Abstract
Teratomas in mice, composed of different tissue types, are derived from primordial germ cells in the fetal gonads. Previously, we identified a locus responsible for experimental testicular teratoma (ETT) formation on chromosome 18, referred to as ett1. The strongest candidate sequence in the ett1 locus was found to be a missense mutation in the melanocortin 4 receptor (Mc4r), Mc4rG25S. We established a strain with a point mutation in the Mc4r gene in the ETT-nonsusceptible LT strain, called LT- Mc4rG25S, by genome editing. Surprisingly, highly developed ovarian teratomas (OTs), rather than testicular teratomas, appeared in the LT-Mc4rG25S strain. The results demonstrated that Mc4r is also one of the genes responsible for OT formation and suggested that missense mutations in Mc4r promote teratoma formation in both sexes. In this study, we performed ETT experiments in different host–graft combinations of the LT-Mc4rG25S and LT strains. Furthermore, the expression of MC4R in germ cells in the testis was demonstrated. Expression of Mc4r in testis was also confirmed by RT-PCR. The results demonstrated that MC4R is expressed in germ cells in the testis and that a point mutation in the Mc4r gene is responsible for ETT formation.
Similar content being viewed by others
Introduction
A teratoma is a tumor of germ cell origin containing tridermic tissues. Teratoma formation has been reported in mice as well as in humans and birds. Teratomas occur congenitally but can also be induced experimentally. Teratoma formation is also used to verify whether stem cells retain pluripotency. Teratomas are formed when pluripotent cells (embryonic carcinoma cells (EC cells), embryonic stem cells (ES cells), induced pluripotent stem cells (iPS cells)) are transplanted into differentiated tissues1,2,3,4. Spontaneous testicular teratoma (STT), an inborn teratoma formed in the mouse testis, is a differentiated pluripotent tumor containing tridermic tissues derived from primordial germ cells (PGCs). It has been suggested that 6–8 recessive loci are involved in the formation of STTs in the 129/Sv mouse strain5. One of these strains, the 129/Sv-+/Ter strain bearing the Ter mutation on chromosome 18, is known to have an increased rate of STT formation6,7. Noguchi and Noguchi established that the 129/Sv-+/Ter strain develops STTs at a rate of almost 100%8. The ter mutation was identified in 2005 as a nonsense mutation in the dead end 1 (Dnd1) gene, a homolog of the dead end (dnd) gene that causes primordial germ cell defects in zebrafish9,10.
Furthermore, the formation of experimental testicular teratoma (ETT) can be induced in the 129 subline by implanting the testis of a 12.5-day embryo (E12.5) into an adult testis11. Stevens performed an experiment in which fetal testes from 129/Sv-C P Sl mice that carried the Steel (Sl) mutation in the kit gene were implanted into adult testes. Mice with a homozygous Sl/Sl genotype develop germ cell defects. However, the wild-type and Sl/ + genotypes had a teratoma formation rate of 70–80%, while mice with the Sl/Sl genotype almost never formed teratomas12. This experiment showed that the teratoma was of germline origin. The rate of ETT formation also depends on the age of the transplanted testes. It was reported that 82% of 12.5-day fetal testes transplanted into adult testes formed ETTs, while the rate of ETT formation decreased to 8% when E13.5 fetal testes were transplanted11. Noguchi et al. transplanted fetal testes from a Ter congenic line (B6-+/Ter) and found that ETTs were not formed13. These results indicate that other genes are responsible for ETT formation14,15. In addition, when fetal testes from 12.5-day embryos of 129/Sv-+/Ter wild-type (+/+) mice were transplanted, ETTs formed 80–90% of the time, even though they did not carry the Ter mutation. However, transfer of fetal testes from B6 or LTXBJ(LT)16 rather than the 129 strain does not result in the formation of ETTs. The fact that STTs and ETTs are not formed in the Ter/Ter and + /Ter congenic lines and that ETTs are formed when a fetal testis from the wild-type Ter 129 is transplanted into an adult testis indicates that there are other genes in the genetic background of line 129 that are responsible for ETT formation. We have focused on identifying the causative genes and elucidating the mechanism of teratoma formation in ETT.
