Abstract
Fungi respond to antifungal drugs by increasing their antioxidant stress response. How this impacts antifungal efficacy remains controversial and not well understood. Here we examine the role of catalase activity in the resistance of Saccharomyces cerevisiae to the common antifungals, fluconazole and miconazole, for which we report minimum inhibitory concentrations (MICs) of 104 and 19 μM, respectively. At sub-MIC concentrations, fluconazole and miconazole stimulate catalase activity 2-3-fold but, unexpectedly, deletion of cytosolic catalase (ctt1) makes cells more resistant to these azoles and to clotrimazole, itraconazole and posaconazole. On the other hand, upregulating Ctt1 activity by preconditioning with 0.2 mM H2O2 potentiates miconazole 32-fold and fluconazole 4-fold. Since H2O2 preconditioning does not alter the resistance of ctt1Δ cells, which possess negligible catalase activity, we link azole potentiation with Ctt1 upregulation. In contrast, sod2Δ cells deleted for mitochondrial superoxide dismutase are 4–8-fold more azole sensitive than wild-type cells, revealing that Sod2 activity protects cells against azole toxicity. In fact, the ctt1Δ mutant has double the Sod2 activity of wild-type cells so ctt1 deletion increases azole resistance in part by Sod2 upregulation. Notably, deletion of peroxisomal/mitochondrial cta1 or cytosolic sod1 does not alter fluconazole or miconazole potency.
Similar content being viewed by others
Introduction
Antimicrobial challenge appears to induce the rewiring of microbial metabolic networks and stress-response pathways regardless of the primary drug-target interaction1,2. Drug lethality increases when major stress responses are disrupted3,4,5,6,7,8,9,10,11 whereas resistance increases in fungi12,13,14 and bacteria3,4,7,9,15,16,17 when antioxidant defenses are boosted. Susceptibility to antimicrobial killing decreases when cells are treated with antioxidants1,2,13. Moreover, deletion of respiratory enzymes or inhibition of cellular respiration, a major source of reactive oxygen species (ROS)1,2,18, decreases antimicrobial lethality. Combined, these observations are consistent with the belief that cidal antibiotics1,19 and antifungals2,20,21 increase ROS levels. Hence, understanding the roles of ROS-metabolizing enzymes in antimicrobial efficacy is of critical importance in treating infection.
Several studies have examined the contribution of key ROS-metabolizing enzymes such as catalases and superoxide dismutases (Sods) to bacterial survival following challenge with antibiotics. For example, the ΔrelA ΔspoT mutant of Pseudomonas aeruginosa, which is deficient in the (p)ppGpp alarmone, exhibits depressed catalase and superoxide dismutase (Sod) activities and is hypersensitive to antibiotics3,4,22. This can be reversed by overexpression in the mutant of KatA, the dominant P. aeruginosa catalase4, or by restoration of Sod activity22. Many other bacteria become more susceptible to antibiotics on Sod deletion, including Enterococcus faecalis9,23,24, Campylobacter jejuni8, Acinetobacter baumanii7, Staphylococcus aureus24 and E. coli in stationary-phase25 but maybe not in exponentially growing cultures15. Deletion of the catalase-peroxidase katG or the alkyl hydroperoxide reductase ahpC also potentiates some antibiotics in E. coli15.
Antioxidant enzymes also are associated with antifungal potency. For example, sirtuin Hst1 deletion increases catalase activity and lowers multidrug sensitivity in Candida glabrata5. Deletion of membrane-associated CuSod4 and CuSod526 (Fig. 1) or inhibition of Cu-dependent Sod activity in Candida albicans increases the anitbiofilm activity of miconazole13 and amphotericin B27. Fluconazole induces a number of genes responsive to oxidative- and nitrosative-stress in C. albicans28 and both fluconazole- and amphotericin B-resistant C. albicans and Candida dubliniensis exhibit increased catalase and Sod activities12.
Subcellular localization of catalases and superoxide dismutases (SODs) in S. cerevisiae and C. albicans. Cytosolic Ctt1 is found in the cytoplasm of S. cerevisiae, whereas Cta1 is co-targeted to peroxisomes and mitochondria in respiring S. cerevisiae48 and inferred to be associated with these two compartments in C. albicans. CuZnSod1 is localized in the cytoplasm and the mitochondrial intermembrane space of both yeasts, while MnSod2 is present in the mitochondrial matrix. C. albicans possesses an extra MnSod3 in the cytosol and cell-membrane-associated CuSod4-6, which are absent in S. cerevisiae. Note that C. glabrata possess only Cta15, CuZnSod1 and MnSod274.
Previously, it was shown that exposure to a fungistatic dose of miconazole induces catalase activity in both C. albicans and Saccharomyces cerevisiae14. Notably, the catalase and superoxide dismutase isoforms present in these yeasts differ considerably (Fig. 1). C. albicans possesses a single peroxisomal/mitochondrial catalase (Cta1)29 together with six Sods13 while S. cerevisiae produces cytosolic Ctt1 in addition to Cta1 but just two Sods, cytosolic CuZnSod1, which also localizes to the mitochondrial intermembrane space30,31, and mitochondrial MnSod232 (for clarity, we indicate Sod metal dependence throughout the text). Thus, a comparison of how deletion of specific antioxidant enzymes alters antifungal potency in these well-characterized yeasts provides an excellent opportunity to gain new insights into pathogen survival strategies and the evolution of antifungal resilience.
In this work, we focus on the role of catalase activity in the response of S. cerevisiae (strain BY4741; Table 1) to challenge with common antifungal azoles. The primary target of these drugs is ergosterol biosynthesis33, a sterol found in the cell membrane of fungi. Specifically, we report on the azole resistance of single ctt1 and cta1 knockouts (Table 1) as well as on wild-type cells preconditioned with a low dose of H2O2 to stimulate catalase activity34,35. Furthermore, since MnSod2 is induced by the H2O2 stress response17,36,37, we also monitored the Sod activity and azole sensitivity of sod1 and sod2 mutants (Table 1) with and without catalase inhibition. Combined, our unprecedented results shed new light on antioxidant defense and azole resistance in S. cerevisiae, which itself is an emerging opportunistic pathogen38,39,40,41.
Results
MICs of azoles for S. cerevisiae and their classification as fungicidal vs. fungistatic
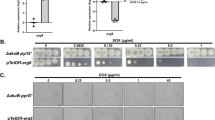
Starting at an initial cell density of 106 cfu/ml and based on cell growth at different drug concentrations (Fig. 2), we determined the minimum inhibitory concentration (MIC µg/mL; µM) for our S. cerevisiae strain (BY4741) of six medically relevant azoles: itraconazole (32; 45), fluconazole (32; 105), posaconazole (32; 46), voriconazole (>256; >730), miconazole (8; 19) and clotrimazole (4; 12) (Table S1). The structures of the azoles, shown as a footnote to Table S1, reveal that the drugs examined can be classified as triazoles (itraconazole, fluconazole, posaconazole and voriconazole) and imidazoles (miconazole and clotrimazole). The imidazoles are more potent antifungals than the triazoles and, in fact, cells are refractory to voriconazole (Table S1). An azole is classified as fungicidal if 1xMIC or 2xMIC promotes a ≥103-fold reduction in the viable cfu/mL and Table S2 shows that the imidazoles are fungicidal under the present experimental conditions, whereas the triazoles are fungistatic with the exception of voriconazole.
