Abstract
Chondroitin sulfate (CS) is a family of glycosaminoglycans and have a wide range of applications in dietary supplements and pharmaceutical drugs. In this study, we evaluated the effects of several types of CS, differing in their sulfated positions, on the human colonic microbiota and their metabolites. CS (CSA, CSC, and CSE) and non-sulfated chondroitin (CH) were added into an in vitro human colonic microbiota model with fecal samples from 10 healthy individuals. CS addition showed a tendency to increase the relative abundance of Bacteroides, Eubacterium, and Faecalibacterium, and CSC and CSE addition significantly increased the total number of eubacteria in the culture of the Kobe University Human Intestinal Microbiota Model. CSE addition also resulted in a significant increase in short-chain fatty acid (SCFA) levels. Furthermore, addition with CSC and CSE increased the levels of a wide range of metabolites including lysine, ornithine, and Ile-Pro-Pro, which could have beneficial effects on the host. However, significant increases in the total number of eubacteria, relative abundance of Bacteroides, and SCFA levels were also observed after addition with CH, and the trends in the effects of CH addition on metabolite concentrations were identical to those of CSC and CSE addition. These results provide novel insight into the contribution of the colonic microbiota to the beneficial effects of dietary CS.
Similar content being viewed by others
Introduction
Glycosaminoglycans (GAGs) are linear polysaccharides comprising repeating disaccharide units of amino sugars (N-acetylglucosamine [GlcNAc] or N-acetylgalactosamine [GalNAc]) and either hexuronic acid or hexose that are universally present in all animals, including humans, as major components of the extracellular matrix. GAGs can be classified into several types according to their disaccharide units and the linkages between sugars. Chondroitin sulfate (CS) is a representative family of GAGs that are ubiquitously present on cell surfaces and within extracellular matrices. CS comprises disaccharide units alternating β (1–4)-linked GalNAc and β (1–3)-linked glucuronic acid (GlcUA) bearing sulfate groups at various positions and has been identified based on its characteristic disaccharide composition (Supplementary Fig. S1). CSA is predominantly composed of an A-unit in which the C-4 position of GalNAc is sulfated. CSC mainly comprises a C-unit with a sulfate group at the C-6 position of GalNAc. CSE has a composition rich in E-units in which the C-4 and C-6 positions of GalNAc are sulfated. Non-sulfated chondroitin (CH) comprises only a non-sulfated unit (O-unit).
CS is particularly known as a major component of articular cartilage and has been implicated in chondrocyte proliferation and differentiation1. It has demonstrated therapeutic immunomodulatory and anti-inflammatory effects2 and is used as a dietary supplement and pharmaceutical drug for the treatment of joint diseases, including osteoarthritis3. Moreover, CS has been recently reported involved in the regulation of various physiological events, such as organogenesis, cytokinesis, morphogenesis, and central nervous system development4, 5. Several studies have shown that oral administration of CS reduces the incidence of coronary events in patients with coronary heart disease6,7,8, and Melgar-Lesmes et al.9 suggested that these cardioprotective effects of CS may arise from modulation of proinflammatory activation of endothelium and monocytes and foam cell formation.
Despite its broad physiological roles and wide range of applications, the mechanisms underlying the beneficial effects of dietary CS remain unclear. CS is a high-molecular-weight polysaccharide that is poorly absorbed by the body when administered orally10, 11. However, some intestinal bacteria can degrade and utilize CS for their growth12, 13. Shang et al.14 reported that dietary CS and its derivatives altered the composition of colonic bacteria in mice. Since the gut microbiota exerts a wide effect on host physiology, the beneficial effects of oral administration of CS on the host are considered to be due to the modulation of colonic microbiota and their metabolites. However, ethical considerations have constrained human interventional clinical trials, and the functionality of orally administered CS in humans remains unclear. In particular, the effects of CS sulfation patterns on the colonic microbiota and their metabolites are largely unknown.
In this study, we investigated the possible effects of various CSs, including CH, on human colonic microbiota using an in vitro human colonic microbiota model, the Kobe University Human Intestinal Microbiota Model (KUHIMM)15, 16. KUHIMM is a single-batch anaerobic fermentation system for the metagenomic and metabolic simulation of human colonic microbiota. Fecal samples from 10 healthy individuals were individually cultivated in KUHIMM with or without each CS, and the total eubacterial growth, microbial composition, and metabolites after cultivation were analyzed to evaluate the effects of these CSs on human colonic microbiota.
Results
Consumption of CSs by human colonic microbiota in KUHIMM
Fecal samples from 10 Japanese volunteers were individually cultivated anaerobically in KUHIMM with each CS (CH, CSA, CSC, and CSE) at 37 °C for 48 h. The concentrations of each type of CS in the culture broth at 0 and 48 h are shown in Supplementary Table S1, and the residual ratio of CSs after 48 h of incubation are summarized in Table 1. In most cultivation experiments, the residual ratio of CSs was low (< 20%), indicating that most of the added CSs were degraded or absorbed by the human colonic microbiota simulated by the KUHIMM and consumed. In contrast, no remarkable consumption of CSA was observed in the cultivation with fecal sample HS-02 (Supplementary Table S1), suggesting failure of cultivation. Therefore, the culture sample of the CSA/HS-02 combination was excluded from further analysis.