The F2 generation was obtained by crossing the F1 [LTXBJ × 129/Sv-Ter+/+] x F1 [LTXBJ × 129/Sv-Ter+/+] generations of the teratoma-forming line 129 and the nonteratoma-forming line LT to identify the region in which the gene responsible for experimental testicular teratoma formation resides. Fetal testes were transplanted into F1 adult testes for teratoma formation assays, followed by simple sequence length polymorphism (SSLP) genotyping and linkage analysis using microsatellite markers on chromosomes 1 through 19. Three regions containing candidate causative genes for experimental testicular teratoma, named experimental testicular teratoma 1, 2, 3 (ett1, ett2, and ett3), were identified on chromosomes 18, 3, and 7, respectively17,18. Exome sequencing analysis was performed to search for mutations in these three regions, and 129-lineage-specific SNPs were identified, including one in Mc4r in the ett1 region18. In a previous study, we established a gene-edited strain, LT-Mc4rG25S, by using an electroporation-based gene transfer method (TAKE)20. Surprisingly, highly developed ovarian teratomas (OTs) formed in the female mice of this strain21. This result demonstrated that Mc4r is one of the genes responsible for ovarian teratoma formation.
Mc4r is involved in energy intake and energy expenditure; thus, the gene knockout strain shows an obese phenotype22,23. As already mentioned in a previous paper, we hypothesized that instead of melanocyte-stimulating hormone (MSH), an authentic ligand for MC, agouti and agouti-related protein (AGRP) is thought to be a ligand for MC4R expressed in mouse germ cells21. One piece of supportive knowledge is that agouti gene mutation causes a tenfold lower rate of testicular teratoma formation24. Ay is a dominant mutation in the agouti gene and causes ectopic expression of agouti protein, now it is referred to agouti-signaling protein-1 (ASIP1), and the inhibition of MC1R and MC4R chronic signal transduction systems25,26,27. ASIP1 is an antagonist for MC4R and high expression of ASIP1 in agouti-mice (Ay/a) resulted in obese phenotype as Mc4r-/- mice28. The increased expression of Mc4r in agouti-mice was demonstrated and suggested that it was the compensatory mechanism for prevention of ASIP1-mediated inhibition of the MC4R-signaling in the obese agouti-mice. AGRP was found as antagonist for MC3R and MC4R29. AGRP was subsequently shown to act as an inverse agonist against for MC4R30. AGRP has also been shown to act as a biased agonist for MC4R31. Interestingly, expression of asip1 and agrp detected from fetal testis and its expression continue to adult32. It is possible that these ligands regulate the germ cell survival, division or apoptosis through MC4R-signaling.
In this study, we investigated whether Mc4rG25S is responsible for ETT formation. Transplantation of fetal testes from Mc4rG25S mice into adult testes in Mc4rG25S mice resulted in a high rate of ETT formation. These results demonstrate that Mc4rG25S is responsible for ETT formation.
Results
Strains homozygous for a knock-in mutation substituting G in LT to A in 129 to produce Mc4rG25S were established by the TAKE (Technique for Animal Knockout system by Electroporation) method20 as previously described21 (Fig. 1A). The LT- Mc4rG25S strain females showed OT formation. In contrast, no obvious teratoma formation was observed in the males, with no incidence of STT found in testis sections among hundreds of mice during breeding (Fig. 1B,C). Thus, the SNP Mc4rG25S is not sufficient for STT formation in the LT genetic background. We conducted transplantation experiments to determine whether the SNP Mc4rG25S is responsible for experimental testicular teratoma (ETT) formation.
