Abstract
Gonorrhoea infection rates and the risk of infection from opportunistic pathogens including P. aeruginosa have both risen globally, in part due to increasing broad-spectrum antibiotic resistance. Development of new antimicrobial drugs is necessary and urgent to counter infections from drug resistant bacteria. Aspartate-semialdehyde dehydrogenase (ASADH) is a key enzyme in the aspartate biosynthetic pathway, which is critical for amino acid and metabolite biosynthesis in most microorganisms including important human pathogens. Here we present the first structures of two ASADH proteins from N. gonorrhoeae and P. aeruginosa solved by X-ray crystallography. These high-resolution structures present an ideal platform for in silico drug design, offering potential targets for antimicrobial drug development as emerging multidrug resistant strains of bacteria become more prevalent.
Similar content being viewed by others
Introduction
Aspartate-semialdehyde dehydrogenase (ASADH; EC 1.2.1.11) is an enzyme involved in the aspartate biosynthetic pathway, responsible for catalyzing the reductive dephosphorylation of aspartyl phosphate into aspartate semialdehyde (Fig. 1)1. The pathway is required for production of essential amino acids and metabolites in microorganisms and has proved an attractive target for antimicrobial drugs2,3,4,5,6,7.
An emerging challenge for human health is the increasing microbial resistance against several of the most commonly used antibiotics. Neisseria gonorrhoeae and Pseudomonas aeruginosa are both gram-negative pathogenic bacteria. P. aeruginosa is a gram-negative bacterium and an opportunistic pathogen in humans and plants. In healthy individuals it is responsible for mild or asymptomatic infections, however immunocompromised individuals, patients with cystic fibrosis, newborns and pre-term, or low birth weight neonates are at higher risk of acute P. aeruginosa infection and/or complications8,9. P. aeruginosa displays an adaptive resistance that involves biofilm formation, which can act as a diffusion barrier to antibiotics in the lungs8, and antibiotic resistance is an increasing challenge in the treatment of infection10. P. aeruginosa exhibits antibiotic resistance through a variety of mechanisms including horizontal transfer of resistance genes and/or gene mutations, expression of efflux pumps that remove antibiotics from the cell, production of enzymes that inactivate the antibiotic, and a low outer membrane permeability8. Pseudomonas mallei and Pseudomonas pseudomallei (also known as Burkholderia mallei and Burkholderia pseudomallei) are two pathogenic organisms closely related to P. aeruginosa that are also responsible for the human diseases, including glanders and melioidosis, respectively11.
Neisseria meningitides and N. gonorrhoeae are two pathogenic species responsible for diseases such as gonorrhoea, cervicitis, urethritis, pharyngitis, proctitis and occasionally conjunctivitis. Newborn exposure to an N. gonorrhoeae infection during birth can result in ocular infections (ophthalmia neonatorum). Common antimicrobial drugs used to treat a variety of infectious microorganisms, including doxycycline and erythromycin, increase selective pressure on infectious agents such as N. gonorrhoeae12 and these selective pressures enhance the emergence of resistant strains of N. gonorrhoeae13,14,15,16,17. Of further concern, N. gonorrhoeae is a cofactor that can facilitate HIV-1 infection18. In 2015, in response to the threat of antimicrobial resistance the World Health Organization (WHO) adopted a global action plan. The plan briefly involves increased awareness and understanding, increased surveillance and research, increased preventative measures, increased investment for treatment, diagnosis, vaccination and other interventions, and to ensure that the plan is inclusive for all countries19. The design of new antimicrobial drugs is critical to counter emerging multidrug-resistant pathogenic microorganisms.
ASADH has been identified as a target for antimicrobial drugs in both pathogenic and non-pathogenic microorganisms including gram-negative bacteria20,21, gram-positive bacteria22, Mycobacterium tuberculosis23, and archael24 and fungal species2,25,26. Humans and other mammals do not have a homolog of ASADH or require the aspartate biosynthetic pathway, making this a highly attractive pathway to target in microorganisms. Some organisms possess a second, redundant ASADH enzyme, which despite sharing a high structural similarity has less than 50% sequence identity to other functional ASADH proteins27. Despite ASADH sharing an overall high structural similarity between fungal and bacterial orthologs, secondary structural differences have been identified that offer the potential for the development of species-selective antimicrobial drugs2.
Several antibiotic resistance genes have been identified within the P. aeruginosa genome and antibiotic resistance through horizontal gene transfer has been demonstrated8,28. Therefore, new inhibitors are required to combat antibiotic resistance. For microorganisms that use the aspartate biosynthesis pathway for the production of essential amino acids and metabolites, inhibition of ASADH is fatal29. The asd gene encoding ASADH is one of the minimal genes essential for life in these microorganisms7,30,31,32,33.