Effects of CS addition on structure of human colonic microbiota in KUHIMM
The microbial composition of original fecal samples and that of samples that cultivated in KUHIMM with or without each CS for 48 h were analyzed by sequencing, covering the V3–V4 hypervariable regions of the bacterial 16S rRNA gene. KUHIMM without CS addition was used as the control. An average of 128,649 high-quality leads was obtained for each sample (Supplementary Table S2). The bacterial operational taxonomic unit numbers and Chao1 value for species richness were lower in the CS non-added KUHIMM culture than in the original fecal samples (p = 0.029 and 0.052, for operational taxonomic unit numbers and Chao1, respectively; Mann–Whitney U-test, Supplementary Table S2). However, there was no significant difference in the count of operational taxonomic unit between the CS-added and non-added KUHIMM cultures (CH: p = 0.388; CSA: p = 0.127; CSC: p = 0.325; CSE: p = 0.211; Mann–Whitney U-test, Supplementary Table S2). The Shannon index for species diversity was lower in the CS non-added KUHIMM culture group than in the original fecal sample group (p = 0.002; Mann–Whitney U-test, Supplementary Table S2). However, there was no significant difference in the values between the CS-added and non-added KUHIMM cultures (CH: p = 0.905; CSA: p = 0.651; CSC: p = 0.912; CSE: p = 0.604; Mann–Whitney U-test, Supplementary Table S2). The Simpson index for species diversity was lower in the CS non-added KUHIMM culture group than in the original fecal sample group (p = 0.003; Mann–Whitney U-test, Supplementary Table S2); however, no significant difference was found in the values between the CS-added and non-added KUHIMM cultures (CH: p = 0.968; CSA: p = 0.739; CSC: p > 0.999; CSE: p = 0.661; Mann–Whitney U-test, Supplementary Table S2). These results confirmed that the diversity of the colonic microbiota did not change upon the addition of 0.3% CSs in KUHIMM. On the other hand, CH, CSC, and CSE addition significantly increased the number of total eubacteria in KUHIMM compared to the control culture (CH: p = 0.014; CSC: p = 0.002; CSE: p = 0.037; Wilcoxon matched-pairs signed-rank test, Fig. 1).
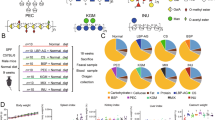
Figure 2a presents the relative abundance of the microbiota of each sample at the genus level. The relative abundance of Bacteroides in the KUHIMM cultures tended to increase with CS addition (Fig. 2a) and was significantly increased by CH, CSA, and CSE addition in comparison to the control culture (CH: p = 0.010; CSA: p = 0.039; CSE: p = 0.027; Wilcoxon matched-pairs signed-rank test, Fig. 2b). The relative abundances of Eubacterium (CH: p = 0.014; CSA: p = 0.012; Wilcoxon matched-pairs signed-rank test, Fig. 2c) and Faecalibacterium (CH: p = 0.039; CSA: p = 0.047; Wilcoxon matched-pairs signed-rank test, Fig. 2d), which belong to the phylum Firmicutes, were significantly increased by CH and CSA addition. Changes in the relative abundances of these genera in the KUHIMM inoculated with each fecal sample are shown in Supplementary Fig. S2. For the other genera, no significant increases in the relative abundance were observed between the CS-added and non-added KUHIMM cultures.
Effect of 0.3% CS addition on the relative abundance of the microbiota of each sample. (a) Genus-level compositional view of bacteria in original feces (FEC) and KUHIMM after 48 h of cultivation with each CS (CH, CSA, CSC, or CSE) or without CS (CS(–)). Data are shown as the average relative abundances in 10 samples (9 for CSA only). Genera with lower abundance (< 1.0%) and lower levels of similarity (< 99%) were indicated as Others and Unclassified bacterium, respectively. (b–d) The relative abundance of Bacteroides (b), Eubacterium (c), and Faecalibacterium (d) in KUHIMM. Data are shown as the median and interquartile range (25th–75th percentiles) of 10 samples (9 for CSA only). *p < 0.05, **p < 0.01. Wilcoxon matched-pairs signed-rank test. (e) The relative abundance of Bacteroides species in KUHIMM. Data are shown as the average relative abundances in 10 samples (9 for CSA only).
While CS addition increased the relative abundance of Bacteroides at the genus level, the breakdown of this increase at the species level differed depending on the CS addition (Fig. 2e). The relative abundance of Bacteroides ovatus and Bacteroides caccae increased after addition with any type of CS, whereas that of Bacteroides plebeius increased only after addition with CSA, CSC, and CSE, but not CH.