STT was not found in LT-Mc4rG25S/G25S male mice. (A) Photographs of LT and LT-Mc4rG25S/G25S male mice. (B) Morphology of the testes of LT-Mc4rG25S/G25S and LT. Scale bar = 5 mm. (C) A section of testis from a LT-Mc4rG25S/G25S mouse. Scale bar = 500 μm. (D) A section of seminiferous tubule. Scale bar = 50 μm.
First, we transplanted LT-Mc4rG25S/G25S fetal testes as donors into LT-Mc4rG25S/G25S adult testes as hosts (Fig. 2). In the ETT of one specimen, various types of tissues from all three germ layers, such as thyroid follicular cells, glandular cells (endodermal), keratin pearls (ectodermal), and cartilage tissue (mesodermal), were observed (Fig. 2B–F). Although one specimen (shown in Fig. 2) produced a teratoma composed of tissues that could be identified as derived from all three germ layers, teratomas with tissues of only two or one germ layers were found in the other specimens (Supplementary Fig. S1).
ETT formation by transplantation of fetal testis from LT-Mc4rG25S/G25S into LT-Mc4rG25S/G25S. (A) Morphology of the ETT-forming testes. The right testis was transplanted and sectioned. Scale bar = 5 mm. (B) A testicular teratoma was found in a testis section (the teratoma area developed from transplanted fetal testis is indicated by the dotted circle). Scale bar = 500 μm. (B–E) Images of tissue-like structures found in a teratoma: (C) thyroid follicular cell (Th); (D) glandular cell (Gl); (E) keratin pearl (Kp); (F) cartilage tissue (Ca). Scale bars = 50 μm.
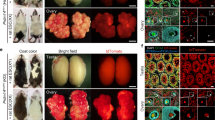
ETTs were also found in another donor–host combination, LT-Mc4 G25S/G25S fetal testis as a donor into LT adult testis as a host (Fig. 3). In the ETT of one specimen, various types of tissues derived from the three germ layers, such as digestive gland (endodermal); keratin pearls (ectodermal); and loose connective tissue (mesodermal), were observed (Fig. 3).
ETT formation by transplantation of fetal testis from LT-Mc4rG25S/G25S into LT. (A) Morphology of the ETT-formed testes. The right testis was transplanted and sectioned. Scale bar = 5 mm. (B) A developed testicular teratoma was found in the section of the testis (the teratoma area developed from transplanted fetal testis is indicated by dotted circle). Scale bar = 500 μm. (B–E) Photographs of tissue-like structures found in a teratoma: (C) digestive gland (Dg); (D) keratin pearl (Kp); (E) loose connective tissue (Lc). Scale bars = 10 μm.
Although one specimen indicated in Fig. 3 produced a teratoma composed of tissues that could be identified as derived from three germ layers, teratoma with tissues derived from only two or one germ layers were found in other specimens (Supplementary Fig. S2).
The incidence of teratoma in each genotype is summarized in Table 1. In total, 26 Mc4rG25S fetal testes were transplanted into adult Mc4rG25S testes, and ETT formation was observed in 6 offspring (23.1%). Similarly, 11 LT-Mc4rG25S fetal testes were transplanted into LT adult testes, and ETT formation was observed in 3 offspring (27.2%). When LT fetal testes were transplanted into LT-Mc4rG25S adult testes (10 cases), no ETT formation was observed (0%), nor were ETTs observed after transplantation of LT fetal testes into LT adult testes17.
The incidence of ETTs was approximately the same for grafts of fetal testes of LT-Mc4rG25S/G25S into LT-Mc4rG25S/G25S and LT testes. These results suggested that the stimulus for ETT development was derived from LT-Mc4G25S/G25S fetal testes.
It was demonstrated by immunofluorescence analysis that MC4R was expressed in germ cells in the fetal testis of LT (Fig. 4).
MC4R is expressed in germ cells in the testis. Immunofluorescence staining of sections of fetal testis from the LT strain. Sections were double stained with anti-rabbit MC4R polyclonal antibodies (α-MC4R), anti-mouse MVH polyclonal antibodies (α-MVH) and DAPI. Merged images of both antibodies and DAPI are also shown (Merge). Control staining with peptide-blocked antibodies is shown below each photograph. Forming seminiferous tubules are indicated by dotted circles. Scale bars = 100 μm.
