Abstract
Electromagnetic radiation is an important environmental factor. It has a potential threat to public health and ecological environment. However, the mechanism by which electromagnetic radiation exerts these biological effects remains unclear. In this study, the effect of Microcystis aeruginosa under electromagnetic radiation (1.8 GHz, 40 V/m) was studied by using transcriptomics. A total of 306 differentially expressed genes, including 121 upregulated and 185 downregulated genes, were obtained in this study. The differentially expressed genes were significantly enriched in the ribosome, oxidative phosphorylation and carbon fixation pathways, indicating that electromagnetic radiation may inhibit protein synthesis and affect cyanobacterial energy metabolism and photosynthesis. The total ATP synthase activity and ATP content significantly increased, whereas H+K+-ATPase activity showed no significant changes. Our results suggest that the energy metabolism pathway may respond positively to electromagnetic radiation. In the future, systematic studies on the effects of electromagnetic radiation based on different intensities, frequencies, and exposure times are warranted; to deeply understand and reveal the target and mechanism of action of electromagnetic exposure on organisms.
Similar content being viewed by others
Introduction
Water bloom not only causes serious disasters to the ecosystem, but also poses a huge safety hazard to people's drinking water. Microcystis aeruginosa (M. aeruginosa) is one of the most common cyanobacterial blooms. M. aeruginosa produces microcystins, which have typical biotoxic effects and can cause serious biosafety hazards1. The effects of chemical substances, such as heavy metal and antibiotics, on the growth of cyanobacteria have been widely reported2,3. Generally, low concentrations of heavy metals and antibiotics promote algal growth, whereas high concentrations have the opposite effects4,5,6,7.
With the rapid development of the electromagnetic environment, excessive electromagnetic radiation has been found to exert potential hazards to animals and plants8,9,10,11. Some studies have reported on the effects of the electromagnetic environment on algal growth. One Study has shown that a specific electromagnetic wave can increase Spirulina platensis chlorophyll content, photosynthetic rate, and biomass12. In our previous study, we found that oxidative stress of M. aeruginosa can be induced under electromagnetic radiation, and regulations on key enzymes of photosynthesis (Rubisco and fructose-1,6-bisphosphate aldolase (FBA)) by electromagnetic radiation indicated that electromagnetic radiation can affect the photosynthesis of M. aeruginosa13.
Some studies have eliminated algae using electromagnetic fields. The pulsed electric field has a significant inhibitory effect on algal growth, and the electric field intensity is the most important impact factor as increasing it can significantly enhance the inhibitory effect on algal growth14. The experimental results indicated that the pulsed magnetic field can kill algae; increase intensity, retention time and that the pulse frequency can enhance the efficiency15. Electromagnetic waves can cause molecular vibrations in algal cells, causing their cell walls to rupture and leading to accumulation of cellular contents, which ultimately leads to cell death16. The influence of electromagnetic radiation on algal growth indicates that the electromagnetic environment may be correlated with aquatic environment safety, which is worthy of further exploration.
Proteomic and transcriptomic methods have been successfully applied to cyanobacteria. These methods have been used to understand the mechanisms of environmental stress (such as antibiotics17, metal ions18, organic pollutants19, nitrogen limitation20) on algae at the molecular level. However, the effects of electromagnetic radiation on the growth of cyanobacteria at the molecular level remain to be elucidated. In our previous study using proteomics, electromagnetic radiation altered the expression levels of photosynthesis-related proteins, and we speculated that the photoreaction system may be the target of electromagnetic radiation on cyanobacteria21. To further determine the effect of electromagnetic radiation on the growth of cyanobacteria, the present study investigated the effects of 1.8 GHz (the most common and widely used mobile communication frequency in mainland China is 1.8 GHz) and 40 V/m electromagnetic radiation on cyanobacteria cells through transcriptomic methods and measured the activity of ATP synthase. Analyze the influence mechanism of the electromagnetic environment on algae growth and explore the methods and means of physical to control algae growth rate, will be of great scientific significance and practical significance to evaluate the potential ecological risk of electromagnetic environment and solve the increasingly serious problem of water bloom.
Results
Differentially expressed genes analysis
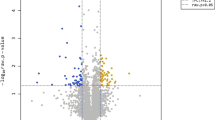
Between the treatment and control groups, 306 differentially expressed genes were determined, 121 and 185 of which were upregulated and downregulated, respectively (Fig. 1). The full list of differentially expressed genes with the fold change and false discovery rate (FDR) detailed can be found as Supplementary Table S1.