Effect of CS addition on short-chain fatty acid (SCFA) production
To evaluate the effect of CSs on the production of SCFAs, which play an essential role in maintaining human health, the levels of three typical SCFAs, namely acetate, propionate, and butyrate, were measured after 48 h of cultivation in KUHIMM with or without CS (Fig. 3). The acetate level was significantly increased by CH addition (p = 0.009; Wilcoxon matched-pairs signed-rank test, Fig. 3a). Meanwhile, the propionate level was significantly increased by CH and CSE addition (p = 0.020 and p = 0.009, respectively; Wilcoxon matched-pairs signed-rank test, Fig. 3b). However, butyrate levels were not significantly altered by addition with any type of CS (Fig. 3c). Changes in the levels of SCFAs in the KUHIMM inoculated with each fecal sample are shown in Supplementary Fig. S2.
Comprehensive quantification of metabolites
To further investigate the effect of CH, CSC, and CSE on bacterial metabolism in the human colon, metabolites in the supernatant of KUHIMM cultures were comprehensively quantified. After 48 h of cultivation in KUHIMM, 348 metabolites were detected in the supernatant, of which 75 showed significant concentration changes after CS addition (Supplementary Table S3). Table 2 lists all metabolites for which a significant > twofold change in concentration was observed after each CS addition. Most of these metabolites increased in concentration after CS addition. Among them, the concentrations of the basic amino acids lysine and ornithine were particularly increased by addition with CH (12.7- and 3.8-fold, respectively, compared to non-addition) and CSE (8.8- and 7.1-fold, respectively, compared to non-addition). However, only 3-(4-hydroxyphenyl)propionic acid, 2-(4-hydroxyphenyl)propionic acid, or 3-(3-hydroxyphenyl)propionic acid significantly decreased in concentration by less than half after CS addition.
Discussion
CS have a wide range of applications in dietary supplements and pharmaceutical drugs, while the mechanisms underlying the beneficial effects of orally administered CS remain unclear. In the present study, we investigated the possible effect of various CSs on human colonic microbiota using the in vitro human colonic microbiota model KUHIMM. The 0.3% CS added into KUHIMM together with the human fecal suspension was mostly consumed after 48 h of cultivation. KUHIMM cultures added with CS showed a substantial increase in the number of total eubacteria compared with those without CS addition, and the increases in cultures added with CH, CSC, and CSE were statistically significant (Fig. 1). These results indicate that these CSs are degraded or absorbed by the human colonic microbiota and utilized for growth. However, of note, the average molecular weights of the CSs used in this study differed (Supplementary Table S4), which could also have contributed to the degradation profiles and bacterial growth observed in this study.
Microbial composition analysis at the genus level revealed that the relative abundance of Bacteroides in the KUHIMM cultures tended to increase with CS addition compared with that in the control culture (Fig. 2b). Various Bacteroides strains possess carbohydrate-active enzymes (CAZymes) for GAG degradation and assimilation in their genomes, and are capable of catabolizing CS13. The results obtained in this study further support those of previous studies that Bacteroides strains play a major role in CS catabolism in human colonic microbiota12, 13.
At the species level, the relative abundances of B. ovatus and B. caccae were increased by addition with any type of CS (Fig. 2e). These species can reportedly grow using CS as the sole carbon source17. Meanwhile, B. plebeius, which increased in relative abundance only with CSA, CSC, and CSE addition, has been reported to lack CS assimilation capacity because its gene cluster for CS degradation is not fully functional17. B. plebeius may have grown by utilizing the saccharides produced by other Bacteroides species that can degrade CS, such as B. ovatus and B. caccae. Syntrophic interactions are widespread among bacteria inhabiting the human intestine, where coexisting microorganisms enable other strains to utilize originally inaccessible polysaccharides18. Raghavan et al.17 suggested that B. plebeius may form a cooperative CS-utilization network with other Bacteroides species that can utilize CS. Our results have implications for understanding the cooperative cross-feeding of CS in the human colonic microbiota.
For the other genera, the relative abundances of Eubacterium and Faecalibacterium belonging to the phylum Firmicutes were significantly increased by CH and CSA addition (Fig. 2c,d). Several Firmicutes, including Faecalibacterium prausnitzii19, have been shown to be CS utilizers13. However, to our knowledge, no CS utilization by Eubacterium has been reported. Further research is needed to determine the causal relationship between CH addition and an increase in Eubacterium.
CS-degrading bacteria liberate sulfuric acid during the CS degradation and assimilation processes20. The sulfate released from CS becomes available to sulfate-reducing bacteria and induces their growth21. Shang et al.14 reported that the abundance of sulfate-reducing bacteria Desulfovibrio slightly increased after oral administration of CS and CS oligomer in mice. In the present study, the relative abundance of sulfate-reducing bacteria was very low (< 1% on average) in the fecal samples of 10 healthy individuals, and no significant increase in these bacteria was observed, even in cultures added with CS. Further studies using fecal samples containing relatively high abundances of sulfate-reducing bacteria are needed to elucidate the effects of the presence and position of sulfate groups in CS on their growth and metabolites.