Currently, inhibitors are used in combination with various antibiotics on antimicrobial-resistant pathogens34 and the use of antibiotic adjuvants can effectively reduce bacterial resistance to commonly used antibiotics35,36. Selective inhibition of key enzymes for biosynthesis pathways in microorganisms can be used to counter antimicrobial drug resistance of pathogenic species such as P. aeruginosa. To provide an enhanced understanding of the structural similarities of ASADH between important microorganisms, as well as to provide a platform for the design of new anti-microbials, here we solve and analyze the structures of two ASADH enzymes from N. gonorrhoeae and P. aeruginosa.
Materials and methods
ASADH proteins were purified using standardized protein expression and purification protocols as described previously37. Briefly, open reading frames encoding ASADH were amplified from genomic DNA and cloned into an expression vector encoding an N-terminal non-cleavable 6-His tag. The proteins were expressed in BL21(DE3) bacterial cells and then purified by nickel affinity and size-exclusion chromatography37. The proteins were concentrated to ~ 20 mg/mL and stored at − 80 °C until used in crystallography experiments. The sparse matrix screening method was used to produce crystals from purified ASADH proteins via the sitting drop vapor diffusion method. Crystals of P. aeruginosa ASADH (SSGCID ID: PsaeA.17885.a.B1.PS02341) were grown at 19.6 mg/ml with 4 mM NADP, and mixed 1:1 with Microlytic MCSG1 screen, well D9 (containing 0.2 M sodium chloride, 0.1 M TRIS:HCl pH 8.50, 25% PEG3350) at 289 K. Crystals harvested were cryo-protected with crystallant supplemented with 15% ethylene glycol and 5 mM NADP and flash‐cooled in liquid nitrogen. X-ray diffraction data with a resolution of 2.1 Å was collected at a temperature of 100 K on a RAYONIX MX-225 CCD detector at a wavelength of 0.9787 Å on the Advanced Photon Source (APS, Argonne, IL) 21‐ID‐F beamline. Crystals from N. gonorrhoeae ASADH (SSGCID ID: NegoA.17885.a.B1.PW37925) were obtained in a condition from Microlytic MCSG1 screen (F11) containing 25% PEG 3350, 200 mM ammonium sulphate, HEPES free acid/NaOH buffer pH 7.5 at temperature 290 K. X-ray diffraction data with a resolution of 2.1 Å was collected at a temperature of 100 K on the RIGAKU SATURN 944 + CCD detector at a wavelength of 1.5418 Å. The diffraction data were reduced using the XDS/XSCALE package38,39. The structures of the P. aeruginosa and N. gonorrhoeae ASADH proteins were solved by molecular replacement using ASADH proteins as starting models from Vibrio cholerae (1MB4) in BALBES40 and Burkholderia thailandensis (3UW3) in MoRDa respectively. Protein structures were refined using PHENIX41. Protein structures shown were modelled using PyMOL and SeeSAR version 9.2 (BioSolveIT GmbH) software42. SeeSAR was utilised to model docking of the SMCS molecule with ASADH proteins. SeeSAR calculates estimated affinities and allows the selection of best poses based on HYDE scoring and analysis of torsion. The default parameter settings were used. The model adopted a similar binding site for docking as defined by “find unoccupied pocket”. From the molecular docking, parameters such as estimated affinity and torsion were acknowledged in selecting the best model. DALI43 (heuristic PDB search) protein structure comparison by alignment of distance matrices were performed. The final protein structures were deposited into the Protein Data Bank (PDB) and assigned the codes 6BAC for N. gonorrhoeae ASADH and 5BNT for P. aeruginosa ASADH.
Results and discussion
Structural analysis of ASADH proteins from N. gonorrhoeae and P. aeruginosa
The structures of ASADH proteins from N. gonorrhoeae and P. aeruginosa were determined using x-ray crystallography, deposited into the Protein Data Bank and assigned the PDB codes 6BAC and 5BNT, respectively (Table 1). Crystals of both ASADH proteins produced x-ray diffraction data with a resolution of 2.1 Å.
The crystallographic structure from N. gonorrhoeae ASADH was solved in the space group P 31 2 1, with unit cell dimensions of a = 109 Å, b = 109 Å, and c = 72.4 Å, and one ASADH protein was present in the asymmetric unit. Continuous electron density allowed modelling of all 371 residues and included two His residues of the non-cleavable N-terminal polyhistidine tag (MAHHHHHH) present in the structure at the N terminus of the protein. The enzyme structure adopted a well-described coenzyme binding domain at the N-terminus (residues 1–150) comprised of six α-helices and seven β-strands, and a catalytic and dimerization domain at the C-terminus (residues 152–351; Fig. 2A,B). The parallel β-sheet flanked by two alpha-helical bundles forms a Rossmann fold, which typically binds enzyme cofactors including β-nicotinamide adenine dinucleotide (NAD) and β-nicotinamide adenine dinucleotide phosphate (NADP)44. The highest structural homology to N. gonorrhoeae ASADH, was the ASADH enzyme from B. thailandensis (3UW3)45, with an RMSD of 0.7 Å (Table 2).