Minimum inhibitory concentration (MIC) of fluconazole and miconazole for wild-type S. cerevisiae cells. Wild-type cells grown to OD600 0.50 (12 h) in YPD at a medium-to-flask ratio of 1:5 at 30 °C/225 rpm were diluted to OD600 0.15 (106 cfu/mL) before challenge with increasing azole concentration in a 96-well plate. OD600 values were read in the plate reader at time t = 0, and cells were incubated at 30 °C without stirring. OD600 values were read again in the plate reader at t = 24 h, and ΔOD600 (24–0 h) values were plotted vs. (A) Fluconazole and (B) miconazole concentration to give MIC of 32 and 8 μg/mL, respectively. The results represent the avg ± SEM of six independent replicates (n = 6).
Cultures of C. albicans (strain SC5314) at the same initial cell density (106 cfu/mL) exhibit MICs of >1 mM for fluconazole42 and 60 µM for miconazole43. Thus, under our culture conditions, S. cerevisiae strain BY4741 is more sensitive to the azoles than C. albicans strain SC531442,43. We note that our initial cell density is higher than that used in standardized methods of antifungal susceptibility testing recommended by the Clinical and Laboratory Standards Institute (103 cfu/mL)44 to provide sufficient cells for the biochemical analyses. At high cell density, the tolerance for the azoles is likely higher than under standard testing conditions42. However, relative MICs are of interest here and in the further studies described here, fluconazole and miconazole represent the fungistatic triazoles and fungicidal imidazoles, respectively.
Fluconazole and miconazole at sub-MIC increase catalase activity and H2O2 levels in S. cerevisiae
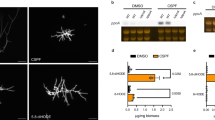
Both azoles increase catalase activity in wild-type S. cerevisiae (Fig. 3A,B)14. However, a 10-fold lower concentration of miconazole (~1 μM; 0.40 μg/mL) vs. fluconazole (~10 μM; 3.2 μg/mL) doubles the catalase activity of wild-type cells as measured at 24 h after azole addition (Fig. 3A,B). Notably, catalase activity declines when cells are treated with increasing miconazole concentrations (Fig. 3B), which concurs with the report that fungicidal doses deplete catalase activity in S. cerevisiae and C. albicans14.
Azoles and H2O2 stimulate catalase activity in wild-type and cta1Δ cells but not in ctt1Δ cells. Total catalase activity was measured without (A,B) and with (D,E) H2O2 preconditioning of wild-type  , cta1Δ
, cta1Δ  and ctt1Δ cells
and ctt1Δ cells  at 24 h after challenge of 3-mL cultures at 106 cfu/mL with (A,D) fluconazole and (B,E) miconazole at concentrations below their minimum inhibitory concentrations (MICs; see Table 2). Relative intracellular H2O2 levels measured by flow cytometry at (C) 8 h and (F) 24 h after DHR-stained wild-type
at 24 h after challenge of 3-mL cultures at 106 cfu/mL with (A,D) fluconazole and (B,E) miconazole at concentrations below their minimum inhibitory concentrations (MICs; see Table 2). Relative intracellular H2O2 levels measured by flow cytometry at (C) 8 h and (F) 24 h after DHR-stained wild-type  , cta1Δ
, cta1Δ  and ctt1Δ cells
and ctt1Δ cells  were exposed to 0.05xMIC miconazole or ethanol only (control). Experimental conditions: Cells at an initial OD600 of 0.15 were grown in YPD at a medium-to-flask ratio of 1:5 at 30 °C/225 rpm. Catalase activity was assayed (see Materials and Methods) at 24 h after 3-mL cultures were challenged with azole in 12 μL of ethanol. For preconditioning, cultures were grown to OD600 0.50 (12 h), 0.2 mM H2O2 was added to the medium, cells were diluted 30 min later to OD600 0.15 (106 cfu/mL) and challenged with azole in 3-mL cultures. To determine relative H2O2 levels, cells grown in 3-mL cultures at initial OD600 of 0.15 were was stained with 5 µM DHR in 1-mL aliquots at 30 °C, pelleted after 120 min, diluted to 106 cells/mL in PBS, fixed with 2% formalin (v/v) and analyzed by flow cytometry (see Materials and Methods). Relative fluorescence units (RFU; ex/em 490/520 nm) of individual cells were measured and the median RFU of 10,000 cells estimates a sample’s relative H2O2 level. All results represent the avg ± SEM of six independent experiments (n = 6). Statistical analyses performed using Student’s t-test compare each sample with the wild-type untreated control. *p < 0.05 and **p < 0.01.
were exposed to 0.05xMIC miconazole or ethanol only (control). Experimental conditions: Cells at an initial OD600 of 0.15 were grown in YPD at a medium-to-flask ratio of 1:5 at 30 °C/225 rpm. Catalase activity was assayed (see Materials and Methods) at 24 h after 3-mL cultures were challenged with azole in 12 μL of ethanol. For preconditioning, cultures were grown to OD600 0.50 (12 h), 0.2 mM H2O2 was added to the medium, cells were diluted 30 min later to OD600 0.15 (106 cfu/mL) and challenged with azole in 3-mL cultures. To determine relative H2O2 levels, cells grown in 3-mL cultures at initial OD600 of 0.15 were was stained with 5 µM DHR in 1-mL aliquots at 30 °C, pelleted after 120 min, diluted to 106 cells/mL in PBS, fixed with 2% formalin (v/v) and analyzed by flow cytometry (see Materials and Methods). Relative fluorescence units (RFU; ex/em 490/520 nm) of individual cells were measured and the median RFU of 10,000 cells estimates a sample’s relative H2O2 level. All results represent the avg ± SEM of six independent experiments (n = 6). Statistical analyses performed using Student’s t-test compare each sample with the wild-type untreated control. *p < 0.05 and **p < 0.01.
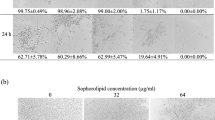
Fungicidal miconazole also induces more ROS formation than fungistatic fluconazole in wild-type S. cerevisiae2, C. albicans2 and C. glabrata45. To examine the rise in intracellular H2O2 induced by the fungicide, we stained cells with the profluorescent dye DHR, which is preferentially oxidized by H2O2 in the presence of cellular catalysts46. At 8 and 24 h after challenge of wild-type cells with 0.4 μg/mL miconazole, we observe a 1.5- and 3-fold increase in probe fluorescence, respectively, compared to untreated control cells (Fig. 3C,F). Hence, a high level of H2O2 accumulates over time in miconazole-treated wild-type cells, affirming that the fungicidal azole increases ROS in yeast2,20.