Additionally, we demonstrated the effects of CS addition on the levels of SCFAs (acetate, propionate, and butyrate), which are the main products of saccharolytic fermentation of nondigestible carbohydrates in the colon. CH addition increased acetate and propionate levels, whereas CSE addition increased propionate levels (Fig. 3a,b). In the human colon, bacteria belonging to the phylum Bacteroidetes, including the Bacteroides genus, mainly produce acetate and propionate22. Since the majority of Bacteroides species, including B. ovatus, B. caccae, and B. plebeius, possess the enzymes involved in the production of these SCFAs23, these bacteria may have contributed to the elevated acetate and/or propionate levels. In contrast, butyrate levels were not significantly altered by CS addition (Fig. 3c). The two most dominant butyrate-producing bacteria in the human colon are Eubacterium rectale and F. prausnitzii belonging to the phylum Firmicutes24. Although CH and CSA addition significantly increased the relative abundance of Eubacterium and Faecalibacterium (Fig. 2c,d), they did not contribute to butyrate production. SCFAs generated in the colon play important roles in maintaining intestinal homeostasis, can be absorbed by the host, and exert various beneficial effects through a wide variety of mechanisms25, 26. Acetate and propionate act as natural ligands for several cell-surface G protein-coupled receptors expressed in a wide range of tissues, and they exert various biological regulatory functions, including the suppression of inflammatory responses27, 28, blood pressure regulation29, 30, and appetite suppression31, 32.
To further investigate the effects of CS addition, we comprehensively quantified metabolites in the supernatants of KUHIMM cultures. The trends in the effects of CH, CSC, and CSE addition on metabolite concentrations were consistent (Table 2). Addition with these CSs had a positive effect on the levels of various metabolites, mainly nitrogen-containing compounds, including amino acids and their derivatives, nucleic acid relatives, and peptides. This result suggests that CS, which contains an amino sugar (GalNAc) as a constituent monosaccharide, plays an important role as a nitrogen source as well as a carbon source for colonic bacteria. Some metabolites whose concentrations were markedly increased by CS addition have been reported to have beneficial effects on the host. Lysine is an essential amino acid for humans. It plays an important role in the human body, not only in proteinogenesis, but also in the cross-linking of collagen polypeptides33 and improvement of intestinal calcium absorption34. Ornithine is a nonproteinogenic amino acid produced in the urea cycle (also known as the ornithine cycle) that plays an important role as a hepatoprotective agent and converts excess ammonia to urea. Ornithine has been shown to promote growth hormone secretion, which eventually leads to an antifatigue effect and improves sleep and waking, skin quality, and muscle and bone development35, 36. Furthermore, other urea cycle intermediates, such as arginine and citrulline, also increased in concentration after CS addition (Supplementary Table S3). Oral administration of L-arginine and L-citrulline can effectively reduce blood pressure by enhancing the production of nitric oxide, a well-known vasodilator produced by the vascular endothelium37,38,39. CS addition also had a positive effect on the concentration of the bioactive peptide Ile-Pro-Pro. Ile-Pro-Pro was first identified in Japanese sour milk fermented by Lactobacillus helveticus and Saccharomyces cerevisiae40. It is a weak competitive angiotensin-converting enzyme inhibitor41, and in addition to its well-known blood pressure-lowering effect42, 43, anti-inflammatory and bone-protective activities have also been reported44, 45. These metabolites and SCFAs may be associated with some of the reported beneficial effects of orally administered CS, including anti-inflammatory46 and cardioprotective effects47. However, it should be noted that the in vitro fermentation system used in this study does not account for interactions with the intestinal tract. Whether the metabolites found in this study are physiologically associated with the beneficial effects of dietary CS requires further investigation.
In conclusion, addition with different CSs showed diverse effects on colonic microbiota and its metabolites in KUHIMM. CSA addition increased the relative abundance of Bacteroides, Eubacterium, and Faecalibacterium but did not increase the total number of eubacteria and SCFA levels. In contrast, CSC addition significantly increased the total number of eubacteria but did not significantly affect the genus-level bacterial composition or SCFA levels. Meanwhile, CSE addition resulted in statistically significant increases in the total number of eubacteria, relative abundance of Bacteroides, and SCFA levels. Furthermore, addition with CSC and CSE had a positive effect on the levels of a wide range of metabolites, including amino acids and peptides, which could have beneficial effects on the host. However, significant increases in the total number of eubacteria, relative abundance of Bacteroides, Eubacterium, and Faecalibacterium, and SCFA levels were also observed with addition of CH, which does not contain any sulfate group, and the trends in the effects of CH addition on metabolite concentrations were identical to those of CSC and CSE additions. This suggests that the sulfate groups of CS are not involved in the beneficial effects attributed to the metabolites of the colonic microbiota. Although further studies including human interventional clinical trials are needed, the results obtained in this study provide novel insights into the contribution of the colonic microbiota to the therapeutic effects of dietary CS.