Volcanic plot of C versus E (The control group mark was C and the electromagnetic radiation exposure group mark was E. The abscissa indicates the logarithm value of the difference multiple between E and C. The ordinate indicates the negative Log10 value of the FDR of the difference between the two groups. The red (E relative to C, gene expression is upregulated) and green (E relative to C, gene expression is downregulated) dots indicate the difference in gene expression (the criterion is FDR < 0.05, and the difference multiple is more than twice), and the black points are not different).
Pathway enrichment analysis of differentially expressed genes
Exposure of M. aeruginosa to electromagnetic radiation caused significant differential gene expression and corresponding enrichment of the ribosome, oxidative phosphorylation and carbon fixation in photosynthetic pathways of organisms (Table 1).
Sixteen differentially expressed genes were present on the ribosome pathway under electromagnetic stress. The corresponding differentially expressed genes are listed in Table 2. The expression of genes regulating 30S ribosomal proteins S3, S8, S9, S10, S17, and S19 and 50S ribosomal proteins L2, L5, L6, L13, L14, L15, L16, L22, L24, and L29 was downregulated.
Fourteen differentially expressed genes were present on the oxidative phosphorylation pathway of M. aeruginosa under electromagnetic stress. Among them, 13 were downregulated, and 1 was upregulated. The corresponding differentially expressed genes are listed in Table 2. The expression of genes regulating NAD(P)H dehydrogenase, subunit NdhF3 family protein, proton-translocating NADH-quinone oxidoreductase, chain M family protein, NADH-ubiquinone/plastoquinone oxidoreductase chain 6 family protein, and NAD(P)H-quinone oxidoreductase subunit 1, 2, I was downregulated. Moreover, the expression of genes regulating ATP synthase subunit alpha, b', b, beta, delta, gamma, and epsilon chains was downregulated. By contrast, the expression of genes regulating succinate dehydrogenase/fumarate reductase and flavoprotein subunit was upregulated.
Seven differentially expressed genes were related to carbon fixation. Among them, 6 were downregulated, and 1 was upregulated. As shown in Table 2, the expression of genes regulating glyceraldehyde-3-phosphate dehydrogenase, fructose-bisphosphate aldolase class 2, Calvin cycle subtype, d-fructose 1,6-bisphosphatase class 2/sedoheptulose 1,7-bisphosphatase, transketolase, phosphoribulokinase, and ribulose-phosphate 3-epimerase was downregulated. By contrast, the expression of genes regulating phosphoenolpyruvate carboxykinase was upregulated.
Moreover, the differentially expressed genes were mainly related to glycolysis/gluconeogenesis, pentose phosphate pathway, RNA polymerase, photosynthesis, starch and sucrose metabolism, carbon metabolism, photosynthesis—antenna proteins, nitrogen metabolism (Table 1).
Genes and proteins with significant differences
Combined with the results of our previous proteomic study21, five genes or proteins showed simultaneous changes in gene and protein levels after electromagnetic radiation. The expression of the genes C789_RS04790, C789_RS04815 was downregulated, whereas that of iron stress-induced chlorophyll-binding protein was upregulated. The gene and protein levels of ATP synthase subunit b showed a downward trend. The expression of two unknown functional genes and proteins was upregulated. The corresponding differentially expressed genes and proteins are listed in Table 3.
Effect of electromagnetic radiation on ATP synthase
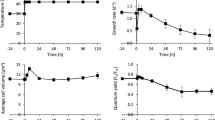
As shown in Fig. 2, 1.8 GHz electromagnetic radiation significantly increased total ATP synthase activity (p < 0.05) and ATP content (p < 0.05). Electromagnetic radiation reduced H+K+-ATP synthase activity, but no significance was observed.
Discussion
Ribosomes are the sites for proteins synthesis. We found that the expression of 16 ribosome-related genes in M. aeruginosa was downregulated after electromagnetic radiation, and the number of 50S ribosomal protein-regulated gene was 10, comprising 62.5% of the total ribosome-related differentially expressed genes. 50S ribosomal proteins were more sensitive to electromagnetic radiation than 30S ribosomal proteins. Significantly downregulated genes S10, L2, S19, L22, S3, L16, and L29 belonged to elongation factor thermo unstable (EF-Tu). EF-Tu is a guanosine nucleotide binding protein and plays a central role in protein synthesis, it is responsible for the extension of the peptide chain during protein synthesis22,23. EF-Tu forms a quaternary complex with the ribosome with aminoacyl-tRNA and guanosine triphosphate (GTP) and is coupled to GTP hydrolysis. As the aminoacyl-tRNA binds to the ribosome, EF-Tu forms a complex with guanosine diphosphate leaving the ribosome. This process continues to circulate so that the peptide chain continues to extend24. EF-Tu also functions as a molecular chaperone; to promote the renaturation of denatured rhodanese, the translation EF-Tu has chaperone-like capacity25 When an organism is exposed to environmental stress, the protein denatures. The protein exposure residues aggregate due to interaction, causing irreversible inactivation of the protein. EF-Tu can prevent the aggregation of exposed hydrophilic groups and protect the structure, which returns to normal after the stress conditions disappear, avoiding permanent denaturation26. After electromagnetic exposure, the downregulation of genes related to EF-Tu may decrease the amount of EF-Tu synthesis, indicating that electromagnetic radiation may inhibit the peptide chain extension of M. aeruginosa cells, affecting protein synthesis, and weaken the function of EF-Tu in preventing protein aggregation.