Ctt1 catalase activity weakly combats miconazole-induced H2O2 accumulation in S. cerevisiae cells
Miconazole induces a rise in intracellular H2O2 (Fig. 3C,F) despite also inducing catalase activity in wild-type cells. This led us to examine catalase activity and H2O2 levels in the cta1Δ and ctt1Δ strains, which lack peroxisomal/mitochondrial and cytosolic catalase, respectively (Fig. 1). Catalase activity (Fig. 3A,B) and H2O2 levels (Fig. 3C) are the same in wild-type and cta1Δ cells, which reflects the strong repression of Cta1 by glucose in the medium34,47,48. In contrast, Ctt1 is not repressed by glucose34,49,50 and confers most of the catalase activity in cells grown in YPD since ctt1Δ cells are virtually devoid of catalase activity (Fig. 3A,B). Moreover, the azoles fail to induce catalase activity in the ctt1Δ strain (Fig. 3A,B) although ~4 times more H2O2 accumulates in miconazole-treated vs. control over 24 h (Fig. 3F). It also is remarkable that the H2O2 levels in wild-type and cta1Δ cells are ~75% those of ctt1Δ cells with negligible catalase activity (Fig. 3C,F). Thus, Ctt1 appears to be an ineffective scavenger of miconazole-induced H2O2.
Deletion of ctt1 or inhibition of catalase activity increases azole resistance in S. cerevisiae
Peroxide-metabolizing enzymes have been associated with protection against cidal antimicrobials4,5,8,14,15,45. However, our observation that Ctt1 does not inhibit miconazole-induced H2O2 accumulation (Fig. 3C,F) led us to ask whether Ctt1 actually protects cells against azole toxicity. As shown in Table 2 and Fig. S1, ctt1Δ cells display 4- and 8-fold higher MICs for fluconazole and miconazole, respectively, than the two strains with Ctt1 activity (Tables 2, S1). Given this surprising observation, we additionally determined the fold-change in MIC when ctt1 was deleted for the four other azoles. Both wild-type and ctt1Δ cells are refractory to voriconazole (Table S1) but the ctt1Δ strain is 8-fold less sensitive to posaconazole and 2-fold less sensitive to clotrimazole and itraconazole than wild-type cells (Table S1). Thus, Ctt1 appears to potentiate both fungistatic and fungicidal azoles in S. cerevisiae.
Aminotriazole is a well-documented inhibitor of catalase activity in S. cerevisiae51. Thus, to directly probe the effect of inhibition of catalase activity on miconazole resistance we added aminotriazole to the cells. This compound did not inhibit the growth of any strain at concentrations as high as 100 mM (data not shown) but treatment with 25 mM lowers catalase activity to undetectable levels in wild-type cells (Fig. S5). In the presence of 25 mM aminotriazole, both wild-type and ctt1Δ cells have the same miconazole MIC (32 μg/mL; Table 2), which links miconazole potentiation in wild-type cells with Ctt1 catalase activity.
H2O2 preconditioning stimulates Ctt1 catalase activity and lowers the fluconazole and miconazole resistance of S. cerevisiae cells
Prompted by the link between miconazole potentiation and catalase activity, we questioned whether stimulating this activity before azole addition would further sensitize cells to the drug. As we previously reported34, preconditioning wild-type or cta1Δ cells with a low dose of H2O2 (e.g., 0.2 mM) in YPD medium doubles their Ctt1 activity (Fig. 3D,E vs. 3A,B; no azole). The combination of H2O2 preconditioning and azole challenge (0.5xMIC fluconazole or 0.05xMIC miconazole) increases Ctt1 activity by 4–6-fold above basal levels in wild-type and cta1Δ cells (Fig. 3D,E vs. A,B) and increases their azole sensitivity 4–32-fold (Table 2). In contrast, H2O2 preconditioning has little or no effect on the catalase activity (which remains barely detectable; Fig. 3) or azole sensitivity of the ctt1Δ strain (Table 2), affirming that azole potentiation is linked to increased Ctt1 activity and is not augmented by H2O2 exposure. Moreover, H2O2 preconditioning potentiates miconazole significantly more than fluconazole (Table 2) presumably because the fungicide is the more potent stimulator of Ctt1 activity (Fig. 3B,E vs. 3A,D).
Deletion of ctt1 elevates MnSod2 activity in early log phase and increases miconazole resistance
Although fungicide-dependent ROS production reportedly leads to fungal cell death2,13,20, we find no link here between elevated H2O2 levels and miconazole sensitivity. In fact, ctt1Δ cells, which are the most miconazole resistant (Table 2), accumulate more H2O2 on challenge with this azole (Fig. 3C,F). However, the miconazole resistance of C. albicans biofilms is dependent on the ROS-detoxifying activity of Sods22, and we37 and others36 have shown previously that suppressing or deleting catalase activity in S. cerevisiae upregulates mitochondrial MnSod2. Thus, we hypothesized that increased MnSod2 activity contributes to the enhanced azole resistance of ctt1Δ cells (Table 2). There are two Sod isoforms in S. cerevisiae (Fig. 1), and we find that the three strains exhibit similar total Sod activity, which doubles between 8 and 24 h but does not increase upon miconazole challenge (Fig. 4A,D). Since MnSod2 accounts for only 10–20% of the total Sod activity in cells growing on glucose52, to unmask any variation in this activity, we selectively inhibited CuZnSod1 with KCN53. This revealed 1.7-fold higher MnSod2 activity in untreated ctt1Δ cells vs. wild-type or cta1Δ cells (Fig. 4B,E).
MnSod2 activity is higher and O2•− levels lower in ctt1Δ cells vs. wild-type and cta1Δ cells. (A,D) Total Sod activity (CuZnSod1 plus MnSod2), (B,E) MnSod2 activity, and (C,F) relative intracellular O2•− levels measured by flow cytometry with DHE staining at (C) 8 h and (F) 24 h after wild-type  , cta1Δ
, cta1Δ  and ctt1Δ cells
and ctt1Δ cells  (see Fig. 2) at 8 h (top panels) and 24 h (bottom panels) after the addition of 0.05xMIC (0.4 μg/mL) miconazole (+miconazole) or ethanol vehicle (control) to the cultures. The Sod activity assay is described under Materials and methods. KCN (5 mM) was added to selectively inhibit CuZnSod2 activity in (B,E). To determine relative O2•− levels, cells were grown in 3 mL cultures at an initial OD600 of 0.15, and 1 mL of culture was stained with 5 µM DHE at 30 °C, pelleted after 60 min, diluted to 106 cells/mL in PBS, fixed with 2% formalin (v/v) and analyzed using flow cytometry (see Materials and Methods). Relative fluorescence units (RFU; ex/em 490/580 nm) of individual cells were measured and the median RFU of 10,000 cells estimates a sample’s relative O2•− levels. Results represent the avg ± SEM of six independent experiments (n = 6). Statistical analyses performed using Student’s t-test compare each sample with the wild-type untreated control. **p < 0.01.