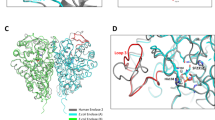
Schematic representation of the structure of ASADH from N. gonorrhoeae (6BAC). (A) Monomeric N. gonorrhoeae ASADH, with α helices colored cyan and β-sheets colored purple. The N- and C-termini are shown in yellow and green, respectively. (B) Topology map of N. gonorrhoeae ASADH showing helices (cylindrical) and directional beta sheets (arrows) analyzed in PDBsum47. (C) N. gonorrhoeae ASADH dimer, with one monomer colored as for A, and the second monomer colored in yellow.
To determine whether N. gonorrhoeae ASADH was likely to form a multimer, we examined the crystal structure by expanding the asymmetric unit. The biological assembly for the ASADH enzyme (6BAC), predicted using Proteins, Interfaces, Structures and Assemblies (PISA), was a dimer with a large interface present between two protomers46 (Fig. 2C). Overall, the protein interface exhibits 52 hydrogen bonds, 18 salt bridges and 3406 Å2 of buried surface area. This biological interface was structurally equivalent to the ASADH protein dimer from B. thailandensis (PDB 3UW3) which exhibited 47 hydrogen bonds, 17 salt bridges and 3463 Å2 of buried surface area.
The structure of the P. aeruginosa ASADH was solved in the space group P 41 21 2, with unit cell dimensions of a = 125.68, b = 125.68, and c = 199.44. The asymmetric unit contained four ASADH protomers arranged as two dimers. The electron density allowed for modelling of residues 1–370. Each monomer is comprised of an N-terminal coenzyme-binding domain (residues 1–133) and a C-terminal catalytic and dimerization domain (residues 135–351), with similar architecture and topology to the ASADH from N. gonorrhoeae (Fig. 3A,B). PISA analysis identified a dimer as the biological unit (Fig. 3C), with the protein interface comprised of 56 hydrogen bonds, 23 salt bridges and 3558 Å2 of buried surface area46. This interface was comparable to ASADH enzymes exhibiting NADP binding including Haemophilus influenzae ASADH (1PQU) involving 52 hydrogen bonds, 18 salt bridges and 3415 Å2 of buried surface area, and V. cholerae ASADH (3PZR) involving 60 hydrogen bonds, 24 salt bridges and 3451 Å2 of buried surface area. In addition, there was a high structural similarity between P. aeruginosa ASADH (5BNT) and the Haemophilus influenzae (1PQU) and V. cholerae (3PZR) ASADH proteins, with RMSD of 0.6 Å and 0.5 Å respectively as determined in DALI48.
Schematic representation of the structure of ASADH from P. aeruginosa (5BNT). (A) P. aeruginosa ASADH structure with α helices colored pink and β-sheets colored green. The N terminus (pale yellow) and C terminus (pale green) are depicted. (B) Topology map of P. aeruginosa ASADH showing helices (cylindrical) and directional beta sheets (arrows) obtained from PDBsum47. (C) P. aeruginosa ASADH formed a dimer with one NADP molecule (yellow) bound within each protomer.
NADP binding in P. aeruginosa ASADH
P. aeruginosa ASADH was co-crystallized with NADP. Each monomer of P. aeruginosa ASADH bound a single NADP molecule (Fig. 4A,B). To examine the conservation of NADP binding residues, structurally similar ASADH proteins exhibiting NADP binding were compared (Fig. 4C). We found many NADP binding residues to be strictly conserved between P. aeruginosa (5BNT) and two structurally similar ASADH proteins; H. influenzae (1PQU) and V. cholerae (3PZR) (Fig. 4D). Overall, the P. aeruginosa ASADH-NADP interface was mediated through 14 hydrogen bonds with a buried surface area of 582.6 Å2. Comparable ASADH-NADP binding was observed in H. influenzae (1PQU) and V. cholerae (3PZR) involving 13 hydrogen bonds, with buried surface areas of 505.8 Å2 and 502.8 Å2 respectively.
NADP binding in P. aeruginosa (5BNT), H. influenzae (1PQU) and V. cholerae (3PZR) ASADH proteins. (A) Schematic of hydrogen bonds (H-bond) between bonding residues of P. aeruginosa ASADH and NADP. (B) NADP (yellow) bound within the P. aeruginosa ASADH dimer and active site residues within the NADP binding site cavity that are important for H-bonding (green). (C) Residues responsible for NADP H-bonding in P. aeruginosa (5BNT, purple), H. influenzae (1PQU, blue) and V. cholerae (3PZR, green) show several conserved residues in NADP binding. (D) Superimposed structures of P. aeruginosa (5BNT, purple), H. influenzae (1PQU, blue) and V. cholerae (3PZR, green) showing the binding surface of the ASADH proteins (grey) modelled in SeeSAR Version 9.242.