The signals representing MC4R and a germ cell-specific protein, mouse vasa-homolog (MVH), overlapped well. In a previous study, we showed that MC4R proteins are expressed in oocytes21. We also examined expression of Mc4r in testis by RT-PCR. The results indicated that Mc4r was expressed in both fetal and adult testis of LT and LT-Mc4rG25S/G25S (Fig. 5). These results demonstrated that mouse MC4R is expressed in germ cells.
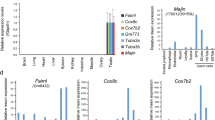
Expression of Mc4r in the fetal and adult testis. RT-PCR was performed with total RNAs extracted from fetal and adult testis from the LT and LT-Mc4rG25S/G25S strains. The amplified band of Mc4r is indicated by arrowhead. PCR amplification is also conducted by using genomic DNA from LT strain as template (gDNA). Genomic DNA was isolated from the ear of LT mouse. Full-length gel images for this figure in different capture setting are indicated in Supplemental figures.
Discussion
In this study, we identified Mc4r as the ett1 gene responsible for ETT formation. We mapped the ett1 region, which is a candidate region for genes involved in ETT formation, and found that among the 8 genes existing in the ett1 region, there is a SNP in Mc4r that causes an amino acid change between the 129 strain and the LT strains 18. Thus, the single amino acid change in Mc4r is proposed to be a candidate for ett1. Then, to validate Mc4r as the ett1 gene, a knock-in strain, LT-Mc4rG25S/G25S, in which this single base substitution was introduced into the LT strain, was created by genome editing. The original LT strain was established as a strain susceptible to OTs in which testicular teratomas (STTs and ETTs) do not develop at all. Surprisingly, we found that a knock-in strain, LT-Mc4rG25S/G25S, developed OTs at an extremely high rate21. The results suggested that Mc4r is a gene responsible for OT formation.
The suggested role of MC4R in germ cells is in the antiapoptotic signaling pathway. As indicated in this study, MC4R is expressed in germ cells in fetal testes. Expression of Mc4r is also demonstrated by RT-PCR. Furthermore, we already reported of the detection of Mc4r expression by qPCR both in fetal and adult testis in 129 and LT strains18. It is hypothesized that the modification of the antiapoptotic pathway of testicular germ cells enhances the transdifferentiation of germ cells into somatic cells or increases the rate of parthenogenesis and causes testicular teratoma formation. Regarding the role of DND1 in suppressing the apoptosis of germ cells33 or maintaining germ cell fate by suppressing the transdifferentiation of germ cells into somatic cells34, the MC4R-mediated signal cascade is involved in germ cell survival, apoptosis and control of the transdifferentiation of germ cells into somatic cells.
These results strongly suggest that Mc4r, an ett1 candidate gene, may be one of the causative genes of STT formation.
In our ETT formation experiments, tridermic teratomas rarely developed. In addition, highly developed teratomas that could be observed externally were not formed. Because the genetic background of this transplantation experiment did not include the Ter gene mutation, it is suggested that Ter gene mutation is required for the development of highly proliferative and differentiated tridermic teratomas. We also identified two additional ETT-related loci, ett2 and ett3, on chromosomes 3 and 7, respectively. It should be clarified whether and how these loci are involved in the development of ETT.
