Abstract
The fungus Candida albicans frequently colonizes the human gastrointestinal tract, from which it can disseminate to cause systemic disease. This polymorphic species can transition between growing as single-celled yeast and as multicellular hyphae to adapt to its environment. The current dogma of C. albicans commensalism is that the yeast form is optimal for gut colonization, whereas hyphal cells are detrimental to colonization but critical for virulence1,2,3. Here, we reveal that this paradigm does not apply to multi-kingdom communities in which a complex interplay between fungal morphology and bacteria dictates C. albicans fitness. Thus, whereas yeast-locked cells outcompete wild-type cells when gut bacteria are absent or depleted by antibiotics, hyphae-competent wild-type cells outcompete yeast-locked cells in hosts with replete bacterial populations. This increased fitness of wild-type cells involves the production of hyphal-specific factors including the toxin candidalysin4,5, which promotes the establishment of colonization. At later time points, adaptive immunity is engaged, and intestinal immunoglobulin A preferentially selects against hyphal cells1,6. Hyphal morphotypes are thus under both positive and negative selective pressures in the gut. Our study further shows that candidalysin has a direct inhibitory effect on bacterial species, including limiting their metabolic output. We therefore propose that C. albicans has evolved hyphal-specific factors, including candidalysin, to better compete with bacterial species in the intestinal niche.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
The 16S sequencing data are publicly available at the NCBI Sequence Read Archive (SRA) and can be accessed under BioProject PRJNA1008281. Microarray data for analysis of C. albicans expression are available at ArrayExpress under accession MTAB-13349. Source data are provided with this paper.
References
Ost, K. S. et al. Adaptive immunity induces mutualism between commensal eukaryotes. Nature 596, 114–118 (2021).
Tso, G. H. W. et al. Experimental evolution of a fungal pathogen into a gut symbiont. Science 362, 589–595 (2018).
Witchley, J. N. et al. Candida albicans morphogenesis programs control the balance between gut commensalism and invasive infection. Cell Host Microbe 25, 432–443.e436 (2019).
Moyes, D. L. et al. Candidalysin is a fungal peptide toxin critical for mucosal infection. Nature 532, 64–68 (2016).
Li, X. V. et al. Immune regulation by fungal strain diversity in inflammatory bowel disease. Nature 603, 672–678 (2022).
Doron, I. et al. Mycobiota-induced IgA antibodies regulate fungal commensalism in the gut and are dysregulated in Crohn’s disease. Nat Microbiol 6, 1493–1504 (2021).
Rao, C. et al. Multi-kingdom ecological drivers of microbiota assembly in preterm infants. Nature 591, 633–638 (2021).
Iliev, I. D. & Cadwell, K. Effects of intestinal fungi and viruses on immune responses and inflammatory bowel diseases. Gastroenterology 160, 1050–1066 (2021).
Swidergall, M. & LeibundGut-Landmann, S. Immunosurveillance of Candida albicans commensalism by the adaptive immune system. Mucosal Immunol. 15, 829–836 (2022).
Shao, T. Y., Haslam, D. B., Bennett, R. J. & Way, S. S. Friendly fungi: symbiosis with commensal Candida albicans. Trends Immunol. 43, 706–717 (2022).
Li, Q. et al. Dysbiosis of gut fungal microbiota is associated with mucosal inflammation in Crohn’s disease. J. Clin. Gastroenterol. 48, 513–523 (2014).
Sokol, H. et al. Fungal microbiota dysbiosis in IBD. Gut 66, 1039–1048 (2017).
Bacher, P. et al. Human anti-fungal Th17 immunity and pathology rely on cross-reactivity against Candida albicans. Cell 176, 1340–1355.e1315 (2019).
Shao, T. Y. et al. Commensal Candida albicans positively calibrates systemic Th17 immunological responses. Cell Host Microbe 25, 404–417.e406 (2019).
Yeung, F. et al. Altered immunity of laboratory mice in the natural environment is associated with fungal colonization. Cell Host Microbe 27, 809–822.e806 (2020).
Zhai, B. et al. High-resolution mycobiota analysis reveals dynamic intestinal translocation preceding invasive candidiasis. Nat. Med. 26, 59–64 (2020).
Pappas, P. G., Lionakis, M. S., Arendrup, M. C., Ostrosky-Zeichner, L. & Kullberg, B. J. Invasive candidiasis. Nat. Rev. Dis. Primers 4, 18026 (2018).