ASADH protein is previously described as having two states, apoenzyme and the NADP-inhibitor complex (with SMCS)20. Changes in enzyme conformation are attributed to NADP binding and involve the repositioning of two surface loops adjacent to the NADP binding site. Here, we show that the NADP binding of ASADH in P. aeruginosa shows a different orientation of the nicotinamide ring relative to other ASADH structures (Fig. 4C). The position of the nicotinamide ring is constrained through hydrogen bonds (Fig. 4A). As binding at the active site is predicted to cause a conformational change in the conserved loops enclosing the bound NADP, the difference in positioning likely affects the ability of NADP to stabilise the ASADH protein dimer. However, as the active site is relatively unchanged with NADP binding, active site binding is likely not significantly affected. This should be further explored using modelling approaches of docking inhibitors.
Comparison to known structures
To determine the sequence similarity of the ASADH proteins to other ASADH proteins of known structures, Clustal Omega49 sequence alignment tools were used to generate a phylogenetic tree (Fig. 5). The selection of ASADH protein sequences for comparison was dependent on the availability of ASADH crystallized structures, which represent primarily organisms of pathogenic interest. P. aeruginosa and N. gonorrhoeae ASADH protein sequences are most closely related to ASADH proteins from B. thailandensis and V. cholerae.
To determine the most structural similar ASADH proteins available from those with structures available on the PDB, a homology search using DALI (heuristic PDB search) was performed48 using N. gonorrhoeae ASADH (6BAC) and P. aeruginosa (5BNT, chain B). Only one representative structure is listed for each organism (Table 2). The protein with the most sequence similarity to ASADH proteins from P. aeruginosa and N. gonorrhoeae is B. thailandensis ASADH (3UW3)45. This protein had a sequence similarity of 73% and RMSD of 0.8 Å and a sequence similarity of 71% and RMSD of 0.7 Å for P. aeruginosa and N. gonorrhoeae ASADH proteins respectively. This was expected for P. aeruginosa ASADH as Burkholderia (previously a part of Pseudomonas) is a gram-negative pathogenic group of bacteria that are closely related to Pseudomonas50. Notably, when comparing the two structures presented here, the NADP-bound ASADH from P. aeruginosa (5BNT) had an RMSD 0.8 Å and 68% sequence similarity to N. gonorrhoeae ASADH (6BAC).
The ASADH protein structures with the next most similar structural homology to the P. aeruginosa ASADH (5BNT) is V. cholerae ASADH. The V. cholerae ASADH protein was crystallized with NADP and product of S-carbamoyl-L-cysteine (3PZR), S-allyl-L-cysteine sulfoxide (3Q0E) and NADP and S-methyl-L-cysteine sulfoxide (1MB4)6,20. The crystal structures showed an RMSD 0.5 Å and sequence similarity 69% to P. aeruginosa ASADH (5BNT). Comparatively, ASADH protein from N. gonorrhoeae (6BAC) showed RMSD 0.8–0.9 Å and a sequence similarity of 66% to V. cholerae ASADH.
A multiple sequence alignment was generated using the ASADH proteins identified as having a high structural similarity from the DALI heuristic PDB search, highlighting highly conserved residues and binding motifs of ASADH proteins (Fig. 6). Conserved protein sequences include a G–G–G binding motif for nucleoside binding and a T-QA-SG-G binding motif predicted to have a role in active site communication5. ASADH proteins also contain two conserved active site residues; Cys135, which initiates catalysis and His274 (H-bond interaction)51. The STS binding motif previously reported in NADP binding of ASADH proteins is not conserved in the N. gonorrhoeae ASADH (6BAC) sequence, which may affect binding affinity.
Sequence alignment of ASADH proteins. Highly conserved residues are highlighted in grey. The red box represents the helical subdomain region in the bacterial ASADH proteins. The conserved protein sequences for the STS, G–G–G and T-QA-SG-G binding motifs are highlighted in cyan. Protein sequences within the sequence alignment include P. aeruginosa (5BNT), N. gonorrhoeae (6BAC), V. cholerae (3PZR), E. coli (1GL3), H. influenzae (1PQU), B. thailandensis (3UW3), F. tularensis subsp. Tularen (4WOJ), B. dermatitidis (6C85), T. thermophilus HB8 (2YV3), S. pneumoniae (2GYY), M. tuberculosis H37Rv (3TZ6), M. jannaschii (1YS4), T. rubrum (4ZIC), C. neoformans (5CEF), A. fumigatus (5JW6) and C. albicans (3HSK).
Studies to investigate the binding of inhibitors to ASADH both experimentally and using virtual modelling have predicted that a successful ASADH inhibitor will bind through electrostatic interactions with two highly conserved active site arginine residues4. Arginine residues within the active site from S. pneumoniae ASADH (Arg99, Arg245) and V. cholerae ASADH (Arg101, Arg267) bind the phosphate and the carboxyl group moieties of the aspartate β-semialdehyde (ASA) substrate, respectively. The residue Arg99-103 in bacterial ASADH proteins corresponds to Arg112-116 in fungal and archaeal ASADH proteins (Fig. 6, green box).