(see Fig. 2) at 8 h (top panels) and 24 h (bottom panels) after the addition of 0.05xMIC (0.4 μg/mL) miconazole (+miconazole) or ethanol vehicle (control) to the cultures. The Sod activity assay is described under Materials and methods. KCN (5 mM) was added to selectively inhibit CuZnSod2 activity in (B,E). To determine relative O2•− levels, cells were grown in 3 mL cultures at an initial OD600 of 0.15, and 1 mL of culture was stained with 5 µM DHE at 30 °C, pelleted after 60 min, diluted to 106 cells/mL in PBS, fixed with 2% formalin (v/v) and analyzed using flow cytometry (see Materials and Methods). Relative fluorescence units (RFU; ex/em 490/580 nm) of individual cells were measured and the median RFU of 10,000 cells estimates a sample’s relative O2•− levels. Results represent the avg ± SEM of six independent experiments (n = 6). Statistical analyses performed using Student’s t-test compare each sample with the wild-type untreated control. **p < 0.01.
We next compared the relative levels of O2•− in the three strains. Staining cells with the profluorescent dye, DHE, which is preferentially oxidized by O2•− 54, we uncovered 2-fold less O2•− in the ctt1Δ strain relative to wild-type or cta1Δ cells (Fig. 4C). O2•− levels were a factor of ~1.3 higher in the cultures challenged with miconazole but remained significantly lower in ctt1Δ cells (Fig. 4C). The O2•− levels tripled between 8 and 24 h such that the 24-h miconazole-challenged cells contained > 10-fold more O2•− than the untreated 8-h cells (Fig. 4F vs. 4C). Also, the 24-h cultures have comparable O2•− levels and MnSod2 activity (Fig. 4E,F) so azole resistance must be associated with the O2•–detoxifying activity of MnSod2 during exponential growth. Thus, we conclude that ctt1Δ cells are more azole resistant (Table 2) because they possess the higher MnSod2 activity in early log phase (Fig. 4B,C).
We additionally examined if the Sod mimetics, TEMPO• or mito-TEMPO•, protect wild-type S. cerevisiae against miconazole toxicity. These radicals are well-established O2•− scavengers and mito-TEMPO• is targeted to mitochondria55 but not TEMPO• 55,56. Addition of 1 mM TEMPO• did not change the MIC of miconazole (Table 3) and 1 mM mito-TEMPO• afforded only modest protection, doubling the MIC of miconazole in four of the six independent cultures examined (Table 3). It is possible that O2•− scavenging by mito-TEMPO• is offset by the miconazole-induced increase in respiration in wild-type cells (Fig. S6). Increased respiration is not detected in the ctt1Δ mutant (Fig. S6) so its elevated MnSod2 activity may provide better protection against miconazole-dependent O2•− production than MnSod2 plus mito-TEMPO• in wild-type cells. Also, the efficacy of mito-TEMPO• may be lowered by its reaction with mitochondrial reductases55.
S. cerevisiae cells deleted for sod2Δ exhibit decreased fluconazole and miconazole resistance
To further explore the importance of Sod activity in azole resistance, we measured the fluconazole and miconazole MICs for sod1Δ and sod2Δ cells. MICs are the same for sod1Δ and wild-type cells, revealing that CuZnSod1 deletion does not impact miconazole resistance (Table 2), which is consistent with 1 mM TEMPO• having no protective effect (Table 3). However, the sod2Δ strain possesses fluconazole and miconazole MICs that are 4- and 8-fold lower, respectively (Table 2). These results confirm that MnSod2 activity protects cells from azole toxicity and upregulation of MnSod2 activity in the ctt1Δ strain (Fig. 4B,E) increases its azole resistance (Table 2).
Inhibiting catalase activity in the sod2Δ strain enhances miconazole resistance less than in wild-type cells
If Ctt1 activity potentiates the azoles by suppressing MnSod2, then inhibiting catalase activity in the sod2Δ strain should not enhance resistance. Treatment of sod2Δ cells with 25 mM aminotriazole resulted in undetectable catalase activity as seen for wild-type cells (Fig. S5). The MIC for miconazole increased from 1 to 4 µg/mL vs. the increase to 32 µg/mL seen on aminotriazole treatment of wild-type cells (Table 2). Hence, Ctt1 activity potentiates miconazole in large part by depressing MnSod2 activity or in other words, the O2•− detoxifying activity of MnSod2 combats azole toxicity and its deletion or suppression by Ctt1 activity lowers azole resistance in S. cerevisiae.
Discussion
Cytosolic Ctt1 catalase activity, not elevated intracellular H2O2, potentiates azole toxicity
De Nollin et al. found that fungistatic doses of miconazole stimulate catalase activity in S. cerevisiae14 and proposed that this rescues cells from H2O2 intoxication. We report here that sub-MIC concentrations of miconazole induce Ctt1 catalase activity up to 3-fold in our wild-type S. cerevisiae strain (BY4741) (Fig. 3) but this does prevent cells from accumulating ~4-fold more H2O2 over 24 h than untreated cells (Fig. 3F). Furthermore, ctt1Δ cells with negligible catalase activity, accumulate more miconazole-induced H2O2 than wild-type or cta1Δ cells (Fig. 3) but are more resistant to the azole (Table 2). Therefore, contrary to expectation14, our results reveal that azole-induced H2O2 production does not alter cell viability. In fact, cytosolic Ctt1 ineffectively combats H2O2 accumulation in wild-type and cta1Δ cells, and the increased azole sensitivity of H2O2-preconditioned wild-type cells results from Ctt1 upregulation and not exposure to exogenous H2O2. In sum, the azoles potentiate their own toxicity by induction of Ctt1 and not H2O2.
Cytosolic Ctt1 activity potentiates the azoles partly by depressing MnSod2 activity
Since ctt1Δ cells with the highest MnSod2 activity of the strains examined here (Fig. 4) are 4–8-fold more azole resistant than wild-type cells, we conclude that depression of MnSod2 activity on Ctt1 stimulation potentiates the azoles. MnSod2 is not essential for fermenting S. cerevisiae57 but sod2Δ cells exhibit 4–8-fold lower azole resistance than wild-type cells (Table 2). However, we note that H2O2 preconditioned wild-type cells are 4-fold more miconazole sensitive than sod2Δ cells (Table 2). Thus, strong Ctt1 induction may potentiate miconazole by additional mechanisms. For example, miconazole may bind to the heme of Ctt1 as reported for CYP5158, the 14α-demethylase in the ergosterol biosynthetic pathway33. This could promote heme-catalyzed azole autoxidation with the formation of reactive, cytotoxic species via mechanisms analogous to those we reported for hydrazides59.
Elevated MnSod2 activity in early log phase increases azole resistance
The ctt1Δ cells from 8-h cultures possess higher MnSod2 activity and less O2•− than wild-type and cta1Δ cells (Fig. 4). Respiration is a major source of O2•−, and induction of respiration by miconazole reportedly increases its toxicity in S. cerevisiae whereas genetic blockage of respiration (by deleting TCA-cycle and ETC components) has the opposite effect2. Respiration-derived O2•− inactivates aconitase60,61 with the release of free iron, which catalyzes the production of highly toxic hydroxyl radicals via Fenton chemistry1,37,62.