By this study it was suggested that Mc4r is involved in the development of experimental testicular teratoma in mice. It has become clear that Mc4r, whose function in the central nervous system has been attracting attention, is also expressed in germ cells and is involved in their control. Teratomas also develop in human testes and ovaries35. The Ter gene mutation (non-sense mutation of Dnd1) discovered in mouse teratoma studies have been investigated as a cause of human cancers and have been shown to have inhibitory effects on several cancers such as liver cancer and tongue cancer36,37,38. Large-scale data analysis of MC4R also points out that a SNIP close to MC4R gene was related to endometrial cancer and breast cancer39,40. Thus, the relation between MC4R and human gonadal cancer should be analyzed as a possible causative. Data from cBioPortal, which has been opened as a platform for analyzing gene mutations in actual human tumor samples, shows that DND1 has a mutation rate of 0.84% in testicular germ cell tumors (TGCTs)41, while MC4R has a mutation rate of 2.01% (cbioportal.org, last accessed on 2 Mar 2023)42,43. All alteration (mutation, deletion or amplification) of DND1 in TGCTs samples was mutations, whereas deep deletion was all the case of MC4R. These patterns of alteration in DND1 and MC4R were specific for TGCTs among patterns of alteration found in various cancers. These specific mutational patterns may imply specific functions of DND1 and MC4R in the gonad. A detailed analysis of the relationship between MC4R and human TGCTs should be carried out.
Methods
Mice
All animals were housed under temperature-controlled conditions and had free access to food and water. All animal procedures were performed per the protocols approved by the Institutional Animal Care and Use Committee of Shizuoka University (2021A-9, 2022A-5). All experiments were performed in accordance with the guidelines of the committee. The study was carried out in compliance with the ARRIVE guidelines44.
The 129/Sv-Ter and LTXBJ mice16, generous gifts from Dr. Leroy Stevens (The Jackson Laboratory, Bar Harbor, ME, USA), were bred in the animal facilities at Shizuoka University. Males of the LT strain undergo normal spermatogenesis and do not form STTs or ETTs13. The genome-edited LT- Mc4rG25S/G25S strain was established by the TAKE method, as previously described21. The F1 offspring obtained were genotyped by sequence analysis, and offspring of the same genotypes were submitted to sibling mating.
Transplantation of fetal testis into adult testis
To address ETT formation ability, transplantation of fetal testis into adult testis was performed as previously described17 except that the transplanted testis were excised 2–3 months after transplantation to analyze ETT formation in this study.
Observation of testicular teratomas
For each mouse, the testis morphology to three months after transplantation was observed and photographed after dissection. Whether the teratoma developed on the left side, right side, or both sides was recorded.
Histological analysis
Tissues were dissected and fixed in Bouin's solution, embedded in paraffin, and serially sectioned at 6-μm thickness. Deparaffinized sections were stained with hematoxylin, eosin and alcian blue (H-E-A) and screened for the presence of ETTs under a light microscope.
Immunofluorescence observation
The testes were fixed with 4% PFA, dehydrated, cleared, and embedded in paraffin to prepare 5 μm-thick sections. After deparaffinization, the slides were autoclaved with 10 mM sodium citrate buffer (pH 6.0) for 10 min at 120 ℃ for antigen retrieval. Slides were washed 3 times with TBST, and then the tissue was surrounded with a Pap Pen liquid blocker (DAIDO SANGYO CO., LTD, Tokyo Japan). Sections were blocked with 3% BSA/TBST for 1 h at room temperature. The primary antibodies anti-MC4R (1:80, Cayman Chemical) and anti-DDX4/MVH (1:50, Abcam) were diluted in 3% BSA/TBST and incubated with the tissue sections overnight at 4 °C. Sections were washed 3 times with TBST for 10 min each. Then, the secondary antibodies, anti-rabbit IgG (H + L), F(ab')2 fragment (Alexa Fluor 488 conjugate) (1:500, Cell Signaling), anti-mouse IgG (H + L), and F(ab')2 fragment (Alexa Fluor 555 conjugate) (1:500, Cell Signaling), diluted in 3% BSA/TBST, were applied at room temperature for 1 h. After 3 washes with TBST for 10 min each, sections were mounted in a DAPI-containing water-soluble mountant (ProLong). A Carl Zeiss confocal laser scanning microscope (LSM700) was used for fluorescence observation.