Cyanobacteria NAD(P)H dehydrogenase is a multi-subunit photosynthetic membrane protein complex located on the thylakoid membrane and plays a vital role in energy metabolism. NAD(P)H-quinone oxidoreductase is involved in many important energy reactions. It was responsible for many functions including respiration, cyclic electron flow around photosystem I (PSI) and CO2 uptake27,28,29. Bernát et al. showed that active electron flow from metabolites to plastoquinone is suppressed upon deletion of ndhF1 and PSI-mediated cyclic electron transport is dependent on NdhF3/F4-type NDH-1 complexes30. Ogawa showed that the ndhB gene is required for Ci transport, the inactivation of the ndhB gene also depressed dark respiration, and NADH dehydrogenase is essential to photoheterotrophic growth and inorganic carbon transport31,32. Stress conditions may alter the NAD(P)H dehydrogenase; for example, the contents of NAD(P)H-dehydrogenase are increased in cells grown in an environment with high salinity33, high light conditions also regulate NADH genes34. Our study shows that electromagnetic radiation also causes downregulation of NAD(P)H-quinone oxidoreductase regulatory gene expression, indicating that electromagnetic radiation may have a certain effect on the NAD(P)H complex of cyanobacteria, which in turn affects a series of physiological processes.
ATP synthase is widely present in the mitochondria and chloroplasts of eukaryotic cells and cytoplasmic membrane of heterotrophic bacteria and photosynthetic bacteria. It participates in oxidative phosphorylation and photosynthetic phosphorylation and catalyzes ATP synthesis under the promotion of transmembrane proton potential. ATP synthase consists of Fo unit and F1 unit, also known as FoF1-ATPase, the Fo unit acts as a proton channel, whereas F1 unit catalyzes ATP synthesis24. The following factors affect the function of ATP synthase: ① ATP synthase function is affected through the subunit of Fo; such as oligomycin binding to the ATP synthase Fo subunit, thereby inhibiting H+ through Fo; dicyclohexylcarbodiimide also inhibits the action of protons through Fo24. ② ATP synthase function is affected by the subunit of F1; when 1 mol of IF1 (the natural ATPase inhibitor from beef heart mitochondria) is bound to 1 mol of F1, the ATPase activity is fully inhibited35. ③ ATP synthase function is affected by the linkage; Subunit ε is a natural endogenous inhibitor of ATP synthase. It plays a dual role in FoF1 from bacteria and chloroplasts. Subunit ε is indispensable for coupling between proton translocation though Fo and ATP synthesis/hydrolysis in F1, and it has a regulatory role inhibiting the ATPase activity of the enzyme36. Stimulation of the external environment has a certain effect on ATPase activity, and ATPase is also part of a stress response mechanism of the organism against external stimuli. Plasma membrane H+ ATPase mitigates of physiological disturbances imposed by salt press37. Our study showed that under the exposure of electromagnetic radiation, the expression of ATP synthase subunit gene was downregulated, it is indicated that electromagnetic radiation may have effects on ATP synthase function. The total ATP synthase activity and the ATP content of cyanobacteria increased significantly. H+K+-ATPase activity was decreased, but no significant change was observed. This may be because the total ATP synthase contains P-type ATPase, F-type ATPase, and many factors affecting ATPase activity. The specific reasons need further study.