We have previously reported on the positive biochemical and physiological effects of elevated MnSod2 activity in young cells deleted for cytochrome c peroxidase (ccp1Δ)48. Like ctt1Δ, the ccp1Δ mutant exhibits low O2•− and high H2O2 levels plus it possesses stable aconitase activity, accumulates low amounts of free iron and hydroxyl radicals, amasses mitochondrial damage more slowly and lives longer than wild-type cells48. These traits arise from the beneficial mitochondrial H2O2 stress response known as mitohormesis, which requires MnSod2 upregulation17,36,37,63. Presumably, the advantages of elevated MnSod2 activity in early log phase contribute to the increased miconazole resistance of ctt1Δ cells. At 24 h after miconazole treatment, the three strains possess comparable MnSod2 activity and O2•− levels (Fig. 4). Nonetheless, based on Rhod123 staining64, miconazole does not increase respiration in ctt1Δ cells (Fig. S6), suggesting that mitohormesis protects mitochondrial function65.
Catalase and azole resistance in S. cerevisiae vs. C. albicans and C. glabrata
Given their different catalase and Sod isozymes (Fig. 1), it is informative to compare azole sensitivity in S. cerevisiae and C. albicans. It was reported in the 1970s that fungistatic doses of miconazole induce catalase activity in S. cerevisiae and C. albicans whereas fungicidal doses inhibit this activity14. We confirm these results for S. cerevisiae but show that only Ctt1 activity is induced (Fig. 3B) since peroxisomal/mitochondrial Cta1 is repressed by glucose66. Cta1 is the only catalase isoform in C. albicans29 (Fig. 1), and synergistic killing of C. albicans biofilms by fluconazole and H2O2 has been reported but no molecular mechanism was suggested67.
Like S. cerevisiae, the opportunistic yeast C. glabrata possesses Cta1, cytosolic CuZnSod1 and mitochondrial MnSod2. Antifungals also induce ROS production and stimulate catalase, Sod and glutathione peroxidase activities in C. glabrata21,45. Azole resistance is associated with increased catalase activity5 and increased protein levels of thiol peroxidases68, but whether deletion of these antioxidant enzymes alters azole resistance in C. glabrata remains to be seen.
Conclusions
Although high catalase activity has been linked to azole resistance in C. albicans, C. glabrata and S. cerevisiae, the present study reveals that azole-induced upregulation of Ctt1 activity potentiates azole toxicity by depressing MnSod2 activity in S. cerevisiae, Hence, MnSod2 is an interesting antifungal target in this yeast but target antioxidant enzymes are likely to be species dependent. Therefore, to expand our knowledge of the role of a given antioxidant activity in fungal survival strategies, we need to establish the potency of antifungal drugs in yeasts singly deleted for the antioxidant enzyme of interest as performed here for S. cerevisiae.
Materials and Methods
Reagents
Suppliers of chemicals/biochemical were as follows: Peptone, yeast extract, microbiological agar, phenylmethylsulfonyl (PMSF), tetramethylethylenediamine (TEMED), tris(hydroxymethyl)aminomethane (Tris, electrophoresis grade 99%), glycine and sodium chloride (Bioshop); glucose and 30% H2O2 (v/v) (Fisher Scientific); buffer salts, 3-amino-1,2,4-triazole (aminotriazole or ATZ), 2,2,6,6-tetramethyl-1-piperidinyloxyl (TEMPO•) and 2-(2,2,6,6-tetramethyl-1-piperidinyloxyl-4-ylamino)-2-oxoethyl triphenylphosphonium chloride (Mito-TEMPO•) (Sigma); fluconazole and miconazole nitrate (Santa Cruz Biotechnology); itraconazole, clotrimazole, posaconazole and voriconazole (Cayman Chemicals). The Bradford reagent and other electrophoresis reagents were obtained from Biorad.
Yeast strains
The Saccharomyces cerevisiae wild-type and mutant BY4741 strains used in this work are listed in Table 1. The wild-type strain was purchased from the EUROSCARF. The cta1Δ, ctt1Δ, sod1Δ and sod2Δ mutant strains are derived from the Yeast Deletion Project69,70 and were kindly provided by Professor Christopher Brett (Department of Biology, Concordia University).
Growth conditions and H2O2 preconditioning
Precultures (10 mL) were obtained by growing single colonies of each strain in YPD (1% yeast extract, 2% peptone and 2% dextrose) for 24 h at 30 °C with high aeration (medium-to-flask ratio of 1:5 and shaking at 225 rpm). These cultures were used to inoculate 25 mL of fresh YPD in 125-mL flasks to give the experimental cultures at an initial OD600 of 0.01 (OD600 was measured at a 1.0-cm pathlength unless otherwise indicated). Cells (3 mL) were grown under the same conditions to mid-log phase (OD600 0.50; 12 h) and preconditioned with 0.2 mM H2O2 for 30 min at 30 °C/225 rpm where indicated.
Determination of azole minimum inhibitory concentration (MIC)
The solid azoles were dissolved in 100% ethanol to give stocks of 50 mg/mL fluconazole, 10 mg/mL voriconazole, 1 mg/mL miconazole and clotrimazole; and in 100% dimethyl sulfoxide (DMSO) to give stocks of 1 mg/mL posaconazole and itraconazole. Since H2O2 preconditioning causes a 25–30% reduction in viable ctt1Δ cells34, the liquid cultures were diluted to OD600 0.15 (106 cfu/mL) in fresh YPD before MIC determination. Our initial cell density is higher than suggested by the Clinical and Laboratory Standards Institute (103 cfu/m)44 to provide sufficient cells for the biochemical analyses. Cells were exposed to different azole concentrations in 96-well plates (final volume of 200 μL per well) and MICs were determined as described44. Briefly, cells were mixed with the drug and OD600 was measured on a SpectraFluor Plus Tecan plate reader at t = 0 and t = 24 h after growth at 30 °C without shaking. The MIC for each azole was determined from a plot of OD600 at t = 24 h minus that t = 0 vs. [azole]. The MIC is the lowest antifungal concentration that results in no detectable growth after 24 h incubation44. MICs for cultures simultaneously treated with the azole and 1 mM TEMPO•, 1 mM mito-TEMPO• (Sod mimetics)55,56 or 25 mM aminotriazole (catalase inhibitor)51 were determined the same way in 96-well plates. To establish if an azole was fungicidal or fungistatic, wells containing 1xMIC and 2xMIC of the drug were serially diluted 10x after 24 h at 30 °C, plated onto YPD agar and grown for 2 days at 30 °C to measure the viable cfu/mL. A drug was considered fungicidal if 1xMIC or 2xMIC promoted a ≥103-fold reduction in viable cfu44.