Koh, A. Y., Kohler, J. R., Coggshall, K. T., Van Rooijen, N. & Pier, G. B. Mucosal damage and neutropenia are required for Candida albicans dissemination. PLoS Pathog. 4, e35 (2008).
Noble, S. M., Gianetti, B. A. & Witchley, J. N. Candida albicans cell-type switching and functional plasticity in the mammalian host. Nat. Rev. Microbiol. 15, 96–108 (2017).
Kadosh, D. Morphogenesis in Candida albicans: Cellular and Molecular Biology (ed. Prasad, R.) 41–62 (Springer, 2017).
Saville, S. P., Lazzell, A. L., Monteagudo, C. & Lopez-Ribot, J. L. Engineered control of cell morphology in vivo reveals distinct roles for yeast and filamentous forms of Candida albicans during infection. Eukaryot. Cell 2, 1053–1060 (2003).
Lo, H. J. et al. Nonfilamentous C. albicans mutants are avirulent. Cell 90, 939–949 (1997).
Carlisle, P. L. et al. Expression levels of a filament-specific transcriptional regulator are sufficient to determine Candida albicans morphology and virulence. Proc. Natl Acad. Sci. USA 106, 599–604 (2009).
Bohm, L. et al. The yeast form of the fungus Candida albicans promotes persistence in the gut of gnotobiotic mice. PLoS Pathog. 13, e1006699 (2017).
Liang, S. H. et al. Hemizygosity enables a mutational transition governing fungal virulence and commensalism. Cell Host Microbe 25, 418–431.e416 (2019).
Mogavero, S. et al. Candidalysin delivery to the invasion pocket is critical for host epithelial damage induced by Candida albicans. Cell Microbiol. 23, e13378 (2021).
Naglik, J. R., Gaffen, S. L. & Hube, B. Candidalysin: discovery and function in Candida albicans infections. Curr. Opin. Microbiol. 52, 100–109 (2019).
Stoldt, V. R., Sonneborn, A., Leuker, C. E. & Ernst, J. F. Efg1p, an essential regulator of morphogenesis of the human pathogen Candida albicans, is a member of a conserved class of bHLH proteins regulating morphogenetic processes in fungi. EMBO J. 16, 1982–1991 (1997).
Braun, B. R. & Johnson, A. D. TUP1, CPH1 and EFG1 make independent contributions to filamentation in Candida albicans. Genetics 155, 57–67 (2000).
Wakade, R. S., Huang, M., Mitchell, A. P., Wellington, M. & Krysan, D. J. Intravital imaging of Candida albicans identifies differential in vitro and in vivo filamentation phenotypes for transcription factor deletion mutants. mSphere 6, e0043621 (2021).
Fan, D. et al. Activation of HIF-1α and LL-37 by commensal bacteria inhibits Candida albicans colonization. Nat. Med. 21, 808–814 (2015).
Yamaguchi, N. et al. Gastric colonization of Candida albicans differs in mice fed commercial and purified diets. J. Nutr. 135, 109–115 (2005).
McDonough, L. D. et al. Candida albicans isolates 529L and CHN1 exhibit stable colonization of the murine gastrointestinal tract. mBio 12, e0287821 (2021).
Braun, B. R., Kadosh, D. & Johnson, A. D. NRG1, a repressor of filamentous growth in C. albicans, is down-regulated during filament induction. EMBO J. 20, 4753–4761 (2001).
Murad, A. M. et al. NRG1 represses yeast-hypha morphogenesis and hypha-specific gene expression in Candida albicans. EMBO J. 20, 4742–4752 (2001).
Wakade, R. S., Kramara, J., Wellington, M. & Krysan, D. J. Candida albicans filamentation does not require the cAMP–PKA pathway in vivo. mBio 13, e0085122 (2022).
Vautier, S. et al. Candida albicans colonization and dissemination from the murine gastrointestinal tract: the influence of morphology and Th17 immunity. Cell Microbiol 17, 445–450 (2015).
Miller, B. M., Liou, M. J., Lee, J. Y. & Baumler, A. J. The longitudinal and cross-sectional heterogeneity of the intestinal microbiota. Curr. Opin. Microbiol. 63, 221–230 (2021).
Brugiroux, S. et al. Genome-guided design of a defined mouse microbiota that confers colonization resistance against Salmonella enterica serovar Typhimurium. Nat Microbiol 2, 16215 (2016).
Trexler, P. C., & Orcutt, R.P. Chapter Sixteen: Development of Gnotobiotics and Contamination Control in Laboratory Animal Science. In: 50 Years of Laboratory Animal Science. Memphis, TN: Am Assoc Lab Anim Sci, 121–128 (2000).