SMCS binding to ASADH
To determine whether the binding pocket for an active site inhibitor of ASADH, S-methyl-L-cysteine sulfoxide (SMCS), in a structurally similar ASADH protein is conserved in P. aeruginosa (5BNT) and N. gonorrhoeae (6BAC) ASADH proteins, we performed a structural alignment of the binding pockets. Binding of SMCS was modelled from the V. cholerae ASADH (1MB4) structure (Fig. 7A,B). Inactivation of ASADH by SMCS (Cys) is a result of disulfide bond formation at Cys134. The H-bond interaction at His274 is also highly conserved and critical for mediating SMCS interaction. Other H-bond interactions with SMCS in V. cholerae ASADH (1MB4) also occur at Asn133, Gln161, Glu240 and Arg267 (Fig. 7C,D). The docking results show binding site similarity between existing literature and support the prediction that the SMCS molecule could be used to target the P. aeruginosa and N. gonorrhoeae ASADH proteins.
Comparison of active site inhibitor S-methyl-L-cysteine sulfoxide (SMCS) Cys binding in V. cholerae (1MB4), P. aeruginosa (5BNT) and N. gonorrhoeae (6BAC) ASADH proteins. (A) Schematic of H-bond interactions between active site residues of V. cholerae (1MB4, blue) ASADH and SMCS, conserved in P. aeruginosa (5BNT, purple) and N. gonorrhoeae (6BAC, cyan) ASADH proteins. (B) SMCS (Cys, green) bound within the V. cholerae (3PZR) ASADH dimer. Important active site residues for H-bonding within the NADP binding site cavity are shown (yellow). (C) Active site residues responsible for SMCS binding through H-bond interactions in V. cholerae ASADH (1MB4, blue). Cys (green) represents the covalently bound inhibitor, SMCS, in V. cholerae ASADH (1MB4) through the formation of a disulfide bond. Important active site residues are conserved in P. aeruginosa (5BNT, purple) and N. gonorrhoeae (6BAC, cyan) ASADH proteins. (D) Superimposed structures of P. aeruginosa (5BNT, purple), N. gonorrhoeae (6BAC, cyan) and V. cholerae (1MB4, blue).
Several small molecule inhibitors of ASADH proteins with high binding efficiency have been identified (Table 3). Where some ASADH inhibitors are able to selectively inhibit ASADH in gram negative bacteria, other ASADH inhibitors have been shown to selectively inhibit fungal ASADH proteins52. There is an increasing need to design small molecule inhibitors that can selectively target pathogenic microorganisms that display antibiotic resistance.
P. aeruginosa has demonstrated antibiotic resistance to β-lactams, aminoglycosides, quinolones and other antibiotics8. Recently, whole genome sequencing of Pseudomonas spp. revealed antibiotic resistance genes to various antibiotics28. New antimicrobial drugs are required to treat P. aeruginosa infection and to combat antibiotic resistance10. New therapies that offer selective inhibition of key enzymes in pathogenic microorganisms may be an alternative or used in combination with current antibiotics8. Docking software can be used to predict structural and kinetic binding of small molecules. The ASADH protein from V. cholerae (1MB4) was used here to model SMCS binding within the newly solved structures of P. aeruginosa (5BNT) and N. gonorrhoeae (6BAC) ASADH proteins. This ASADH protein showed high structural and sequence similarity to the protein structures presented here, in particular, within conserved active site residues and binding site cavities. Both P. aeruginosa and N. gonorrhoeae ASADH proteins are potential targets for the small molecule inhibitor SMCS. Here, SMCS binding is modelled within ASADH, however, there are various other inhibitors, catalytic intermediates and substrate analogues including phosphonamidites analogues of the substrate aspartyl phosphate53 that could be modelled to predict binding. Selective inhibitors of ASADH proteins have been identified using screening tools and a fragment library containing 378 compounds52,54,55, virtual screening of databases23 and molecular docking approaches4,56. Due to structural similarities with other characterised ASADH proteins, we predict that various other ligands could be modelled within ASADH protein structures presented in this paper via screening of inhibitors shown to bind ASADH proteins.
Conclusion
Antimicrobial drug resistance is an urgent and ongoing medical threat. High-resolution structures of enzymes that are critical for microbial survival represent an important starting point for the development of new antibiotics. Here we present the crystallographic structures P. aeruginosa and N. gonorrhoeae ASADH proteins, that represent ideal platforms for potential in silico drug screening.
References
Viola, R. E. The central enzymes of the aspartate family of amino acid biosynthesis. Acc. Chem. Res. 34, 339–349. https://doi.org/10.1021/ar000057q (2001).
Dahal, G. P. & Viola, R. E. Structure of a fungal form of aspartate-semialdehyde dehydrogenase from Aspergillus fumigatus. Acta Crystallogr. F Struct. Biol. Commun. 73, 36–44. https://doi.org/10.1107/s2053230x16020070 (2017).