Soluble protein extracts
Cells ±H2O2 preconditioning and ± aminotriazole exposure were diluted to OD600 0.15 in 3 mL of fresh YPD ± azole in a 15-mL Falcon tube, grown at 30 °C/225 rpm for 24 h, OD600 values were measured, and soluble proteins were extracted as described previously17,34. Briefly, after centrifugation at 2000 × g, cells were washed 2x with 100 mM potassium phosphate buffer at pH 7.0 (KPi) containing 0.1 mM PMSF, the pellets were diluted into KPi/PMSF, and mixed with an equal volume of acid-washed glass beads (400-600 µm). Cells were disrupted by vortexing 4 × 15 s, the homogenates were spun at 13000 × g for 10 min at 4 °C, and the total protein concentration in the supernatants was determined by the Bradford assay with BSA as a standard71.
Catalase and Sod activity assays
Cells exposed to azole concentrations below the MIC (sub-MIC) were used in the biochemical analyses to avoid the general metabolic collapse and down regulation of multiple enzyme activities seen at lethal drug concentrations14,34. To assay for catalase activity, 25–150 µL aliquots of soluble protein extract72 containing 20–100 μg protein were added to 1.0 mL of 20 mM H2O2 in 50 mM KPi in a cuvette. H2O2 decomposition was monitored at 240 nm (ε240 = 43.6 M−1 cm−1)72. One unit of catalase activity catalyzes the degradation of 1 µmol of H2O2 per min34,72. Sod activity was assayed using the Superoxide Dismutase Detection Kit (Cell Technologies, CSOD100), where O2•− is generated by xanthine/xanthine oxidase and oxidized by XTT (2,3-bis(2-methoxy-4-nitro-5-sulfophenyl)-2H-tetrazolium-5-carboxanilide)73. One unit of Sod activity inhibits the rate of XTT reduction by O2•− by 50% and was assayed according to the manufacturer’s instructions using 2–10 µg of total protein in 96-well plates. To determine MnSod2 activity only, lysates at 2 mg/mL total soluble protein were preincubated with 5 mM KCN for 30 min at room temperature to fully inhibit CuZnSod1 prior to assaying 5–50 µg of total protein for Sod activity.
Relative ROS levels
Relative levels were estimated as we described before17,37 using the fluorescent probes, dihydrorhodamine 123 (DHR) for H2O246 and dihydroethidine (DHE) for O2•− 54. Cultures were diluted to OD600 0.15 in 3 mL of fresh YPD in a 15-mL Falcon tube, vehicle (ethanol) or 0.05xMIC (0.4 μg/mL or ~1 μM) miconazole was added, cells were incubated at 30 °C/225 rpm for 8 or 24 h, harvested at 2000 × g for 10 min, washed once and resuspended in PBS (10 mM NaPi and 150 mM NaCl, pH 7.0) to a final density of 107 cells/mL. One mL of suspension was stained with 5 µM DHR or 5 µM DHE at 30 °C for 120 and 60 min, respectively, the cells were pelleted, diluted to 106 cells/mL in PBS, fixed with 2% formalin (v/v) and analyzed by flow cytometry (BD Accuri C6, BD Biosciences). The fluorescence from individual cells was measured and expressed as relative fluorescence units (RFU). Relative H2O2 and O2•− levels are estimated from the median RFU of 10,000 cells for each sample.
Statistical analyses
These were performed using the two-tailed Student’s t-test calculated using Graph Pad Prism 7 software. The analyses compare each sample with the wild-type untreated control (see figure legends). Probabilities < 5% are considered significant (p < 0.05).
References
Kohanski, M. A., Dwyer, D. J., Hayete, B., Lawrence, C. A. & Collins, J. J. A common mechanism of cellular death induced by bactericidal antibiotics. Cell 130, 797–810 (2007).
Belenky, P., Camacho, D. & Collins, J. J. Fungicidal drugs induce a common oxidative-damage cellular death pathway. Cell Rep. 3, 350–358 (2013).
Nguyen, D. et al. Active starvation responses mediate antibiotic tolerance in biofilms and nutrient-limited bacteria. Science (80-.). 334, 982–986 (2011).
Khakimova, M., Ahlgren, H. G., Harrison, J. J., English, A. M. & Nguyen, D. The stringent response controls catalases in Pseudomonas aeruginosa and is required for hydrogen peroxide and antibiotic tolerance. J. Bacteriol. 195, 2011–2020 (2013).
Orta-Zavalza, E. et al. Local silencing controls the oxidative stress response and the multidrug resistance in Candida glabrata. Mol. Microbiol. 88, 1135–1148 (2013).
Poole, K. Bacterial stress responses as determinants of antimicrobial resistance. J. Antimicrob. Chemother. 67, 2069–2089 (2012).
Heindorf, M., Kadari, M., Heider, C., Skiebe, E. & Wilharm, G. Impact of Acinetobacter baumannii superoxide dismutase on motility, virulence, oxidative stress resistance and susceptibility to antibiotics. PLoS One 9, e101033, https://doi.org/10.1371/journal.pone.0101033 (2014).
Hwang, S., Ryu, S. & Jeon, B. Roles of the superoxide dismutase SodB and the catalase KatA in the antibiotic resistance of Campylobacter jejuni. J. Antibiot. (Tokyo). 66, 351–353 (2013).
Bizzini, A., Zhao, C., Auffray, Y. & Hartke, A. The Enterococcus faecalis superoxide dismutase is essential for its tolerance to vancomycin and penicillin. J. Antimicrob. Chemother. 64, 1196–1202 (2009).
Dörr, T., Lewis, K. & Vulić, M. SOS Response Induces Persistence to Fluoroquinolones in Escherichia coli. PLoS Genet. 5, e1000760, https://doi.org/10.1371/journal.pgen.1000760 (2009).
Djorić, D. & Kristich, C. J. Oxidative stress enhances cephalosporin resistance of Enterococcus faecalis through activation of a two-component signaling system. Antimicrob. Agents Chemother. 59, 159–169 (2015).
Blanco Linares, C. E. et al. Fluconazole and amphotericin-B resistance are associated with increased catalase and superoxide dismutase activity in Candida albicans and Candida dubliniensis. Rev. Soc. Bras. Med. Trop. 46, 752–758 (2013).
Bink, A. et al. Superoxide dismutases are involved in Candida albicans biofilm persistence against miconazole. Antimicrob. Agents Chemother. 55, 4033–4037 (2011).
de Nollin, S., Van Belle, H., Goossens, F., Thone, F. & Borgers, M. Cytochemical and biochemical studies of yeasts after in vitro exposure to miconazole. Antimicrob. Agents Chemother. 11, 500–513 (1977).
Wang, X. & Zhao, X. Contribution of oxidative damage to antimicrobial lethality. Antimicrob. Agents Chemother. 53, 1395–1402 (2009).
Britigan, B. E. et al. Antioxidant enzyme expression in clinical isolates of Pseudomonas aeruginosa: identification of an atypical form of manganese superoxide dismutase. Infect. Immun. 69, 7396–7401 (2001).
Martins, D., Kathiresan, M. & English, A. M. Cytochrome c peroxidase is a mitochondrial heme-based H2O2 sensor that modulates antioxidant defense. Free Radic. Biol. Med. 65, 541–551 (2013).