Methods
Characterization of CSs
CSs (CH, CSA, CSC, and CSE) were provided by Seikagaku Co. (Tokyo, Japan). Their characteristics are listed in Supplementary Table S4. The weight-average molecular weight was determined via high-performance liquid chromatography (HPLC) (Shimadzu, Kyoto, Japan). A size exclusion column (Ultrahydrogel linear 7.8 × 300 mm; Waters, Milford, MS, USA) was used together with a refractive index detector (RID-10A; Shimadzu). The HPLC system was operated at 40 °C with 0.2 M NaCl (flow rate, 0.6 mL/min) as the mobile phase. Chromatograms of the size exclusion chromatography of CSs are shown in Supplementary Fig. S3. The weight-average molecular weights were calculated using a standard curve determined with molecular weight-defined pullulan standards (Shodex, Tokyo, Japan) (Supplementary Fig. S4). The disaccharide composition was analyzed using an HPLC apparatus equipped with a post-column fluorescent labelling system as reported previously48 with a slight modification. In brief, each CS was solubilized in water, and chondroitinase ABC (Seikagaku Corp.) and chondroitinase ACII (Seikagaku Corp.) were then added, followed by incubation at 37 °C for 16–18 h to completely digest the CS into their disaccharide units. The reaction solutions were ultrafiltered with a Nanosep centrifuge device with a molecular weight cutoff of 10,000 Da (Pall Corporation, Port Washington, NY, USA). The filtrates were then injected into a C22-bound silica column (SenShu Pak DOCOSIL SP400; Senshu Scientific Co., Tokyo, Japan) and eluted with a gradient of 0–140 mM sodium chloride containing 1.45 mM tetrabutylammonium monohydroxysulfate for 65 min (flow rate, 1.1 mL/min). The effluent was fluorescently labelled with 2-cyanoacetamide through a T-connector. The labelled eluates were monitored at wavelengths of 346 nm/410 nm (Ex/Em), and the area of the peak corresponding to the disaccharides was used to determine the disaccharide composition of each CS. Chromatograms of the disaccharide analysis of CSs are shown in Supplementary Fig. S5. The sulfur content was calculated as the weight of the sulfur atom (S) per weight of each CS (based on the disaccharide composition) as follows:
where α and β are the contents (%) of unsaturated disaccharides in sodium form with one sulfate group (Δ-4,5-unsaturated hexuronic acid [ΔΗexUA] − C4-sulfated GalNAc and ΔΗexUA − C6-sulfated GalNAc) and two sulfate groups (C2-sulfated ΔΗexUA − C6-sulfated GalNAc and ΔΗexUA − C4,6-disulfated GalNAc), respectively.
Fecal samples
Fresh fecal samples were obtained from 10 healthy Japanese volunteers, 60% of whom were female. The inclusion criteria were as follows: Japanese ancestry, no pre-existing illness (according to patient interviews), aged 20–60 years, nonsmoker, and no antibiotic treatment for at least 6 months prior to sampling. The study design was approved by the institutional ethics review board of Kobe University Hospital Clinical and Translational Research Center (research code 1902, approval date May 10, 2016), and all participants provided written informed consent before fecal sample collection. Immediately after collection, each fecal sample was stored in an anaerobic condition with a BD BBL Culture Swab (Becton, Dickinson and Company, NJ, USA) and used within 24 h. This study was conducted following the principles of the Declaration of Helsinki.
Cultivation of fecal samples in KUHIMM
A Bio Jr.8 fermenter (ABLE, Tokyo, Japan) comprising eight parallel and independent anaerobic culturing vessels was used for fecal sample cultivation as described previously49. Briefly, 0.5 g of fecal samples were suspended in 2 mL of PBS buffer (nacalai tesque, Kyoto, Japan). Each vessel containing 100 mL of Gifu anaerobic medium (Nissui Pharmaceutical Co., Ltd, Tokyo, Japan) was inoculated with either 100 μL of fecal suspension alone or with 0.3% (3 g/L) of CSs and then cultivated anaerobically at 37 °C (n = 10). The culture broth was stirred at 300 rpm and continuously purged with an anaerobic gas mixture (N2:CO2 = 80:20) to maintain anaerobic conditions. After 48 h of cultivation, the culture broths were collected and used for subsequent analyses.
Measurement of CS concentration
The supernatant of the culture broths collected at 0 and 48 h was diluted four times with water, and CS concentrations were determined by analyzing the disaccharide composition as described above. The residual CS ratio after 48 h of fermentation was calculated as follows:
DNA extraction
Microbial genomic DNA was extracted from fecal suspensions and culture broths as described previously16.