Caballero, S. et al. Cooperating commensals restore colonization resistance to vancomycin-resistant Enterococcus faecium. Cell Host Microbe 21, 592–602.e594 (2017).
Dambuza, I. M. & Brown, G. D. Managing the mycobiota with IgA. Nat. Microbiol. 6, 1471–1472 (2021).
Kasper, L. et al. The fungal peptide toxin candidalysin activates the NLRP3 inflammasome and causes cytolysis in mononuclear phagocytes. Nat. Commun. 9, 4260 (2018).
Swidergall, M. et al. Candidalysin is required for neutrophil recruitment and virulence during systemic Candida albicans infection. J. Infect. Dis. 220, 1477–1488 (2019).
White, S. J. et al. Self-regulation of Candida albicans population size during GI colonization. PLoS Pathog. 3, e184 (2007).
Hoyer, L. L., Payne, T. L., Bell, M., Myers, A. M. & Scherer, S. Candida albicans ALS3 and insights into the nature of the ALS gene family. Curr. Genet. 33, 451–459 (1998).
Phan, Q. T. et al. Als3 is a Candida albicans invasin that binds to cadherins and induces endocytosis by host cells. PLoS Biol. 5, e64 (2007).
Almeida, R. S. et al. The hyphal-associated adhesin and invasin Als3 of Candida albicans mediates iron acquisition from host ferritin. PLoS Pathog. 4, e1000217 (2008).
Martchenko, M., Alarco, A. M., Harcus, D. & Whiteway, M. Superoxide dismutases in Candida albicans: transcriptional regulation and functional characterization of the hyphal-induced SOD5 gene. Mol. Biol. Cell 15, 456–467 (2004).
Fradin, C. et al. Granulocytes govern the transcriptional response, morphology and proliferation of Candida albicans in human blood. Mol. Microbiol. 56, 397–415 (2005).
Hube, B. Fungal adaptation to the host environment. Curr. Opin. Microbiol. 12, 347–349 (2009).
Dewhirst, F. E. et al. Phylogeny of the defined murine microbiota: altered Schaedler flora. Appl. Environ. Microbiol. 65, 3287–3292 (1999).
Guthrie, C. & Fink, G. R. Guide to Yeast Genetics and Molecular Biology (Academic Press, 1991).
Liu, H., Kohler, J. & Fink, G. R. Suppression of hyphal formation in Candida albicans by mutation of a STE12 homolog. Science 266, 1723–1726 (1994).
Park, S. O., Frazer, C. & Bennett, R. J. An adjuvant-based approach enables the use of dominant HYG and KAN selectable markers in Candida albicans. mSphere 7, e0034722 (2022).
Reuss, O., Vik, A., Kolter, R. & Morschhauser, J. The SAT1 flipper, an optimized tool for gene disruption in Candida albicans. Gene 341, 119–127 (2004).
Noble, S. M. & Johnson, A. D. Strains and strategies for large-scale gene deletion studies of the diploid human fungal pathogen Candida albicans. Eukaryot. Cell 4, 298–309 (2005).
Mancera, E. et al. Genetic modification of closely related Candida species. Front. Microbiol. 10, 357 (2019).
Gerami-Nejad, M., Zacchi, L. F., McClellan, M., Matter, K. & Berman, J. Shuttle vectors for facile gap repair cloning and integration into a neutral locus in Candida albicans. Microbiology 159, 565–579 (2013).
Hollomon, J. M. et al. The Candida albicans Cdk8-dependent phosphoproteome reveals repression of hyphal growth through a Flo8-dependent pathway. PLoS Genet. 18, e1009622 (2022).
Dallari, S. et al. Enteric viruses evoke broad host immune responses resembling those elicited by the bacterial microbiome. Cell Host Microbe 29, 1014–1029.e1018 (2021).
Thompson, L. R. et al. A communal catalogue reveals Earth’s multiscale microbial diversity. Nature 551, 457–463 (2017).
Callahan, B. J. et al. DADA2: high-resolution sample inference from Illumina amplicon data. Nat. Methods 13, 581–583 (2016).
Bolyen, E. et al. Reproducible, interactive, scalable and extensible microbiome data science using QIIME 2. Nat. Biotechnol. 37, 852–857 (2019).
Katoh, K., Misawa, K., Kuma, K. & Miyata, T. MAFFT: a novel method for rapid multiple sequence alignment based on fast Fourier transform. Nucleic Acids Res. 30, 3059–3066 (2002).
