame (described in section “One-cardiac-cycle OCT videos”), followed by a fine A-scan alignment. The latter step serves to correct a progressive rotational misalignment (tilt) that can affect the individual A-scans. We correct the A-scan tilt by calculating the correlation between pairs of A-scans from the reference and moving images. The maximum correlation between pairs of A-scans is the axial shift that could be applied to the moving A-scan to optimally register it to the reference image. This shift is calculated for all pairs of A-scans and a linear fit is obtained. Axial shifts are applied based on the linear fits computed for every image. In the interest of speed, correlations are only computed every 20 A-scans.
An outlier detection scheme is used to detect frames where the registration process would not have worked. The Mattes Mutual Information (MMI)40 metric is computed between all frames corresponding to the same cardiac phase, where the first frame of each bin is taken as the reference. This similarity metric allows to detect images where the registration would have diverged, resulting in artifacts. Otsu thresholding is used to split the whole set of MMIs computed on every frame of the movie, after which a binomial distribution is fitted on the split. Goodness of fit indicates whether frames can be detected as outliers as such the lower MMIs corresponding frames are removed.
Global image quality quantification
Different metrics are used to quantify image quality before and after denoising has been applied.
We use Mattes Mutual Information (MMI)40 to compute the similarity between whole image pairs before and after denoising:
Mattes Mutual Information:
where P represents the probability distribution of pixels in an image, and x and y are pixel intensity values of images X and Y.
We use the signal-to-noise ratio (SNR) (Eq. (2)) and the contrast-to-noise ratio (CNR) (Eq. (3)) to quantify the noise present in structures of interest:
Signal-to-noise Ratio:
Contrast-to-noise Ratio:
where \(\mu\) and \(\sigma\) are the mean and the standard deviation of the intensity, respectively, and index i refers to a region of interest (ROI) and b to the background. SNR and CNR are calculated in manually segmented ROIs corresponding to the choroid, retinal nerve fiber layer (RNFL) and retinal pigment epithelium (RPE). ROIs were identified on one B-scan for every video by the same observer. The ROI corresponding to the background was located in the bottom portion of the B-scan on both sides of the ONH.
We use the 2-dimensional Fast Fourier Transform (FFT) to compute the signal content of the images in the frequency domain. Based on the specifications that determine axial and transverse resolutions of the OCT machine, we define an ellipse that contains useful information. We calculated the semi-axes of this ellipse at the cut-off frequencies corresponding to the axial and transverse resolution of the OCT machine. Signal outside this ellipse necessarily corresponds to noise, thus the average spectral intensity in that region measures the overall noise level. See Fig. 2D for a visual representation of the FFT spectrum and the ellipse.
Region-specific image quality quantification
To allow more meaningful comparisons between different denoising strategies, we designed several metrics to describe the image clarity of different ocular structures. ROI-specific metrics are defined and computed for the sclera, vitreous humor and retinal layers. All ROIs are manually segmented on one frame in each video by the same observer.
At the resolution provided by an OCT, the sclera is a homogeneous collagenous layer. Thus, variation of intensities in this region should be minimal; the standard deviation is computed and expected to be close to zero.
As the OCT signal in the vitreous humor should be null, the mean and the standard deviation of the pixel intensities are computed in this region, and both are expected to be close to zero.
The different retinal layers are expected to be clearly defined and distinguishable. Starting from the inner limiting membrane and ending with the Bruch’s membrane, we count 11 interfaces that should be found in denoised OCT scans. These interfaces include the transitions between the retinal nerve fiber layer, ganglion cell layer, inner plexiform layer, inner nuclear layer, outer plexiform layer, outer nuclear layer, external limiting membrane (very thin layer that would be counted as one delimitation), photoreceptor inner and outer segments (that are visibly divided into three sub-layers) and finally the retinal pigment epithelium. The thicknesses of these layers vary along the retina and depend on the exact position of the scan, as well as on inter-subject variability. We considered as optimal an average thickness of approximately 50 µm41. To measure this, we extract the locations and the distances between peaks of the vertical gradient across the retina, from the RNFL to the RPE (Fig. 3F). An average of the number of detected layers and their average thickness was computed over all lines extracted in the ROI.