Yan, L. et al. The alternative oxidase of Candida albicans causes reduced fluconazole susceptibility. J. Antimicrob. Chemother. 64, 764–773 (2009).
Dwyer, D. J., Kohanski, M. A., Hayete, B. & Collins, J. J. Gyrase inhibitors induce an oxidative damage cellular death pathway in Escherichia coli. Mol. Syst. Biol. 3, 91, https://doi.org/10.1038/msb4100135 (2007).
Kobayashi, D. et al. Endogenous reactive oxygen species is an important mediator of miconazole antifungal effect. Antimicrob. Agents Chemother. 46, 3113–3117 (2002).
Mesa-Arango, A. C. et al. The production of reactive oxygen species is a universal action mechanism of Amphotericin B against pathogenic yeasts and contributes to the fungicidal effect of this drug. Antimicrob. Agents Chemother. 58, 6627–6638 (2014).
Martins, D. et al. Superoxide dismutase activity confers (p)ppGpp-mediated antibiotic tolerance to stationary-phase Pseudomonas aeruginosa. Proc. Natl. Acad. Sci. USA 115, 9797–9802 (2018).
Ladjouzi, R. et al. Loss of Antibiotic Tolerance in Sod-Deficient Mutants Is Dependent on the Energy Source and Arginine Catabolism in Enterococci. J. Bacteriol. 197, 3283–3293 (2015).
Ladjouzi, R. et al. Analysis of the tolerance of pathogenic enterococci and Staphylococcus aureus to cell wall active antibiotics. J. Antimicrob. Chemother. 68, 2083–2091 (2013).
Wang, J. H. et al. Sigma S-dependent antioxidant defense protects stationary-phase Escherichia coli against the bactericidal antibiotic gentamicin. Antimicrob. Agents Chemother. 58, 5964–5975 (2014).
Martchenko, M., Alarco, A. M., Harcus, D. & Whiteway, M. Superoxide dismutases in Candida albicans: transcriptional regulation and functional characterization of the hyphal-induced SOD5 gene. Mol. Biol. Cell. 15, 456–467 (2004).
De Brucker, K., Bink, A., Meert, E., Cammue, B. P. & Thevissen, K. Potentiation of antibiofilm activity of amphotericin B by superoxide dismutase inhibition. Oxid. Med. Cell. Longev. 2013, 704654, https://doi.org/10.1155/2013/704654 (2013).
Arana, D. M., Nombela, C. & Pla, J. Fluconazole at subinhibitory concentrations induces the oxidative- and nitrosative-responsive genes TRR1, GRE2 and YHB1, and enhances the resistance of Candida albicans to phagocytes. J. Antimicrob. Chemother. 65, 54–62 (2009).
Wysong, D. R., Christin, L., Sugar, A. M., Robbins, P. W. & Diamond, R. D. Cloning and sequencing of a Candida albicans catalase gene and effects of disruption of this gene. Infect. Immun. 66, 1953–1961 (1998).
Okado-Matsumoto, A. & Fridovich, I. Subcellular distribution of superoxide dismutases (SOD) in rat liver. Cu,Zn-SOD in mitochondria. J. Biol. Chem. 276, 38388–38393 (2001).
Klöppel, C., Michels, C., Zimmer, J., Herrmann, J. M. & Riemer, J. In yeast redistribution of Sod1 to the mitochondrial intermembrane space provides protection against respiration derived oxidative stress. Biochem. Biophys. Res. Commun. 403, 114–119 (2010).
Ghaemmaghami, S. et al. Global analysis of protein expression in yeast. Nature 425, 737–741 (2003).
Lupetti, A., Danesi, R., Campa, M., Tacca, M. D. & Kelly, S. Molecular basis of resistance to azole antifungals. Trends Mol. Med. 8, 76–81 (2002).
Martins, D. & English, A. M. Catalase activity is stimulated by H2O2 in rich culture medium and is required for H2O2 resistance and adaptation in yeast. Redox Biol. 2, 308–313 (2014).
Izawa, S., Inoue, Y. & Kimura, A. Importance of catalase in the adaptive response to hydrogen peroxide: analysis of acatalasaemic Saccharomyces cerevisiae. Biochem. J. 320, 61–67 (1996).
Mesquita, A. et al. Caloric restriction or catalase inactivation extends yeast chronological lifespan by inducing H2O2 and superoxide dismutase activity. Proc. Natl. Acad. Sci. USA 107, 15123–15128 (2010).
Martins, D., Titorenko, V. I. & English, A. M. Cells with impaired mitochondrial H2O2 sensing generate less •OH radicals and live longer. Antioxid. Redox Signal. 21, 1490–1503 (2014).
Muñoz, P. et al. Saccharomyces cerevisiae fungemia: an emerging infectious disease. Clin. Infect. Dis. 40, 1625–1634 (2005).
Chitasombat, M. N. et al. Rare opportunistic (non-Candida, non-Cryptococcus) yeast bloodstream infections in patients with cancer. J. Infect. 64, 68–75 (2012).
Pfaller, M. A. & Diekema, D. J. Epidemiology of invasive mycoses in North America. Crit. Rev. Microbiol. 36, 1–53 (2010).
Enache-Angoulvant, A. & Hennequin, C. Invasive Saccharomyces infection: a comprehensive review. Clin. Infect. Dis. 41, 1559–1568 (2005).
Perumal, P., Mekala, S. & Chaffin, W. L. Role for Cell Density in Antifungal Drug Resistance in Candida albicans biofilms. Antimicrob. Agents Chemother. 51, 2454–2463 (2007).
Thevissen, K. et al. Miconazole Induces Changes in Actin Cytoskeleton prior to Reactive Oxygen Species Induction in Yeast. J. Biol. Chem. 282, 21592–21597 (2007).
C L S I. Reference Method for Broth Dilution antifungal susceptibility testing of yeasts, 3rd edition, Approved Standard M27-A3. (Clinical Laboratory Standards Institute, Wayne, PA, 2008).
Mahl, C. D. et al. Induction of ROS generation by fluconazole in Candida glabrata: Activation of antioxidant enzymes and oxidative DNA damage. Diagn. Microbiol. Infect. Dis. 82, 203–208 (2015).
Royall, J. A. & Ischiropoulos, H. Evaluation of 2′,7′-dichlorofluorescin and dihydrorhodamine 123 as fluorescent probes for intracellular H2O2 in cultured endothelial cells. Arch. Biochem. Biophys. 302, 348–355 (1993).
Cross, H. S. & Ruis, H. Regulation of catalase synthesis in Saccharomyces cerevisiae by carbon catabolite repression. Mol. Gen. Genet. 166, 37–43 (1978).
Petrova, V. Y., Drescher, D., Kujumdzieva, A. V. & Schmitt, M. J. Dual targeting of yeast catalase A to peroxisomes and mitochondria. Biochem. J. 380, 393–400 (2004).