The Calvin cycle of photosynthesis consists mainly of three phases: fixation of CO2, reduction of 3-phosphoglycerate, and regeneration of ribulose-1,5-diphosphate. The external environment affects the photosynthetic carbon-fixing enzyme. Studies have shown that suboptimal temperature and low light intensity significantly decreased the growth, photosynthetic rate, activities and mRNA expressions of ribulose 1,5-biphosphate carboxylase/oxygenase, fructose-1,6-bisphosphatase (FBP), glyceraldehyde-3-phosphate dehydrogenase (GAPDH), FBA, transketolase38. Cucumber CsGAPDH gene was significantly induced at 2 h after waterlogging treatment, and the peak level of gene expression was observed at 12 h39. Adversity environment (high temperature stress, salt stress) will affect the expression of CpFBA gene, creating a “short-term inhibition” effect, where short-term stress will be expressed in large quantities and long-term stress will reduce gene expression levels40,41,42. Studies have also shown that CpFBA exhibits periodic variation under adverse conditions43. In our previous study, electromagnetic radiation significantly reduced the activity of FBA enzymes of M. aeruginosa13. The different response modes of cpFBPase mRNA levels to desiccation stress and high temperature indicated that cpFBPase played an important role in responsing to abiotic stress44. The expression of transketolase in rice leaves was upregulated in response to heat stress45, whereas the expression of transketolase in Manihot esculenta Crantz at 5 and 15 days of drought stress was downregulated46. SBPase overexpression enhances photosynthesis under high temperature or salt stress in transgenic rice plants, indicating SBPase plays an important role in plants in stress47,48. Studies have shown that the superior water-deficit tolerance in bermudagrass (Tifway) could be mainly associated with Phosphoribulokinase49. Phosphoribulokinase was downregulated by drought and abscisic acid in maize (Zea mays L.)50. A study has shown that phosphoenolpyruvate carboxylase activity was three times higher in infected plant leaves compared to healthy plants51. In summary, external stress can affect the gene or protein expression of photosynthetic carbon fixation enzymes or enzyme activity. Our study showed that under the exposure of electromagnetic radiation, seven differentially expressed genes were related to carbon fixation; among them, 6 (gap2, Fba, glpX, Tkt, PRK, Rpe) were downregulated, and 1 (pckA) was upregulated, it indicated that electromagnetic radiation may affect the related synthesis of photosynthetic carbon fixation enzymes through the regulation of gene expression.
After electromagnetic radiation, five genes or proteins showed simultaneous changes in gene level and protein levels. Iron stress-induced protein A (IsiA) is the main chlorophyll-binding protein in the thylakoid membrane, and significantly induced under iron deficiency conditions; dynamic changes in IsiA-containing complexes during long-term iron deficiency in cyanobacteria may represent an adaptation to iron limitation stress for flexible light energy distribution, which can balance electron transfer between PS I and PS II, thus minimizing photooxidative damage52. Havaux also showed that IsiA protects cyanobacteria from photooxidativestress53. Our results indicate that electromagnetic radiation may affects the function of iron stress-induced chlorophyll-binding protein. In the study, IsiA gene expression is downregulated, whereas the corresponding protein levels are increased. We speculate that it may be a negative feedback regulation mechanism. CP43′ protein functions as a nonradiative dissipator of light energy, which is encoded by the IsiA gene54. Electromagnetic radiation is a kind of energy, which may promote the upregulation of CP43′ protein. The control and exposure groups were placed in the dark for 24 h. The lack of light may cause CP43′ to be downregulated. Electromagnetic radiation upregulated CP43′ protein. In order to keep CP43′ downregulated, the IsiA gene is downregulated, forming a negative feedback regulation mechanism. The influence of electromagnetic radiation on IsiA gene and related regulatory protein needs further study.
The gene and protein levels of ATP synthase subunit b showed a downward trend after eletcromagnetic exposure. ATP synthase acts on photosynthetic and oxidative phosphorylation to synthesize ATP. ATP synthase b subunit plays an important role in the growth, reproduction and adaptability of organisms. Chen55 and Zheng56 found that the ATP synthase b subunit gene is related to the testis development of drosophila, which will reduce the fecundity of male flies and cause male sterility. Li et al.57 found that FoATP synthase b-chain is involved in white spot syndrome virus infection. Feng et al. found that the ATP synthase b subunit gene is essential to the survival of rice brown planthopper (BPH), and the RNAi of ATP synthase b shows an effective inhibition effect of the BPH. ATP synthase b may serve as a potential target gene for BPH control58. ATP synthase b subunit is essential for normal function of ATP synthase. The gene and protein level of ATP synthase subunit b are downregulated by electromagnetic radiation. This finding indicated that electromagnetic radiation will affect the photosynthesis and energy metabolism of M. aeruginosa, which may reduce the ability of these biological processes.
This experiment is an exploratory experiment that found some response of cyanobacteria to electromagnetic radiation. Previous studies in our laboratory have found that electromagnetic waves have effects on the energy metabolism of Caenorhabditis elegans59,60 and yeast61, which is similar to the results of this experiment. Next, we need to further study the mechanism of electromagnetic radiation's influence on energy metabolism pathway and changes in energy metabolism-related metabolites and so on. Further experiments investigating the altered pathways would be beneficial to reveal a mechanism by which electromagnetic radiation exerts these biological effects. In the future, it is necessary to base on different intensities (energy domain), frequencies (frequency domain), exposure times (time domain), from the level of cells lines, tissues, organs, individuals, colonies, and adopt different methods (appearance characteristics analysis, epigenetic analysis, genomic analysis, cytology atlas, etc.), systematically study the effects of electromagnetic radiation. It will help the scientific community to deeply understand and reveal the target sites, signaling pathway and mechanism of action of electromagnetic exposure on organisms. Future research outlook is shown in Fig. 3. From study the effects of electromagnetic radiation on organisms to electromagnetic dose effects, clarify the mechanism of electromagnetic radiation, and finally establish electromagnetic radiation protection programs, in order to ensure the safe development of the electromagnetic environment.