Kumar, R., Garg, P. & Bharatam, P. V. Pharmacoinformatics analysis to identify inhibitors of Mtb-ASADH. J. Biomol. Struct. Dyn. 34, 1–14. https://doi.org/10.1080/07391102.2015.1005137 (2016).
Luniwal, A., Wang, L., Pavlovsky, A., Erhardt, P. W. & Viola, R. E. Molecular docking and enzymatic evaluation to identify selective inhibitors of aspartate semialdehyde dehydrogenase. Bioorg. Med. Chem. 20, 2950–2956. https://doi.org/10.1016/j.bmc.2012.03.013 (2012).
Mank, N. J. et al. Structure of aspartate β-semialdehyde dehydrogenase from Francisella tularensis. Acta Crystallogr. F Struct. Biol. Commun. 74, 14–22. https://doi.org/10.1107/s2053230x17017241 (2018).
Pavlovsky, A. G., Liu, X., Faehnle, C. R., Potente, N. & Viola, R. E. Structural characterization of inhibitors with selectivity against members of a homologous enzyme family. Chem. Biol. Drug Des. 79, 128–136. https://doi.org/10.1111/j.1747-0285.2011.01267.x (2012).
Kobayashi, K. et al. Essential Bacillus subtilis genes. Proc. Natl. Acad. Sci. USA 100, 4678–4683. https://doi.org/10.1073/pnas.0730515100 (2003).
Pang, Z., Raudonis, R., Glick, B. R., Lin, T. J. & Cheng, Z. Antibiotic resistance in Pseudomonas aeruginosa: Mechanisms and alternative therapeutic strategies. Biotechnol. Adv. 37, 177–192. https://doi.org/10.1016/j.biotechadv.2018.11.013 (2019).
Foca, M. et al. Endemic Pseudomonas aeruginosa Infection in a Neonatal Intensive Care Unit. N. Engl. J. Med. 343, 695–700. https://doi.org/10.1056/nejm200009073431004 (2000).
Pachori, P., Gothalwal, R. & Gandhi, P. Emergence of antibiotic resistance Pseudomonas aeruginosa in intensive care unit; a critical review. Genes Dis. 6, 109–119. https://doi.org/10.1016/j.gendis.2019.04.001 (2019).
Nguyen, H. V. N., Smith, M.E., & Hayoun, M.A. Glanders and Melioidosis, (2020).
Ng, L. K. & Martin, I. E. The laboratory diagnosis of Neisseria gonorrhoeae. Can. J. Infect. Dis. Med. Microbiol. 16, 15–25. https://doi.org/10.1155/2005/323082 (2005).
Bala, M., Ray, K., Gupta, S. M., Muralidhar, S. & Jain, R. K. Changing trends of antimicrobial susceptibility patterns of Neisseria gonorrhoeae in India and the emergence of ceftriaxone less susceptible N. gonorrhoeae strains. J. Antimicrob. Chemother. 60, 582–586. https://doi.org/10.1093/jac/dkm238 (2007).
Bala, M. & Sood, S. Cephalosporin resistance in Neisseria gonorrhoeae. J. Glob. Infect. Dis. 2, 284–290. https://doi.org/10.4103/0974-777x.68537 (2010).
Tanaka, M. et al. Analysis of mutations within multiple genes associated with resistance in a clinical isolate of Neisseria gonorrhoeae with reduced ceftriaxone susceptibility that shows a multidrug-resistant phenotype. Int. J. Antimicrob. Agents 27, 20–26. https://doi.org/10.1016/j.ijantimicag.2005.08.021 (2006).
Tapsall, J. W., Ndowa, F., Lewis, D. A. & Unemo, M. Meeting the public health challenge of multidrug- and extensively drug-resistant Neisseria gonorrhoeae. Expert Rev. AntiInfect. Ther. 7, 821–834. https://doi.org/10.1586/eri.09.63 (2009).
Unemo, M., Del Rio, C. & Shafer, W. M. Antimicrobial resistance expressed by Neisseria gonorrhoeae: A major global public health problem in the 21st century. Microbiol. Spectr. https://doi.org/10.1128/microbiolspec.EI10-0009-2015 (2016).
Jarvis, G. A. & Chang, T. L. Modulation of HIV transmission by Neisseria gonorrhoeae: Molecular and immunological aspects. Curr. HIV Res. 10, 211–217. https://doi.org/10.2174/157016212800618138 (2012).
Organization, W. H. Global Action Plan on Antimicrobial Resistance, (2015).
Blanco, J., Moore, R. A., Kabaleeswaran, V. & Viola, R. E. A structural basis for the mechanism of aspartate-beta-semialdehyde dehydrogenase from Vibrio cholerae. Prot. Sci. 12, 27–33. https://doi.org/10.1110/ps.0230803 (2003).
Hadfield, A. et al. Structure of aspartate-beta-semialdehyde dehydrogenase from Escherichia coli, a key enzyme in the aspartate family of amino acid biosynthesis. J. Mol. Biol. 289, 991–1002. https://doi.org/10.1006/jmbi.1999.2828 (1999).