Movement preservation quantification
A good denoising strategy removes noise while preserving tissue anatomy. Furthermore, using temporal data allows the analysis of the dynamics and biomechanical properties of tissues. As such, we designed a metric assessing whether physiological movement induced in scans was still visible after denoising.
We followed a methodology we have used in the past12: a known displacement field simulating the expansion of the choroid was created. 100 pairs of consecutive scans in the original and the one-cycle videos were identified, after which the second scan of each pair was warped with the known deformation field (for the one-cycle videos, those hundred scans were simply the one-cycle video itself). All pairs of scans were denoised using their respective denoising strategies, after which the displacement field was computed again between both denoised scans with the demons algorithm42,43. The average absolute median error was computed between the known and computed displacement fields in the retina.
We also quantified the displacement occurring between pairs of consecutive frames after denoising (without inducing artificial deformation). The average median displacement was computed in the retina over 100 pairs of images for each denoising strategy.
BM3D and NLM denoising approaches
The Non-Local Means (NLM) approach denoises images based on averaging neighborhood pixels based on their similarity. The similarity is computed using the Euclidean distance between the pixel values. We used the MATLAB implementation of the algorithm provided by the authors38.
The Block-matching and 3D filtering (BM3D) algorithm37 was originally inspired by the NLM approach, and is based on the Wiener filtering of similar groupings of pixels called blocks. The MATLAB implementation of the algorithm published by the authors was used.
Neural networks implementation
We implemented supervised and unsupervised training schemes for the Noise2Noise architecture. The model architecture, training sets, hyperparameters and metrics used are described in the following subsections.
Noise2Noise
The Noise2Noise (N2N) neural network framework denoises images in an unsupervised manner. The method is based on training the network using pairs of images representing the same structures with different noise levels. In the original article25, synthetic noise was added to the images to simulate these pairs.
Using the same assumption as for the one-cycle videos, we suppose that registered frames corresponding to the same instant in the cardiac cycle represent the same structures but with different noise distributions. Hence, frames detected to be in the same temporal bin are considered as pairs and used as input and output of the N2N framework (registration is described in section “Image registration”). All combinations of these pairs of frames are used as the N2N training set. Images are cropped to facilitate training to a fixed size of 512 by 512. Cropped patches are located randomly in the images, 20 patches are extracted from each image to introduce variation to the training set, and a total of 16,812,180 patches are used. We use a U-net-like architecture as the network backbone44. The convolutional blocks are composed of two convolutional layers as well as a max pooling (resp. upsampling) layer on the encoder (resp. decoder) pathway. Skip connections are used, as in the original U-net. No drop-out is used. The ReLU activation is used whenever applicable. We choose the Adam optimizer with \({\beta }_{1}=0.9, {\beta }_{2}=0.99, \varepsilon ={10}^{-8}\).
Noise2Clean implementation
As in other works that tested N2N, we compared the performance of the unsupervised framework to a supervised counterpart, which we named Noise2Clean (N2C). The N2C model architecture and gradient descent optimizer are the same as in the N2N workflow, the difference lying in the training dataset. In N2C, the network is trained to transform a noisy B-scan into its corresponding one-cycle averaged frame. As such, N2C corresponds to a supervised training scheme where the inputs are the noisy frames and the outputs are the corresponding denoised frames, using the one-cycle workflow to provide the ground truth samples. All combinations of these pairs of frames are used to compose the N2C training set. The training set comprises a total of 1,196,480 patches.
Training
We use a batch size of 4 cropped images. The total image sets are split into training, validation, and testing, with proportions 64%, 16% and 20%, respectively. We choose a grid search approach to vary several parameters such as the number of convolutional blocks used in the U-net-like architecture, the number of features present at each convolutional layer, the starting learning rate and the loss function used. Monitoring of the training uses the weights and biases library. The model architectures and training were implemented using the Keras library in Python. A fixed number of 300 epochs is used for training, with early stopping to end the training when the validation loss stops improving. The same training sets are used for all combinations in the grid search.