Sequencing of 16S rRNA genes
The V3–V4 region of bacterial 16S rRNA genes was amplified using the extracted DNA samples as the template, as previously described15, 50. Following the manufacturer’s instructions, polymerase chain reaction (PCR) was performed with an Nextera XT index adapter added to the gene sequence (Illumina Inc., San Diego, CA, USA). Amplicons were purified using AMPure XP DNA purification beads in accordance with the manufacturer’s instructions (Beckman Coulter, Brea, CA, USA). The concentration of the purified amplicons was measured using a Qubit fluorometer (Thermo Fisher Inc., Waltham, MA, USA). The amplicons were pooled at an equimolar concentration of 5 nM. The 16S rRNA genes and internal PhiX control (Illumina) were analyzed for paired-end sequencing using MiSeq (Illumina) with Reagent Kit v3 (Illumina) for 600 cycles. Pair-end reads with a Q score of 20 or higher were combined using automated CASAVA 1.8 pair-end demultiplexing FASTQ, with the FASTQ Generation in Basespace Sequence Hub (https://basespace.illumina.com/). The sequences were subjected to quality control and corrected with the DADA2 pipeline using QIIME 2 version 2022.251. The OTUs were classified using the naive Bayes classifier trained on the Greengenes 13_8 99% OTU full-length sequence database. The OTUs and taxonomic metadata were used for α-diversity estimation.
Quantification of total eubacterial growth and microbial composition analysis
Quantitative real-time PCR for the quantification of total bacterial growth and for microbial composition analysis were conducted as described previously16. A LightCycler 96 system (Roche, Basel, Switzerland) and the primer sets targeting all eubacteria52, 53 were used for the quantitative real-time PCR.
SCFA analysis
The concentrations of acetate, propionate, and butyrate in the culture supernatants were determined via HPLC (Shimadzu), as described previously54.
Metabolome analysis
The supernatants of the KUHIMM culture were filtered through a 5-kDa cut-off filter (ULTRAFREE-MC-PLHCC; Human Metabolome Technologies, Yamagata, Japan), and the filtrates were concentrated by centrifugation and resuspended in 50 μL of ultrapure water immediately before measurement. All metabolome measurements were performed using Capillary Electrophoresis Time-of-Flight Mass Spectrometry (CE-TOFMS) at a facility service at Human Metabolome Technologies Inc55.
Statistical analyses
All statistical analyses in this study were performed using Prism 9 (GraphPad Software, Inc., San Diego, CA, USA). Results with p value < 0.05 were considered statistically significant.
Data availability
All 16S rRNA gene sequences obtained in this study have been deposited at the MG-RAST server (http://metagenomics.anl.gov) as “Model Culture System of Human Colonic Microbiota_CSs” under accession numbers mgm4990853.3-mgm4990912.3.
References
López-Senra, E. et al. Impact of prevalence ratios of chondroitin sulfate (CS)-4 and -6 isomers derived from marine sources in cell proliferation and chondrogenic differentiation processes. Mar. Drugs 18, 94. https://doi.org/10.3390/md18020094 (2020).
du Souich, P., García, A. G., Vergés, J. & Montell, E. Immunomodulatory and anti-inflammatory effects of chondroitin sulphate. J. Cell. Mol. Med. 13, 1451–1463 (2009).
Wang, S. J., Wang, Y. H. & Huang, L. C. The effect of oral low molecular weight liquid hyaluronic acid combination with glucosamine and chondroitin on knee osteoarthritis patients with mild knee pain: An 8-week randomized double-blind placebo-controlled trial. Medicine (Baltimore) 100, e24252. https://doi.org/10.1097/MD.0000000000024252 (2021).
Mikami, T. & Kitagawa, H. Biosynthesis and function of chondroitin sulfate. Biochim. Biophys. Acta 1830, 4719–4733 (2013).
Volpi, N. Condrosulf®: Structural characterization, pharmacological activities and mechanism of action. Curr. Med. Chem. 21, 3949–3961 (2014).
de Abajo, F. J. et al. Risk of nonfatal acute myocardial infarction associated with non-steroidal antiinflammatory drugs, non-narcotic analgesics and other drugs used in osteoarthritis: A nested case-control study. Pharmacoepidemiol. Drug Saf. 23, 1128–1138 (2014).
Morrison, L. M. Response of ischemic heart disease to chondroitin sulfate-A. J. Am. Geriatr. Soc. 17, 913–923 (1969).
Nakazawa, K. & Murata, K. Comparative study of the effects of chondroitin sulfate isomers on atherosclerotic subjects. Z. Alternsforsch. 34, 153–159 (1979).
Melgar-Lesmes, P. et al. Chondroitin sulphate attenuates atherosclerosis in ApoE knockout mice involving cellular regulation of the inflammatory response. Thromb. Haemost. 118, 1329–1339 (2018).
Baici, A. et al. Analysis of glycosaminoglycans in human serum after oral administration of chondroitin sulfate. Rheumatol. Int. 12, 81–88 (1992).
Conte, A. et al. Metabolic fate of exogenous chondroitin sulfate in man. Arzneimittelforschung. 41, 768–772 (1991).
Shang, Q. et al. Degradation of chondroitin sulfate by the gut microbiota of Chinese individuals. Int. J. Biol. Macromol. 86, 112–118 (2016).
Rawat, P. S., Seyed Hameed, A. S., Meng, X. & Liu, W. Utilization of glycosaminoglycans by the human gut microbiota: participating bacteria and their enzymatic machineries. Gut Microbes 14, 2068367. https://doi.org/10.1080/19490976.2022.2068367 (2022).