Faehnle, C. R., Le Coq, J., Liu, X. & Viola, R. E. Examination of key intermediates in the catalytic cycle of aspartate-beta-semialdehyde dehydrogenase from a gram-positive infectious bacteria. J. Biol. Chem. 281, 31031–31040. https://doi.org/10.1074/jbc.M605926200 (2006).
Kumar, R., Garg, P. & Bharatam, P. V. Shape-based virtual screening, docking, and molecular dynamics simulations to identify Mtb-ASADH inhibitors. J. Biomol. Struct. Dyn. 33, 1082–1093. https://doi.org/10.1080/07391102.2014.929535 (2015).
Faehnle, C. R., Ohren, J. F. & Viola, R. E. A new branch in the family: structure of aspartate-beta-semialdehyde dehydrogenase from Methanococcus jannaschii. J .Mol. Biol. 353, 1055–1068. https://doi.org/10.1016/j.jmb.2005.09.027 (2005).
Arachea, B. T., Liu, X., Pavlovsky, A. G. & Viola, R. E. Expansion of the aspartate beta-semialdehyde dehydrogenase family: The first structure of a fungal ortholog. Acta Crystallogr. D Biol. Crystallogr. 66, 205–212. https://doi.org/10.1107/s0907444909052834 (2010).
Dahal, G. & Viola, R. E. Structure of a fungal form of aspartate semialdehyde dehydrogenase from Cryptococcus neoformans. Acta Crystallogr. F Struct. Biol. Commun. 71, 1365–1371. https://doi.org/10.1107/s2053230x15017495 (2015).
Viola, R. E., Liu, X., Ohren, J. F. & Faehnle, C. R. The structure of a redundant enzyme: A second isoform of aspartate beta-semialdehyde dehydrogenase in Vibrio cholerae. Acta Crystallogr. D Biol. Crystallogr. 64, 321–330. https://doi.org/10.1107/s0907444907068552 (2008).
Meng, L. et al. Antibiotic resistance patterns of Pseudomonas spp. isolated from raw milk revealed by whole genome sequencing. Front. Microbiol. https://doi.org/10.3389/fmicb.2020.01005 (2020).
Galán, J. E., Nakayama, K. & Curtiss, R. 3rd. Cloning and characterization of the asd gene of Salmonella typhimurium: Use in stable maintenance of recombinant plasmids in Salmonella vaccine strains. Gene 94, 29–35. https://doi.org/10.1016/0378-1119(90)90464-3 (1990).
Akerley, B. J. et al. A genome-scale analysis for identification of genes required for growth or survival of Haemophilus influenzae. Proc. Natl. Acad. Sci. USA 99, 966–971. https://doi.org/10.1073/pnas.012602299 (2002).
Salama, N. R., Shepherd, B. & Falkow, S. Global transposon mutagenesis and essential gene analysis of Helicobacter pylori. J. Bacteriol. 186, 7926–7935. https://doi.org/10.1128/jb.186.23.7926-7935.2004 (2004).
Gerdes, S. Y. et al. Experimental determination and system level analysis of essential genes in Escherichia coli MG1655. J. Bacteriol. 185, 5673–5684. https://doi.org/10.1128/jb.185.19.5673-5684.2003 (2003).
Becker, D. et al. Robust Salmonella metabolism limits possibilities for new antimicrobials. Nature 440, 303–307. https://doi.org/10.1038/nature04616 (2006).
Breijyeh, Z., Jubeh, B. & Karaman, R. Resistance of gram-negative bacteria to current antibacterial agents and approaches to resolve it. Molecules 25, 1340 (2020).
González-Bello, C. Antibiotic adjuvants: A strategy to unlock bacterial resistance to antibiotics. Bioorg. Med. Chem. Lett. 27, 4221–4228. https://doi.org/10.1016/j.bmcl.2017.08.027 (2017).
Laws, M., Shaaban, A. & Rahman, K. M. Antibiotic resistance breakers: Current approaches and future directions. FEMS Microbiol. Rev. 43, 490–516. https://doi.org/10.1093/femsre/fuz014 (2019).
Bryan, C. M. et al. High-throughput protein production and purification at the Seattle structural genomics center for infectious disease. Acta Crystallogr. Sect. F Struct. Biol. Cryst. Commun. 67, 1010–1014. https://doi.org/10.1107/S1744309111018367 (2011).
Kabsch, W. XDS. Acta Crystallogr. Sect. D 66, 125–132. https://doi.org/10.1107/S0907444909047337 (2010).
Kabsch, W. Integration, scaling, space-group assignment and post-refinement. Acta Crystallogr. Sect. D 66, 133–144. https://doi.org/10.1107/S0907444909047374 (2010).
Long, F., Vagin, A. A., Young, P. & Murshudov, G. N. BALBES: a molecular-replacement pipeline. Acta. Crystallogr. D Biol. Crystallogr. 64, 125–132. https://doi.org/10.1107/s0907444907050172 (2008).