In conclusion, electromagnetic radiation affects the expression of ribosome-related regulatory genes, which may inhibit the protein synthesis of M. aeruginosa. Electromagnetic radiation affects the expression of Calvin cycle-related regulatory genes, and alters photoreaction system related protein expression levels, thereby disturbing the photosynthesis of M. aeruginosa. Moreover, it influences oxidative phosphorylation, thereby affecting cyanobacterial energy metabolism. Our finding suggests that energy metabolism pathway may respond positively to electromagnetic radiation.
Materials and methods
Experimental materials
The experiment involved species from M. aeruginosa (FACHB-905), and came from the Institute of Aquatic Biology, Chinese Academy of Sciences. The culture medium was BG 11 medium21.
Experimental treatment
M. aeruginosa were cultured in the normal light (Culture temperature was 25 ± 1 °C with 12 L: 12 D light–to-dark ratio, light intensity was 1000 ± 100 lx) for a period. Then, M. aeruginosa suspension were divided into two parts for exposure and control experiments. Based on our previous study result (13,21), a dark condition was chosen for the experiment. The exposure group M. aeruginosa was treated with 1.8 GHz and 40 V/m electromagnetic radiation in the dark for 24 h, temperature was 25 ± 1 °C, whereas the control group was not exposed to electromagnetic radiation and other conditions remained constant.
Electromagnetic radiation was generated using a vector signal generator (AgilentE8267DPSG) and a signal amplifier (AV38701E), and emitted from an antenna (ETS3180B). The antenna was placed at 24 cm above the sample area. The signal at the sample position was measured using an electromagnetic radiation analyzer (PMM8053B, Narta-STS, Italy) and a signal analyzer (AgilentN9030A). M. aeruginosa suspension were exposed to 1.8 GHz radio frequency electromagnetic radiation through a continuous sine wave, and at the position of the M. aeruginosa suspension, the radio frequency electromagnetic field strength was 40 V/m and the temperature was 25 °C. Experimental exposure device is the same as the device in the reference21.
Each experiment was repeated three times.
Transcriptomic analysis
Three samples each from the treatment and control groups were sent to GENE DENOVO Company to extract RNA and Transcriptome sequencing. The ligation products were size selected by agarose gel electrophoresis, PCR amplified, and sequenced using Illumina HiSeqTM 4000 by Gene Denovo Biotechnology Co. (Guangzhou, China).
To get high quality clean reads, reads were further filtered by fastp (version 0.18.0). The edgeR package (version 3.12.1) (http://www.r-project.org/) was used to identify differentially expressed genes (DEGs) across samples or groups. We identified genes with a fold change ≥ 2 and a false discovery rate (FDR) < 0.05 in a comparison as significant DEGs. DEGs were then subjected to enrichment analysis of KEGG pathways (https://www.kegg.jp/kegg/kegg1.html). The calculated p-value has undergone FDR correction, taking FDR ≤ 0.05 as a threshold. Pathways meeting this condition were defined as significantly enriched pathways in DEGs.
Determination of enzymes activity or content
Three samples each from the treatment and control groups were determinated total ATP synthase, H+K+-ATPase activity and ATP content.
Approximately 20 mL of M. aeruginosa suspension was taken, and algal cells were first collected by using a refrigerated centrifuge (Eppendorf 5810R). The cells were centrifuged at 10,000 r·min-1 and 4 °C for 3 min. Second, 1 mL of PBS or 0.9% saline solution and quartz sand were added to the suspension. Subsequently, a high-throughput tissue crusher (Shanghai Jingxin tiss-48) was used to crush the suspension for 6 min. Finally, the suspension was centrifuged at 10,000 r·min-1 for 5 min to obtain a test solution.
Total protein content, total ATP synthase activity, H+K+-ATPase activity, and ATP content were determined by kits (all kits used in the experiment were purchased from Nanjing Jiancheng Bioengineering Institute). Sample determination was performed in accordance with the kits’ instructions. The experiment was repeated three times.
Statistical analysis of the data was performed using SPSS 18 software. The significance analysis was performed using an independent sample t- test, and the data were expressed as mean ± SD). When p < 0.05, a significant difference was present between the control group and the exposed group.
Data availability
All data generated or analysed during this study are included in this published article.