Shang, Q. et al. Structural modulation of gut microbiota by chondroitin sulfate and its oligosaccharide. Int. J. Biol. Macromol. 89, 489–498 (2016).
Sasaki, K. et al. Taurine does not affect the composition, diversity, or metabolism of human colonic microbiota simulated in a single-batch fermentation system. PLoS ONE 12, e0180991. https://doi.org/10.1371/journal.pone.0180991 (2017).
Takagi, R. et al. A single-batch fermentation system to simulate human colonic microbiota for high-throughput evaluation of prebiotics. PLoS ONE 11, e0160533. https://doi.org/10.1371/journal.pone.0160533 (2016).
Raghavan, V. & Groisman, E. A. Species-specific dynamic responses of gut bacteria to a mammalian glycan. J. Bacteriol. 197, 1538–1548 (2015).
Rakoff-Nahoum, S., Coyne, M. J. & Comstock, L. E. An ecological network of polysaccharide utilization among human intestinal symbionts. Curr. Biol. 24, 40–49 (2014).
Desai, M. S. et al. A dietary fiber-deprived gut microbiota degrades the colonic mucus barrier and enhances pathogen susceptibility. Cell 167, 1339-1353.e21 (2016).
Ulmer, J. E. et al. Characterization of glycosaminoglycan (GAG) sulfatases from the human gut symbiont Bacteroides thetaiotaomicron reveals the first GAG-specific bacterial endosulfatase. J. Biol. Chem. 289, 24289–24303 (2014).
Rey, F. E. et al. Metabolic niche of a prominent sulfate-reducing human gut bacterium. Proc. Natl. Acad. Sci. USA 110, 13582–13587 (2013).
Louis, P. & Flint, H. J. Formation of propionate and butyrate by the human colonic microbiota. Environ. Microbiol. 19, 29–41 (2017).
Horvath, T. D. et al. Bacteroides ovatus colonization influences the abundance of intestinal short chain fatty acids and neurotransmitters. iScience 25, 104158. https://doi.org/10.1016/j.isci.2022.104158 (2022).
Louis, P. & Flint, H. J. Diversity, metabolism and microbial ecology of butyrate-producing bacteria from the human large intestine. FEMS Microbiol. Lett. 294, 1–8 (2009).
Chambers, E. S., Preston, T., Frost, G. & Morrison, D. J. Role of gut microbiota-generated short-chain fatty acids in metabolic and cardiovascular health. Curr. Nutr. Rep. 7, 198–206 (2018).
Parada Venegas, D. et al. Short chain fatty acids (SCFAs)-mediated gut epithelial and immune regulation and its relevance for inflammatory bowel diseases. Front. Immunol. 10, 277. https://doi.org/10.3389/fimmu.2019.00277 (2019).
Macia, L. et al. Metabolite-sensing receptors GPR43 and GPR109A facilitate dietary fibre-induced gut homeostasis through regulation of the inflammasome. Nat. Commun. 6, 6734. https://doi.org/10.1038/ncomms7734 (2015).
Maslowski, K. M. et al. Regulation of inflammatory responses by gut microbiota and chemoattractant receptor GPR43. Nature 461, 1282–1286 (2009).
Natarajan, N. et al. Microbial short chain fatty acid metabolites lower blood pressure via endothelial G protein-coupled receptor 41. Physiol. Genom. 48, 826–834 (2016).
Pluznick, J. L. et al. Olfactory receptor responding to gut microbiota-derived signals plays a role in renin secretion and blood pressure regulation. Proc. Natl. Acad. Sci. USA 110, 4410–4415 (2013).
Chambers, E. S. et al. Effects of targeted delivery of propionate to the human colon on appetite regulation, body weight maintenance and adiposity in overweight adults. Gut 64, 1744–1754 (2015).
Frost, G. et al. The short-chain fatty acid acetate reduces appetite via a central homeostatic mechanism. Nat. Commun. 5, 3611. https://doi.org/10.1038/ncomms4611 (2014).
Shoulders, M. D. & Raines, R. T. Collagen structure and stability. Annu. Rev. Biochem. 78, 929–958 (2009).
Civitelli, R. et al. Dietary L-lysine and calcium metabolism in humans. Nutrition 8, 400–405 (1992).
Ho, Y. Y. et al. l-Ornithine stimulates growth hormone release in a manner dependent on the ghrelin system. Food Funct. 8, 2110–2114 (2017).
Miyake, M. et al. Randomised controlled trial of the effects of L-ornithine on stress markers and sleep quality in healthy workers. Nutr. J. 13, 53. https://doi.org/10.1186/1475-2891-13-53 (2014).
Barkhidarian, B., Khorshidi, M., Shab-Bidar, S. & Hashemi, B. Effects of L-citrulline supplementation on blood pressure: A systematic review and meta-analysis. Avicenna J. Phytomed. 9, 10–20 (2019).
Khalaf, D., Krüger, M., Wehland, M., Infanger, M. & Grimm, D. The effects of oral l-arginine and l-citrulline supplementation on blood pressure. Nutrients 11, 1679. https://doi.org/10.3390/nu11071679 (2019).