Reverse transcription-polymerase chain reaction (RT-PCR)
Total RNA was isolated from fetal or adult testis using pre-made reagents for RNA extraction (ISOGEN, Nippon Gene, Tokyo, Japan) in accordance with the manufacturer’s protocol. RT-PCR was carried out using Ready-to-Go RT-PCR Beads (Amersham Biosciences) in accordance with the manufacturer’s instructions. The gene-specific primers used for Mc4r were as follows: (sense 5′-AAAGGAGGCAAAAACAGGCG-3′ and antisense 5′-CAGGACCAAATGTGTCCGTT-3′). PCR conditions involved incubation for 30 s at 95 °C, 30 s at 67 °C and 60 s at 72 °C for a total of 40 cycles.
Data availability
The datasets generated and/or analyzed during the current study are available from the corresponding author on reasonable request.
References
Evans, M. J. The isolation and properties of a clonal tissue culture strain of pluripotent mouse teratoma cells. J. Embryol. Exp. Morphol. 28, 163–176 (1972).
Evans, M. J. & Kaufman, M. H. Establishment in culture of pluripotential cells from mouse embryos. Nature 292, 154–156. https://doi.org/10.1038/292154a0 (1981).
Martin, G. R. Isolation of a pluripotent cell line from early mouse embryos cultured in medium conditioned by teratocarcinoma stem cells. Proc. Natl. Acad. Sci. USA 78, 7634–7638. https://doi.org/10.1073/pnas.78.12.7634 (1981).
Takahashi, K. & Yamanaka, S. Induction of pluripotent stem cells from mouse embryonic and adult fibroblast cultures by defined factors. Cell 126, 663–676. https://doi.org/10.1016/j.cell.2006.07.024 (2006).
Lam, M. Y., Heaney, J. D., Youngren, K. K., Kawasoe, J. H. & Nadeau, J. H. Trans-generational epistasis between Dnd1Ter and other modifier genes controls susceptibility to testicular germ cell tumors. Hum. Mol. Genet. 16, 2233–2240 (2007).
Asada, Y., Varnum, D. S., Frankel, W. N. & Nadeau, J. H. A mutation in the Ter gene causing increased susceptibility to testicular teratomas maps to mouse chromosome 18. Nat. Genet. 6, 363–368 (1994).
Sakurai, T., Katoh, H., Moriwaki, K., Noguchi, T. & Noguchi, M. The ter primordial germ cell deficiency mutation maps near Grl-1 on mouse chromosome 18. Mamm Genome 5, 333–336 (1994).
Noguchi, T. & Noguchi, M. A recessive mutation (ter) causing germ cell deficiency and a high incidence of congenital testicular teratomas in 129/Sv-ter mice. J. Natl. Cancer Inst. 75, 385–392 (1985).
Weidinger, G. et al. Dead end, a novel vertebrate germ plasm component, is required for zebrafish primordial germ cell migration and survival. Curr. Biol. 13, 1429–1434 (2003).
Youngren, K. K. et al. The Ter mutation in the dead end gene causes germ cell loss and testicular germ cell tumours. Nature 435, 360–364. https://doi.org/10.1038/nature03595 (2005).
Stevens, L. C. Experimental production of testicular teratomas in mice. Proc. Natl. Acad. Sci. USA 52, 654–661 (1964).
Stevens, L. C. Origin of testicular teratomas from primordial germ cells in mice. J. Natl. Cancer Inst. 38, 549–552 (1967).
Noguchi, M. et al. The ter mutation responsible for germ cell deficiency but not testicular nor ovarian teratocarcinogenesis in ter/ter congenic mice. Develop. Growth Differ. 38, 59–69 (1996).
Jiang, L. I. & Nadeau, J. H. 129/Sv mice–a model system for studying germ cell biology and testicular cancer. Mamm Genome 12, 89–94 (2001).
Sekita, Y., Nakamura, T. & Kimura, T. Reprogramming of germ cells into pluripotency. World J. Stem Cells 8, 251–259. https://doi.org/10.4252/wjsc.v8.i8.251 (2016).