References
Apeldoorn, M. E., Egmond, H. P., Speijers, G. J. A. & Bakker, G. J. I. Toxins of cyanobacteria. Mol. Nutr. Food Res. 51(1), 7–60 (2007).
Liu, Y., Chen, S., Zhang, J. & Gao, B. Growth, microcystin-production and proteomic responses of Microcystis aeruginosa under long-term exposure to amoxicillin. Water Res. 93, 141–152 (2016).
Su, Y. et al. Impact of ageing on the fate of molybdate-zerovalent iron nanohybrid and its subsequent effect on cyanobacteria (Microcystis aeruginosa) growth in aqueous media. Water Res. 140, 135–147 (2018).
Gonzalez-Pleiter, M. et al. Toxicity of five antibiotics and their mixtures towards photosynthetic aquatic organisms: implications for environmental risk assessment. Water Res. 47(6), 2050–2064 (2013).
Pomati, F., Netting, A. G., Calamari, D. & Neilan, B. A. Effects of erythromycin, tetracycline and ibuprofen on the growth of Synechocystis sp. and Lemna minor. Aquat. Toxicol. 67, 387–396 (2004).
Wu, Y., Wang, Y., Du, J., Wang, Z. & Wu, Q. Effects of yttrium under lead stress on growth and physiological characteristics of Microcystis aeruginosa. J. Rare Earths 34(7), 747–756 (2016).
Shi, W. et al. Effects of Mn2+, Cu2+, Zn2+ and Fe3+ concentrations on growth of Alga Aureococcus anophagefferens. Fish. Sci. 38(03), 62–69 (2019).
Gherardini, L., Ciuti, G., Tognarelli, S. & Cinti, C. Searching for the perfect wave: the effect of radiofrequency electromagnetic fields on cells. Int. J. Mol. Sci. 15, 5366–5387 (2014).
Kouzmanova, M., Dimitrova, M., Dragolova, D., Atanasova, G. & Atanasov, N. Alterations in enzyme activities in leaves after exposure of Plectranthus Sp. plants to 900 MHz electromagnetic field. Biotechnol. Biotechnol. Eq. 23, 611–615 (2009).
Vian, A. et al. Microwave irradiation affects gene expression in plants. Plant Signal Behav. 1(2), 67–69 (2006).
Jangid, R. K. et al. Microwave treatment induced mutations and altered gene expression in Vigna aconitifolia. Biol. Plant. 54(4), 703–706 (2010).
Yang, D. & Zhou, X. Effect of specific electromagnetic wave (TDP) on photosynthesis and biomass in Spirulina platensis. J. Southwest Agri. Univ. 10(1), 112–115 (1988).
Tang, C., Yu, H., Yang, C. & Cai, P. The effects of the electromagnetic environment on oxidative stress and photosynthetic carbon fixation of Microcystis aeruginosa. Acta Sci. Circum. 37(8), 3194–3200 (2017).
Xiong, L. et al. Experimental study on algae inhibition effect of square wave pulsed electric field. High Voltage Engineering 42(2), 368–376 (2016).
Li, M., Qu, J. & Peng, Y. Sterilization and algae killing by the application of pulsed magnetic field. Acta Sci. Circum. 24(2), 260–264 (2004).
Zheng, B. The Study on Inhibition Mechanism of Cyanobacteria Growth by Irradiation Treatment. Ph.D. Dissertation, (Nanjing University, 2012).
Chen, S., Liu, Y., Zhang, J. & Gao, B. iTRAQ-based quantitative proteomic analysis of Microcystis aeruginosa exposed to spiramycin at different nutrient levels. Aquat. Toxicol. 185, 193–200 (2017).
Qian, H. et al. Contrasting silver nanoparticle toxicity and detoxification strategies in Microcystis aeruginosa and Chlorella vulgaris: new insights from proteomic and physiological analyses. Sci. Total Environ. 572, 1213–1221 (2016).
Yang, M. & Wang, X. Interactions between Microcystis aeruginosa and coexisting bisphenol A at different phosphorus levels. Sci. Total Environ. 658, 439–448 (2019).
Zuo, Z., Ni, B. & Yang, L. Production of primary metabolites in Microcystis aeruginosa in regulation of nitrogen limitation. Bioresour. Technol. 270, 588–595 (2018).
Tang, C. et al. Electromagnetic radiation disturbed the photosynthesis of Microcystis aeruginosa at the proteomics level. Sci. Rep. 8(1), 479–486 (2018).
Ludwig, W. et al. Complete nucleotide sequences of seven eubacterial genes coding for the elongation factor Tu: functional, structural and phylogenetic evaluations. Arch. Microbiol. 153, 241–247 (1990).
Zhuang, J. Regulation of NirA and Differential Proteomic Study of Microcystis aerugiuosa Cultured on Different Nitrogen Nutrients and Different 2-OG Concentrations. Master Dissertation, (Xiamen University, 2008).