Shiraseb, F. et al. Effect of l-arginine supplementation on blood pressure in adults: A systematic review and dose-response meta-analysis of randomized clinical trials. Adv. Nutr. 13, 1226–1242 (2022).
Nakamura, Y., Yamamoto, N., Sakai, K. & Takano, T. Antihypertensive effect of sour milk and peptides isolated from it that are inhibitors to angiotensin I-converting enzyme. J. Dairy Sci. 78, 1253–1257 (1995).
Nakamura, Y. et al. Purification and characterization of angiotensin I-converting enzyme inhibitors from sour milk. J. Dairy Sci. 78, 777–783 (1995).
Cicero, A. F. et al. Effect of lactotripeptides (isoleucine-proline-proline/valine-proline-proline) on blood pressure and arterial stiffness changes in subjects with suboptimal blood pressure control and metabolic syndrome: A double-blind, randomized, crossover clinical trial. Metab. Syndr. Relat. Disord. 14, 161–166 (2016).
Siltari, A., Roivanen, J., Korpela, R. & Vapaatalo, H. Long-term feeding with bioactive tripeptides in aged hypertensive and normotensive rats: Special focus on blood pressure and bradykinin-induced vascular reactivity. J. Physiol. Pharmacol. 68, 407–418 (2017).
Chakrabarti, S. & Wu, J. Milk-derived tripeptides IPP (Ile-Pro-Pro) and VPP (Val-Pro-Pro) promote adipocyte differentiation and inhibit inflammation in 3T3-F442A cells. PLoS ONE 10, e0117492. https://doi.org/10.1371/journal.pone.0117492 (2015).
Huttunen, M. M., Pekkinen, M., Ahlström, M. E. & Lamberg-Allardt, C. J. Long-term effects of tripeptide Ile-Pro-Pro on osteoblast differentiation in vitro. J. Nutr. Biochem. 19, 708–715 (2008).
Iovu, M., Dumais, G. & du Souich, P. Anti-inflammatory activity of chondroitin sulfate. Osteoarthritis Cartil. 16(Suppl 3), S14–S18 (2008).
Mazzucchelli, R. et al. Risk of acute myocardial infarction among new users of chondroitin sulfate: A nested case-control study. PLoS ONE 16, e0253932. https://doi.org/10.1371/journal.pone.0253932 (2021).
Takeuchi, R. et al. Effects of vibration and hyaluronic acid on activation of three-dimensional cultured chondrocytes. Arthritis Rheum. 54, 1897–1905 (2006).
Sasaki, D. et al. Low amounts of dietary fibre increase in vitro production of short-chain fatty acids without changing human colonic microbiota structure. Sci. Rep. 8, 435. https://doi.org/10.1038/s41598-017-18877-8 (2018).
Klindworth, A. et al. Evaluation of general 16S ribosomal RNA gene PCR primers for classical and next-generation sequencing-based diversity studies. Nucleic Acids Res. 41, e1. https://doi.org/10.1093/nar/gks808 (2013).
Callahan, B. J. et al. DADA2: High-resolution sample inference from Illumina amplicon data. Nat. methods 13, 581–583 (2016).
Matsuki, T. et al. Quantitative PCR with 16S rRNA-gene-targeted species-specific primers for analysis of human intestinal bifidobacteria. Appl. Environ. Microbiol. 70, 167–173 (2004).
Rinttilä, T., Kassinen, A., Malinen, E., Krogius, L. & Palva, A. Development of an extensive set of 16S rDNA-targeted primers for quantification of pathogenic and indigenous bacteria in faecal samples by real-time PCR. J. Appl. Microbiol. 97, 1166–1177 (2004).
Yoshida, N. et al. Effect of resistant starch on the gut microbiota and its metabolites in patients with coronary artery disease. J. Atheroscler. Thromb. 26, 705–719 (2019).
Ohashi, Y. et al. Depiction of metabolome changes in histidine-starved Escherichia coli by CE-TOFMS. Mol. Biosyst. 4, 135–147 (2008).
Acknowledgements
This study was partly founded by the Japan Society for the Promotion of Science (KAKENHI Grant number 20K05938) and the Japan Agency for Medical Research and Development (AMED Grant number JP21ae0121036).
Author information
Authors and Affiliations
Contributions
K.I., D.S., and M.I. wrote the manuscript. D.S. and A.K. conceived and designed the experiments. D.S. operated and analyzed the model culture system. K.K. performed the characterization and measurement of CS. M.I. and Y.O. contributed to the metabolome analysis. D.S., M.I., and Y.O. revised the manuscript. A.K. conceived and supervised the research. All authors read and approved the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Inokuma, K., Sasaki, D., Kurata, K. et al. Sulfated and non-sulfated chondroitin affect the composition and metabolism of human colonic microbiota simulated in an in vitro fermentation system. Sci Rep 13, 12313 (2023). https://doi.org/10.1038/s41598-023-38849-5
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-023-38849-5
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.