Four different loss functions are tested: mean squared error (MSE), mean absolute error (MAE), cosine similarity function and L0 error function, as defined elsewhere25. We use the peak signal-to-noise ratio (PSNR) and Structural Similarity Index Metric (SSIM)45 (Eqs. (4) and (5)) to evaluate the performance of the models. Those that achieve the highest validation SSIM and PSNR while limiting the presence of artefacts are chosen as best.
Peak signal-to-noise ratio (PSNR):
Structural Similarity Index Metric (SSIM):
The final N2N model consists of a U-net architecture made up of 6 convolutional blocks in each pathway, using a starting number of 64 filters per convolutional layer, and training with an initial learning rate of 10–5. The final N2C model consists of the same optimal hyperparameters, apart from the starting learning rate which is of 10–4.
Ethical compliance
Human subjects were included in this study. This study was approved by the institutional review board of Maisonneuve-Rosemont Hospital and was performed in accordance with the 1964 Declaration of Helsinki and its amendments. Written informed consent was obtained from all participants. No animal subjects were used in this study.
Data availability
The datasets used and analyzed during the current study will be available from the corresponding author on reasonable request.
Code availability
The MATLAB code used to generate the one-cycle videos is available at https://github.com/EmmanuelleRicher/One-cycle-denoising.
References
Bowd, C., Weinreb, R. N., Williams, J. M. & Zangwill, L. M. The retinal nerve fiber layer thickness in ocular hypertensive, normal, and glaucomatous eyes with optical coherence tomography. Arch. Ophthalmol. 118, 22–26 (2000).
Groth, S. L. & Joos, K. M. Primary open-angle glaucoma. In Albert and Jakobiec’s Principles and Practice of Ophthalmology (eds Albert, D. et al.) 1–15 (Springer International Publishing, 2020).
Kass, M. A. et al. The Ocular Hypertension Treatment Study: A randomized trial determines that topical ocular hypotensive medication delays or prevents the onset of primary open-angle glaucoma. Arch. Ophthalmol. 120, 701–713 (2002).
Wang, X. et al. Finite element analysis predicts large optic nerve head strains during horizontal eye movements. Invest. Ophthalmol. Vis. Sci. 57, 2452–2462 (2016).
Sigal, I. A., Flanagan, J. G., Tertinegg, I. & Ethier, C. R. Finite element modeling of optic nerve head biomechanics. Invest. Ophthalmol. Vis. Sci. 45, 4378–4387 (2004).
Sigal, I. A., Yang, H., Roberts, M. D., Burgoyne, C. F. & Downs, J. C. IOP-induced lamina cribrosa displacement and scleral canal expansion: An analysis of factor interactions using parameterized eye-specific models. Invest. Ophthalmol. Vis. Sci. 52, 1896–1907 (2011).
Zhong, F. et al. A high-accuracy and high-efficiency digital volume correlation method to characterize in-vivo optic nerve head biomechanics from optical coherence tomography. Acta Biomater. 143, 72–86 (2022).
Yang, H. et al. Deformation of the early glaucomatous monkey optic nerve head connective tissue after acute IOP elevation in 3-D histomorphometric reconstructions. Invest. Ophthalmol. Vis. Sci. 52, 345–363 (2011).
Wei, J. et al. Comparing acute IOP-induced lamina cribrosa deformations premortem and postmortem. Transl. Vis. Sci. Technol. 11, 1 (2022).
Chuangsuwanich, T. et al. Differing associations between optic nerve head strains and visual field loss in patients with normal- and high-tension glaucoma. Ophthalmology 130, 99–110 (2023).
Girard, M. J. A. et al. In Vivo 3-dimensional strain mapping of the optic nerve head following intraocular pressure lowering by trabeculectomy. Ophthalmology 123, 1190–1200 (2016).
Solano, M. M., Richer, E., Cheriet, F., Lesk, M. R. & Costantino, S. Mapping pulsatile optic nerve head deformation using OCT. Ophthalmol. Sci. 2, 100205 (2022).
Flammer, J. et al. The eye and the heart. Eur. Heart J. 34, 1270–1278 (2013).