Wang, J., Zhu, S.& Xu, C. Biochemistry, 3rd, ed. (Higher Education Press, Beijing, 2002).
Kudlicki, W., Coffman, A., Kramer, G. & Hardesty, B. Renaturation of rhodanese by translational elongation factor (EF) Tu: protein refolding by EF-Tu flexing. J. Biol. Chem. 272(51), 32206–32210 (1997).
Chen, J. Analysis of Physiological Response and Differentially Expressed Gene of Temperature Stress of Arthrospira. Master Dissertation (Tianjin University of Commerce, 2017).
Battchikova, N., Eisenhut, M. & Aro, E. Cyanobacterial NDH-1 complexes: novel insights and remaining puzzles. Biochim. Biophys. Acta-Bioenerg. 1807, 935–944 (2011).
Zhao, J., Rong, W., Gao, F., Ogawa, T. & Ma, W. Subunit Q Is Required to Stabilize the Large Complex of NADPH Dehydrogenase in Synechocystis sp. Strain PCC 6803. Plant Physiol. 168 (2), 443–451 (2015).
Zhang, J. et al. NdhP is an exclusive subunit of large complex of NADPH dehydrogenase essential to stabilize the complex in Synechocystis sp. strain PCC 6803. J. Biol. Chem. 289 (27), 18770–18781 (2014).
Bernat, G., Appel, J., Ogawa, T. & Rogner, M. Distinct roles of multiple NDH-1 complexes in the cyanobacterial electron transport network as revealed by kinetic analysis of P700+ reduction in various ndh-deficient mutants of Synechocystis sp. Strain PCC6803. J. Bacteriol. 193 (1), 292–295 (2011).
Ogawa, T. A gene homologous to the subunit-2 gene of NADH dehydrogenase is essential to inorganic carbon transport of Synechocystis PCC6803. Proc. Natl. Acad. Sci. 88, 4275–4279 (1991).
Ogawa, T. Identification and characterization of the ictA/ndhL gene product essential to inorganic carbon transport of synechocystic PCC6803. Plant Physiol. 99, 1604–1608 (1992).
Hibino, T. et al. Salt enhances photosystem I content and cyclic electron flow via NAD(P)H dehydrogenase in the halotolerant cyanobacterium Aphanothece halophytica. Aust. J. Plant Physiol 23, 321–330 (1996).
Hihara, Y., Kamei, A., Kanehisa, M., Kaplan, A. & Ikeuchi, M. DNA microarray analysis of cyanobacterial gene expression during acclimation to high light. Plant Cell 13, 793–806 (2001).
Klein, G., Satre, M., Dianoux, A. C. & Vignais, P. V. Radiolabeling of natural adenosine triphosphatase inhibitor with phenyl (14C) isothiocyanate and study of its interaction with mitochondrial adenosine triphosphatase. Localization of inhibitor binding sites and stoichiometry of binding. Biochemistry, 19, 2919–2925 (1980).
Feniouk, B. A. & Yoshida, M. Regulatory mechanisms of proton-translocating FOF1-ATP synthase. Results Probl. Cell Differ. 45, 279–308 (2008).
Sahu, B. B. & Shaw, B. P. Salt-inducible isoform of plasma membrane H+ ATPase gene in rice remains constitutively expressed in natural halophyte, Suaeda maritima. J. Plant Physiol. 166, 1077–1089 (2009).
Bi, H., Dong, X., Wang, M. & Ai, X. Foliar spray calcium and salicylic acid improve the activities and gene expression of photosynthetic enzymes in cucumber seedlings under low light intensity and suboptimal temperature. Acta Horticult. Sin. 42(1), 56–64 (2015).
Qi, X. et al. Cloning, characterization and expression of CsGAPDH, a waterlogging responsive gene in cucumber. Acta Horticult. Sin. 38(9), 1693–1698 (2011).
Michelis, R. & Gepstein, S. Identification and characterization of a heat-induced isoform of aldolase in oat chloroplast. Plant Mol. Biol. 44, 487–498 (2000).
Wei, H. et al. Comparative analysis of expressed sequence tags from cold-acclimated and non-acclimated leaves of Rhododendron catawbiense Michx. Planta 221, 406–416 (2005).
Zhang, X. et al. Cloning of NaCl-induced Dunaliella salina Fructose-1,-6-bisphosphate aldolase cDNA and its expression in tobacco. Sci. China (Ser. C) 32(5), 392–398 (2002).
Lv, K. et al. The methods for measuring the activity of chloroplast fructose-1,6-bisphosphate aldolase in common wheat and its application. J. Triticeae Crops 31(1), 82–87 (2011).