Stevens, L. C. & Varnum, D. S. The development of teratomas from parthenogenetically activated ovarian mouse eggs. Dev. Biol. 37, 369–380 (1974).
Miyazaki, T. et al. Identification of genomic locus responsible for experimentally induced testicular teratoma 1 (ett1) on mouse Chr 18. Mamm Genome 25, 317–326. https://doi.org/10.1007/s00335-014-9529-8 (2014).
Miyazaki, T. et al. Identification of Two Additional Genomic Loci Responsible for experimentally induced testicular teratoma 2 and 3 (ett2 and ett3). Zool. Sci. 35, 172–178. https://doi.org/10.2108/zs170176 (2018).
Kaneko, T., Sakuma, T., Yamamoto, T. & Mashimo, T. Simple knockout by electroporation of engineered endonucleases into intact rat embryos. Sci. Rep. 4, 1. https://doi.org/10.1038/srep06382 (2014).
Hashimoto, M. & Takemoto, T. Electroporation enables the efficient mRNA delivery into the mouse zygotes and facilitates CRISPR/Cas9-based genome editing (vol 5, 11315, 2015). Sci. Rep. 5, 1. https://doi.org/10.1038/srep12658 (2015).
Naser, A. A. et al. MC4R mutant mice develop ovarian teratomas. Sci. Rep. 11, 1. https://doi.org/10.1038/s41598-021-83001-w (2021).
Tao, Y. X. The melanocortin-4 receptor: Physiology, pharmacology, and pathophysiology. Endocr. Rev. 31, 506–543. https://doi.org/10.1210/er.2009-0037 (2010).
Huszar, D. et al. Targeted disruption of the melanocortin-4 receptor results in obesity in mice. Cell 88, 131–141 (1997).
Stevens, L. C. The biology of teratomas. Adv. Morphog. 6, 1–31 (1967).
Bultman, S. J., Michaud, E. J. & Woychik, R. P. Molecular characterization of the mouse agouti locus. Cell 71, 1195–1204. https://doi.org/10.1016/s0092-8674(05)80067-4 (1992).
Dinulescu, D. M. & Cone, R. D. Agouti and agouti-related protein: analogies and contrasts. J. Biol. Chem. 275, 6695–6698 (2000).
Miller, M. W. et al. Cloning of the mouse agouti gene predicts a secreted protein ubiquitously expressed in mice carrying the lethal yellow mutation. Genes Dev. 7, 454–467 (1993).
Derkach, K. et al. The evidence of metabolic-improving effect of metformin in Ay/a mice with genetically-induced melanocortin obesity and the contribution of hypothalamic mechanisms to this effect. PLoS ONE 14, e0213779. https://doi.org/10.1371/journal.pone.0213779 (2019).
Ollmann, M. M. et al. Antagonism of central melanocortin receptors in vitro and in vivo by agouti-related protein. Science 278, 135–138. https://doi.org/10.1126/science.278.5335.135 (1997).
Yang, Z. & Tao, Y. X. Biased signaling initiated by agouti-related peptide through human melanocortin-3 and-4 receptors. BBA-Mol. Basis Dis. 1485–1494, 2016. https://doi.org/10.1016/j.bbadis.2016.05.008 (1862).
Buch, T. R., Heling, D., Damm, E., Gudermann, T. & Breit, A. Pertussis toxin-sensitive signaling of melanocortin-4 receptors in hypothalamic GT1-7 cells defines agouti-related protein as a biased agonist. J. Biol. Chem. 284, 26411–26420. https://doi.org/10.1074/jbc.M109.039339 (2009).
Anzai, T., Yamagata, T. & Uosaki, H. Comparative transcriptome landscape of mouse and human hearts. Front. Cell Dev. Biol. 8, 268. https://doi.org/10.3389/fcell.2020.00268 (2020).
Yamaji, M. et al. DND1 maintains germline stem cells via recruitment of the CCR4-NOT complex to target mRNAs. Nature 543, 568–572. https://doi.org/10.1038/nature21690 (2017).