Mozaffarieh, M., Grieshaber, M. C. & Flammer, J. Oxygen and blood flow: Players in the pathogenesis of glaucoma. Mol. Vis. 14, 224–233 (2008).
McClintic, B. R., McClintic, J. I., Bisognano, J. D. & Block, R. C. The relationship between retinal microvascular abnormalities and coronary heart disease: A review. Am. J. Med. 123(374), e1-7 (2010).
Klein, R. et al. Are retinal arteriolar abnormalities related to atherosclerosis? The Atherosclerosis Risk in Communities Study. Arterioscler. Thromb. Vasc. Biol. 20, 1644–1650 (2000).
Klein, R. et al. Fifteen-year cumulative incidence of age-related macular degeneration: The Beaver Dam Eye Study. Ophthalmology 114, 253–262 (2007).
Yu, T., Mitchell, P., Berry, G., Li, W. & Wang, J. J. Retinopathy in older persons without diabetes and its relationship to hypertension. Arch. Ophthalmol. Chic. Ill 1960(116), 83–89 (1998).
de Carlo, T. E., Romano, A., Waheed, N. K. & Duker, J. S. A review of optical coherence tomography angiography (OCTA). Int. J. Retina Vitr. 1, 5 (2015).
Leitgeb, R. A., Werkmeister, R. M., Blatter, C. & Schmetterer, L. Doppler optical coherence tomography. Prog. Retin. Eye Res. 41, 26–43 (2014).
Schmoll, T. & Leitgeb, R. A. Heart-beat-phase-coherent Doppler optical coherence tomography for measuring pulsatile ocular blood flow. J. Biophotonics 6, 275–282 (2013).
Lee, B. et al. Cardiac-gated en face doppler measurement of retinal blood flow using swept-source optical coherence tomography at 100,000 axial scans per second. Invest. Ophthalmol. Vis. Sci. 56, 2522–2530 (2015).
Jin, Y. et al. Effect of changing heart rate on the ocular pulse and dynamic biomechanical behavior of the optic nerve head. Invest. Ophthalmol. Vis. Sci. 61, 27–27 (2020).
Sakamoto, A., Hangai, M. & Yoshimura, N. Spectral-domain optical coherence tomography with multiple B-Scan averaging for enhanced imaging of retinal diseases. Ophthalmology 115, 1071-1078.e7 (2008).
Lehtinen, J. et al. Noise2Noise: Learning image restoration without clean data. ArXiv Prepr. ArXiv180304189 (2018).
Qiu, B. et al. Comparative study of deep neural networks with unsupervised Noise2Noise strategy for noise reduction of optical coherence tomography images. J. Biophotonics 14, e202100151 (2021).
Krull, A., Buchholz, T.-O. & Jug, F. Noise2void-Learning Denoising from Single Noisy Images 2129–2137 (2019).
Nienhaus, J. et al. Live 4D-OCT denoising with self-supervised deep learning. Sci. Rep. 13, 5760 (2023).
Ma, Y. et al. Speckle noise reduction in optical coherence tomography images based on edge-sensitive cGAN. Biomed. Opt. Express 9, 5129–5146 (2018).
Devalla, S. K. et al. A deep learning approach to denoise optical coherence tomography images of the optic nerve head. Sci. Rep. 9, 14454 (2019).
Qiu, B. et al. Noise reduction in optical coherence tomography images using a deep neural network with perceptually-sensitive loss function. Biomed. Opt. Express 11, 817–830 (2020).
Holzinger, A., Langs, G., Denk, H., Zatloukal, K. & Müller, H. Causability and explainability of artificial intelligence in medicine. WIREs Data Min. Knowl. Discov. 9, e1312 (2019).
Holzinger, A., Biemann, C., Pattichis, C. S. & Kell, D. B. What do we need to build explainable AI systems for the medical domain? ArXiv Prepr. ArXiv171209923 (2017).
Ghassemi, M., Oakden-Rayner, L. & Beam, A. L. The false hope of current approaches to explainable artificial intelligence in health care. Lancet Digit. Health 3, e745–e750 (2021).
Begoli, E., Bhattacharya, T. & Kusnezov, D. The need for uncertainty quantification in machine-assisted medical decision making. Nat. Mach. Intell. 1, 20–23 (2019).