Adams, P. D. et al. PHENIX: A comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. Sect. D Biol. Crystallogr. 66, 213–221. https://doi.org/10.1107/S0907444909052925 (2010).
SeeSAR (Sankt Augustin, Germany, 2021).
Holm, L. & Laakso, L. M. Dali server update. Nucleic Acids Res. 44, W351-355. https://doi.org/10.1093/nar/gkw357 (2016).
Hanukoglu, I. Proteopedia: Rossmann fold: A beta-alpha-beta fold at dinucleotide binding sites. Biochem. Mol. Biol. Educ. 43, 206–209. https://doi.org/10.1002/bmb.20849 (2015).
Baugh, L. et al. Combining functional and structural genomics to sample the essential Burkholderia structome. PLoS ONE 8, e53851. https://doi.org/10.1371/journal.pone.0053851 (2013).
Krissinel, E. & Henrick, K. Inference of macromolecular assemblies from crystalline state. J. Mol. Biol. 372, 774–797 (2007).
Laskowski, R. A. PDBsum new things. Nucleic Acids Res. 37, D355-359. https://doi.org/10.1093/nar/gkn860 (2009).
Holm, L. Benchmarking fold detection by DaliLite v.5. Bioinformatics 35, 5326–5327. https://doi.org/10.1093/bioinformatics/btz536 (2019).
Sievers, F. et al. Fast, scalable generation of high-quality protein multiple sequence alignments using Clustal Omega. Mol. Syst. Biol. 7, 539. https://doi.org/10.1038/msb.2011.75 (2011).
Yabuuchi, E. et al. Proposal of Burkholderia gen. nov. and transfer of seven species of the genus Pseudomonas homology group II to the new genus, with the type species Burkholderia cepacia (Palleroni and Holmes 1981) comb. nov. Microbiol. Immunol. 36, 1251–1275. https://doi.org/10.1111/j.1348-0421.1992.tb02129.x (1992).
Blanco, J., Moore, R. A., Faehnle, C. R. & Viola, R. E. Critical catalytic functional groups in the mechanism of aspartate-[beta]-semialdehyde dehydrogenase. Acta Crystallogr. Sect. D 60, 1808–1815. https://doi.org/10.1107/S0907444904020104 (2004).
Gao, G., Liu, X., Pavlovsky, A. & Viola, R. E. Identification of selective enzyme inhibitors by fragment library screening. J. Biomol. Screen. 15, 1042–1050. https://doi.org/10.1177/1087057110381383 (2010).
Cox, R. J., Gibson, J. S. & MayoMartín, M. B. Aspartyl phosphonates and phosphoramidates: the first synthetic inhibitors of bacterial aspartate-semialdehyde dehydrogenase. ChemBioChem 3, 874–886. https://doi.org/10.1002/1439-7633(20020902)3:9%3c874::Aid-cbic874%3e3.0.Co;2-v (2002).
Sarver, J. G. et al. Early stage efficacy and toxicology screening for antibiotics and enzyme inhibitors. J. Biomol. Screen. 17, 673–682. https://doi.org/10.1177/1087057112438769 (2012).
Thangavelu, B., Bhansali, P. & Viola, R. E. Elaboration of a fragment library hit produces potent and selective aspartate semialdehyde dehydrogenase inhibitors. Bioorg. Med. Chem. 23, 6622–6631. https://doi.org/10.1016/j.bmc.2015.09.017 (2015).
Wang, X. et al. IMB-XMA0038, a new inhibitor targeting aspartate-semialdehyde dehydrogenase of Mycobacterium tuberculosis. Emerg. Microbes Infect. 10, 2291–2299. https://doi.org/10.1080/22221751.2021.2006578 (2021).
Acknowledgements
We thank the SSGCID cloning and protein production groups at the Center for Infectious Disease Research and the University of Washington. This research was supported by federal funds from the National Institute of Allergy and Infectious Diseases, National Institutes of Health, Department of Health Services, under Contract Nos.: HHNSN272201200025C and HHNSN272201700059C.
Author information
Authors and Affiliations
Contributions
S.L.T. prepared the manuscript. J.W.F. solved the 5BNT structure and J.A. supervised and reviewed it. M.M.M. solved the 6BAC structure, D.D.M. supervised and peer reviewed it. D.D.L., P.J.M., T.E.E. supervised and managed the project. J.K.F. prepared the manuscript. All authors reviewed the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Teakel, S.L., Fairman, J.W., Muruthi, M.M. et al. Structural characterization of aspartate-semialdehyde dehydrogenase from Pseudomonas aeruginosa and Neisseria gonorrhoeae. Sci Rep 12, 14010 (2022). https://doi.org/10.1038/s41598-022-17384-9
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-022-17384-9
This article is cited by
-
Bridging drug discovery through hierarchical subtractive genomics against asd, trpG, and secY of pneumonia causing MDR Staphylococcus aureus
Molecular Genetics and Genomics (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.