Xiao, H., Chen, C., Xu, Y., Ji, D. & Xie, C. Cloning and expression analysis of the chloroplast fructose-1,6-bisphosphatase gene from Pyropia haitanensis. Acta Oceanol. Sin. 33(4), 92–100 (2014).
Lee, D. et al. A proteomic approach in analyzing heat-responsive proteins in rice leaves. Proteomics 7, 3369–3383 (2007).
Wang, L. Comparative proteomics of Manihot esculenta leaves and chloroplasts under drought stress. Master Dissertation (Hainan University, 2013).
Feng, L. et al. Overexpression of SBPase enhances photosynthesis against high temperature stress in transgenic rice plants. Plant Cell Physiol. 26, 1635–1646 (2007).
Feng, L. et al. Overexpression of sedoheptulose-1,7-bisphosphatase enhances photosynthesis and growth under salt stress in transgenic rice plants. Funct. Plant Biol. 34(9), 822–834 (2007).
Zhao, Y., Du, H., Wang, Z. & Huang, B. Identification of proteins associated with water-deficit tolerance in C4 perennial grass species, Cynodon dactylon × Cynodon transvaalensis and Cynodon dactylon. Physiol. Plant. 141, 40–55 (2011).
Hu, X. et al. Abscisic acid refines the synthesis of chloroplast proteins in maize (Zea mays) in response to drought and light. PLoS ONE 7(11), e49500 (2012).
Müller, K., Doubnerova, V., Synkova, H., Cerovska, N. & Ryslava, H. Regulation of phosphoenolpyruvate carboxylase in PVY(NTN)-infected tobacco plants. Biol. Chem. 390(3), 245–251 (2009).
Ma, F. et al. Dynamic Changes of IsiA-Containing Complexes during Long-Term Iron Deficiency in Synechocystis sp.PCC 6803. Mol. Plant. 10, 143–154 (2017).
Havaux, M. et al. The chlorophyll-binding protein IsiA is inducible by high light and protects the cyanobacterium Synechocystis PCC6803 from photooxidative stress. FEBS Lett. 579, 2289–2293 (2005).
Sandström, S., Park, Y. I., Oquist, G., & Gustafsson, P. CP43', the isiA gene product, functions as an excitation energy dissipator in the cyanobacterium Synechococcus sp. PCC 7942. Photochem. Photobiol. 74 (3), 431–437 (2001).
Chen, Y. The effects of knockdown of ATPsyn-b on spermiogenesis and larval growth in Drosophila. Master Dissertation (Huazhong Normal University, 2014).
Zheng, J. The effects of knockdown of ATPsyn-b and Mst84Db on male fertility in Drosophila. Master Dissertation (Huazhong Normal University, 2013).
Li, Q., Liu, Q. & Huang, J. FOATP synthase b-chain of Litopenaeus vannamei involved in White Spot Syndrome Virus infection. Virus Genes 47, 42–48 (2013).
Feng, Y. et al. Molecular cloning and function analysis of ATP synthase b subunit gene ATPSb in the brown planthopper, Nilaparvata lugens (Hemiptera: Delphacidae). Acta Entomolog Sin 61(5), 519–526 (2018).
Shi, Z. et al. The energy metabolism in Caenorhabditis elegans under the extremely low-frequency electromagnetic field exposure. Sci. Rep. 5, 8471–8481 (2015).
Sun, Y. et al. Coupling of oxidative stress responses to tricarboxylic acid cycle and prostaglandin E2 alterations in Caenorhabditis elegans under extremely low-frequency electromagnetic field. Int. J. Radiat. Biol. 94(12), 1159–1166 (2018).
Lin, K., Yang, C., Lian, H. & Cai, P. Exposure of ELF-EMF and RF-EMF increase the rate of glucose transport and TCA cycle in budding yeast. Front. Microbiol. 7, 1378–1392 (2016).
Acknowledgements
This work was supported by Focus Deploy Project of Chinese Academy of Sciences (Grant No. KGFZD-135-160-024), Xiamen Science and Technology Plans Project (Grant No. 3502Z20162021), Science and Technology Innovation Fund Project, Chinese Academy of Sciences (Grant No. CXJJ-16M115), and Xiamen Key Laboratory of Physical Environment.
Author information
Authors and Affiliations
Contributions
Conceived and designed the experiments: P. C., C. T. Performed the experiments: C. T., S. T., Z. Z. Wrote the paper: C. T. Analyzed data and drew pictures: C. T., P. C. All authors reviewed the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Tang, C., Zhang, Z., Tian, S. et al. Transcriptomic responses of Microcystis aeruginosa under electromagnetic radiation exposure. Sci Rep 11, 2123 (2021). https://doi.org/10.1038/s41598-020-80830-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-020-80830-z
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.