Huang, S., Tang, C., Xu, M., Qiu, Y. & Lei, Z. BM3D-based total variation algorithm for speckle removal with structure-preserving in OCT images. Appl. Opt. 58, 6233–6243 (2019).
Dabov, K., Foi, A., Katkovnik, V. & Egiazarian, K. Image denoising by sparse 3-D transform-domain collaborative filtering. IEEE Trans. Image Process. 16, 2080–2095 (2007).
A. Buades, B. Coll, & J.-M. Morel. A non-local algorithm for image denoising. in 2005 IEEE Computer Society Conference on Computer Vision and Pattern Recognition (CVPR’05) vol. 2 60–65 (2005).
Richer, E., Solano, M. M., Cheriet, F., Lesk, M. R. & Costantino, S. Noise-free one-cardiac-cycle OCT videos for local assessment of retinal tissue deformation. In International Symposium on Biomedical Imaging (ISBI) (IEEE Xplore, 2023).
Mattes, D., Haynor, D. R., Vesselle, H., Lewellyn, T. K. & Eubank, W. Nonrigid multimodality image registration. SPIE 4322, 1609–1620 (2001).
Bagci, A. M. et al. Thickness profiles of retinal layers by optical coherence tomography image segmentation. Am. J. Ophthalmol. 146, 679-687.e1 (2008).
Thirion, J.-P. Image matching as a diffusion process: An analogy with Maxwell’s demons. Med. Image Anal. 2, 243–260 (1998).
Vercauteren, T., Pennec, X., Perchant, A. & Ayache, N. Diffeomorphic demons: Efficient non-parametric image registration. Math. Brain Imaging 45, S61–S72 (2009).
Ronneberger, O., Fischer, P. & Brox, T. U-net: Convolutional networks for biomedical image segmentation. In Medical Image Computing and Computer-Assisted Intervention–MICCAI 2015: 18th International Conference, Munich, Germany, October 5–9, 2015, Proceedings, Part III 18 234–241 (Springer, 2015).
Z. Wang, E. P. Simoncelli, & A. C. Bovik. Multiscale structural similarity for image quality assessment. In The Thrity-Seventh Asilomar Conference on Signals, Systems & Computers, 2003 vol. 2 1398–1402 (2003).
Bechtold, B. Violin Plots for Matlab (2016).
Acknowledgements
Special thanks to Zacharie Legault, Gabriel Lepetit-Aimon and Charles M. Bélanger Nzakimuena for their conceptual and technical help in preparing the figures. We thank Philippe Debanné for manuscript editing. The violin plots were generated using software publicly available46.
Funding
This work was supported by grants from the Canadian Institutes of Health Research (S.C and M.R.L), the Canadian Space Agency (S.C and M.R.L), the Fonds de Recherche du Québec en Santé (M.M.S), and the Glaucoma Research Society of Canada (M.R.L). M.M.S received a Salary Award from the Fonds de Recherche du Québec en Santé. E.R. received bursaries from the Institut de Valorisation des données (IVADO) and the OPSIDIAN scholarship program of Polytechnique Montréal.
Author information
Authors and Affiliations
Contributions
Conceptualization: S.C and E.R; Methodology: S.C and E.R; Software: E.R; Investigation: E.R, M.R.L and M.M.S; Writing—Original Draft: S.C and E.R; Writing—Review & Editing: F.C, S.C, M.R.L, M.M.S and E.R; Supervision: F.C and S.C; Funding Acquisition: F.C.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Supplementary Video 1.
Supplementary Video 2.
Supplementary Video 3.
Supplementary Video 4.
Supplementary Video 5.
Supplementary Video 6.
Supplementary Video 7.
Supplementary Video 8.
Supplementary Video 9.
Supplementary Video 10.
Supplementary Video 11.
Supplementary Video 12.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Richer, E., Solano, M.M., Cheriet, F. et al. Denoising OCT videos based on temporal redundancy. Sci Rep 14, 6605 (2024). https://doi.org/10.1038/s41598-024-56935-0
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-024-56935-0
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.