Bissinger, P. H., Wieser, R., Hamilton, B. & Ruis, H. Control of Saccharomyces cerevisiae catalase T gene (CTT1) expression by nutrient supply via the RAS-cyclic AMP pathway. Mol. Biol. Cell. 9, 1309–1315 (1989).
Godon, C. et al. The H2O2 stimulon in Saccharomyces cerevisiae. J. Biol. Chem. 273, 22480–22489 (1998).
Bayliak, M., Gospodaryov, D., Semchyshyn, H. & Lushchak, V. Inhibition of catalase by aminotriazole in vivo results in reduction of glucose-6-phosphate dehydrogenase activity in Saccharomyces cerevisiae cells. Biochem. 73, 420–426 (2008).
Jamieson, D. J., Rivers, S. L. & Stephen, D. W. S. Analysis of Saccharomyces cerevisiae proteins induced by peroxide and superoxide stress. Microbiology 140, 3277–3283 (1994).
Weydert, C. J. & Cullen, J. J. Measurement of superoxide dismutase, catalase and glutathione peroxidase in cultured cells and tissue. Nat. Protoc. 5, 51–66 (2010).
Nazarewicz, R. R., Bikineyeva, A. & Dikalov, S. I. Rapid and specific measurements of superoxide using fluorescence spectroscopy. J. Biomol. Screen. 18, 498–503 (2013).
Trnka, J., Blaikie, F. H., Smith, R. A. J. & Murphy, M. P. A mitochondria-targeted nitroxide is reduced to its hydroxylamine by ubiquinol in mitochondria. Free Radic. Biol. Med. 44, 1406–1419 (2008).
Du, K., Farhood, A. & Jaeschke, H. Mitochondria-targeted antioxidant Mito-Tempo protects against acetaminophen hepatotoxicity. Arch. Toxicol. 91, 761–773 (2017).
Longo, V. D., Liou, L. L., Valentine, J. S. & Gralla, E. B. Mitochondrial superoxide decreases yeast survival in stationary phase. Arch. Biochem. Biophys. 365, 131–142 (1999).
Warrilow, A. G., Parker, J. E., Kelly, D. E. & Kelly, S. L. Azole Affinity of Sterol 14-Demethylase (CYP51) Enzymes from Candida albicans and Homo sapiens. Antimicrob. Agents Chemother. 57, 1352–1360 (2013).
Aitken, S. M., Ouellet, M., Percival, M. D. & English, A. M. Mechanism of horseradish peroxidase inactivation by benzhydrazide: a critical evaluation of arylhydrazides as peroxidase inhibitors. Biochem. J. 621, 613–621 (2003).
Murakami, K. & Yoshino, M. Inactivation of aconitase in yeast exposed to oxidative stress. Biochem. Mol. Biol. Int. 41, 481–486 (1997).
Longo, V. D., Viola, K. L., Klein, W. L. & Finch, C. E. Reversible inactivation of superoxide-sensitive aconitase in Abeta1-42-treated neuronal cell lines. J. Neurochem. 75, 1977–1985 (2000).
Keyer, K. & Imlay, J. A. Superoxide accelerates DNA damage by elevating free-iron levels. Proc. Natl. Acad. Sci. USA 93, 13635–13640 (1996).
Ristow, M. & Zarse, K. How increased oxidative stress promotes longevity and metabolic health: The concept of mitochondrial hormesis (mitohormesis). Exp. Gerontol. 45, 410–418 (2010).
Baracca, A., Sgarbi, G., Solaini, G. & Lenaz, G. Rhodamine 123 as a probe of mitochondrial membrane potential: evaluation of proton flux through F0 during ATP synthesis. Biochim. Biophys. Acta - Bioenerg. 1606, 137–146 (2003).
Pan, Y. Mitochondria, reactive oxygen species, and chronological aging: A message from yeast. Exp. Gerontol. 46, 847–852 (2011).
Simon, M., Adam, G., Rapatz, W., Spevak, W. & Ruis, H. The Saccharomyces cerevisiae ADR1 gene is a positive regulator of transcription of genes encoding peroxisomal proteins. Mol. Cell. Biol. 11, 699–704 (1991).
Nett, J. E., Guite, K. M., Ringeisen, A., Holoyda, K. A. & Andes, D. R. Reduced biocide susceptibility in Candida albicans biofilms. Antimicrob. Agents Chemother. 52, 3411–3413 (2008).
Rogers, P. D., Vermitsky, J. P., Edlind, T. D. & Hilliard, G. M. Proteomic analysis of experimentally induced azole resistance in Candida glabrata. J. Antimicrob. Chemother. 58, 434–438 (2006).
Giaever, G. et al. Functional profiling of the Saccharomyces cerevisiae genome. Nature 418, 387–391 (2002).
Winzeler, E. A. et al. Functional characterization of the S. cerevisiae genome by gene deletion and parallel analysis. Science 285, 901–906 (1999).
Bradford, M. M. A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein dye binding. Anal. Biochem. 72, 248–254 (1976).
Beers, R. F. & Sizer, I. W. A spectrophotometric method for measuring the breakdown of hydrogen peroxide by catalase. J. Biol. Chem. 195, 133–140 (1952).
Ukeda, H., Maeda, S., Ishii, T. & Sawamura, M. Spectrophotometric Assay for Superoxide Dismutase Based on Tetrazolium Salt 3′-{1-[(Phenylamino)-carbonyl]-3,4-tetrazolium}-bis(4-methoxy-6-nitro)benzenesulfonic Acid Hydrate Reduction by Xanthine–Xanthine Oxidase. Anal. Biochem. 251, 206–209 (1997).
Briones-Martin-Del-Campo, M. et al. The superoxide dismutases of Candida glabrata protect against oxidative damage and are required for lysine biosynthesis, DNA integrity and chronological life survival. Microbiol. (United Kingdom) 161, 300–310 (2015).
Acknowledgements
We thank Dr. Cristopher Brett (Concordia University, Canada) for kindly providing us with the sod1∆ and sod2∆ strains. This work was performed with funding from the Natural Sciences and Engineering Research Council of Canada (NSERC) and Concordia University. A.M.E holds an Honorary Concordia University Research Chair. D.M. acknowledges a postdoctoral scholarship from the NSERC CREATE Training Program in Bionanomachines and additional awards from Concordia University.
Author information
Authors and Affiliations
Contributions
D.M. and A.M.E. designed research; D.M. performed the experimental work. D.N. contributed new reagents and analytical tools; D.M. and A.M.E. analyzed the data; and D.M., A.M.E., and D.N. wrote the paper.
Corresponding author
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
41598_2019_45070_MOESM1_ESM.pdf
Ctt1 catalase activity potentiates antifungal azoles in the emerging opportunistic pathogen <i>Saccharomyces cerevisiae</i>
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Martins, D., Nguyen, D. & English, A.M. Ctt1 catalase activity potentiates antifungal azoles in the emerging opportunistic pathogen Saccharomyces cerevisiae. Sci Rep 9, 9185 (2019). https://doi.org/10.1038/s41598-019-45070-w
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-019-45070-w
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.