Gross-Thebing, T. et al. The vertebrate protein dead end maintains primordial germ cell fate by inhibiting somatic differentiation. Dev. Cell 43, 704. https://doi.org/10.1016/j.devcel.2017.11.019 (2017).
Cunningham, J. J., Ulbright, T. M., Pera, M. F. & Looijenga, L. H. Lessons from human teratomas to guide development of safe stem cell therapies (vol 30, pg 849, 2012). Nat. Biotechnol. 31, 565–565. https://doi.org/10.1038/nbt0613-565a (2013).
Liu, X. Q. et al. MicroRNA-24 targeting RNA-binding protein DND1 in tongue squamous cell carcinoma. Febs. Lett. 584, 4115–4120. https://doi.org/10.1016/j.febslet.2010.08.040 (2010).
Wampfler, J., Federzoni, E. A., Torbett, B. E., Fey, M. F. & Tschan, M. P. The RNA binding proteins RBM38 and DND1 are repressed in AML and have a novel function in APL differentiation. Leukemia Res. 41, 96–102. https://doi.org/10.1016/j.leukres.2015.12.006 (2016).
Xu, W. L., Gong, F. C., Zhang, T., Chi, B. R. & Wang, J. Y. RNA-binding protein Dnd1 inhibits epithelial-mesenchymal transition and cancer stem cell-related traits on hepatocellular carcinoma cells. Biotechnol. Lett. 39, 1359–1367. https://doi.org/10.1007/s10529-017-2375-5 (2017).
Delahanty, R. J. et al. Association of obesity-related genetic variants with endometrial cancer risk: A report from the shanghai endometrial cancer genetics study. Am. J. Epidemiol. 174, 1115–1126. https://doi.org/10.1093/aje/kwr233 (2011).
da Cunha, P. A. et al. Interaction between obesity-related genes, FTO and MC4R, associated to an increase of breast cancer risk. Mol. Biol. Rep. 40, 6657–6664. https://doi.org/10.1007/s11033-013-2780-3 (2013).
Zhang, Y. et al. In silico analysis of DND1 and its co-expressed genes in human cancers. Biochem. Biophys. Rep. 29, 1. https://doi.org/10.1016/j.bbrep.2022.101206 (2022).
Cerami, E. et al. The cBio cancer genomics portal: An open platform for exploring multidimensional cancer genomics data (vol 2, pg 401, 2012). Cancer Discov. 2, 960–960. https://doi.org/10.1158/2159-8290.Cd-12-0326 (2012).
Gao, J. J. et al. Integrative analysis of complex cancer genomics and clinical profiles using the cBioPortal. Sci Signal 6, 1. https://doi.org/10.1126/scisignal.2004088 (2013).
du Sert, N. P. et al. Reporting animal research: Explanation and elaboration for the ARRIVE guidelines 20. PLoS Biol. 18, 1. https://doi.org/10.1371/journal.pbio.3000411 (2020).
Acknowledgements
This study was supported by Grants-in-Aid for Scientific Research in Priority Areas from the Ministry of Education, Culture, Sports, Science and Technology of Japan to Shizuoka University for preserving special mouse lines. We thank Dr. Stevens for providing the 129/Sv and LTXBJ mice.
Author information
Authors and Affiliations
Contributions
S.S. and K.O. performed the E.T.T. experiments and the histological experiments, analyzed the data and drafted the manuscript. T.M. and S.T. conducted the genome editing. A.N., S.S. and K.O. established and maintained the strains. T.M. performed immunofluorescence staining. K.O. and E.T. performed RT-PCR. TT participated in the study design, supervised the study and wrote the paper. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Seki, S., Ohura, K., Miyazaki, T. et al. The Mc4r gene is responsible for the development of experimentally induced testicular teratomas. Sci Rep 13, 6756 (2023). https://doi.org/10.1038/s41598-023-32784-1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-023-32784-1
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.