Price, M. N., Dehal, P. S. & Arkin, A. P. FastTree 2-approximately maximum-likelihood trees for large alignments. PLoS ONE 5, e9490 (2010).
Bokulich, N. A. et al. q2-longitudinal: longitudinal and paired-sample analyses of microbiome data. mSystems 3, e00219–18 (2018).
Quast, C. et al. The SILVA ribosomal RNA gene database project: improved data processing and web-based tools. Nucleic Acids Res. 41, D590–D596 (2013).
Bray, J. R. & Curtis, J. T. An ordination of the upland forest communities of southern Wisconsin. Ecol. Monogr. 27, 325–349 (1957).
Bokulich, N. A. et al. Optimizing taxonomic classification of marker-gene amplicon sequences with QIIME 2’s q2-feature-classifier plugin. Microbiome 6, 90 (2018).
Lozupone, C. & Knight, R. UniFrac: a new phylogenetic method for comparing microbial communities. Appl. Environ. Microbiol. 71, 8228–8235 (2005).
Benjaminii, Y. & Hichberg, Y. Controlling the false discovery rate: a practical and powerful approach to multiple testing. J. R. Stat. Soc. Ser. B 57, 289–300 (1995).
Afgan, E. et al. The Galaxy platform for accessible, reproducible and collaborative biomedical analyses: 2018 update. Nucleic Acids Res. 46, W537–W544 (2018).
Segata, N. et al. Metagenomic biomarker discovery and explanation. Genome Biol. 12, R60 (2011).
Mogavero, S. & Hube, B. Candida albicans interaction with oral epithelial cells: adhesion, invasion, and damage assays. Methods Mol. Biol. 2260, 133–143 (2021).
Gerwien, F. et al. A novel hybrid iron regulation network combines features from pathogenic and nonpathogenic yeasts. mBio 7, e01782-16 (2016).
Ramirez-Zavala, B. et al. The Snf1-activating kinase Sak1 is a key regulator of metabolic adaptation and in vivo fitness of Candida albicans. Mol. Microbiol. 104, 989–1007 (2017).
Miramon, P. et al. A family of glutathione peroxidases contributes to oxidative stress resistance in Candida albicans. Med. Mycol. 52, 223–239 (2014).
Mallick, H. et al. Multivariable association discovery in population-scale meta-omics studies. PLoS Comput. Biol. 17, e1009442 (2021).
Acknowledgements
This work was supported by NIH grants AI166869, AI141893, AI081704 and AI168222 to R.J.B. Y.-H.C. was supported by the Charles H. Revson Senior Fellowship in Biomedical Science. B.H. and T.B.S. were supported by the German Research Foundation (Deutsche Forschungsgemeinschaft (DFG)) within the Cluster of Excellence ‘Balance of the Microverse’, under Germany’s Excellence Strategy, EXC 2051, project ID 390713860. B.H. and S.A. were further supported by the DFG, project HU 528/20-1. J.C.P was supported by AI175081, P.B. was supported by DK125382 and S.P. was supported by a Graduate Research Fellowship from the NSF under award number 1644760. I.V.E. was supported by the Institut Pasteur and is a CIFAR Azrieli Global Scholar in the CIFAR Program Fungal Kingdom: Threats & Opportunities. We thank E. Pamer, I. Jacobsen and T. Hohl for sharing strains and the National Gnotobiotic Rodent Resource Center (NIH grants P40 OD010995 and P30 DK034987) for support.
Author information
Authors and Affiliations
Contributions
R.J.B. conceived the majority of the experiments and wrote the initial manuscript draft, with input from S.-H.L. and S.S. S.-H.L. and S.S. performed the majority of the experiments. P.K. performed the microscopic analysis of yeast and hyphal cells. L.D.M. constructed strains and analysed in vivo phenotypes. J.D. performed analyses of C. albicans strains with individual bacterial strains in mice. C.F. assisted with design and construction of C. albicans mutant strains. S.V. provided gnotobiotic mice and advised on the project. J.C.P., K.C. and Y.-H.C. performed consortia experiments. S.P. and P.B. analysed 16S data and generated related figures. T.B.S., S.A., M.H., S.M. and O.E. performed analyses of C. albicans with candidalysin. O.E. performed in vitro C. albicans competition experiments. S.A. and S.M. performed transcriptional profiling of the ece1 mutant. B.H. conceived the in vitro experiments, provided the C. albicans ece1 mutant strains and advised on the project. I.V.E. performed in vivo experiments and advised on the project.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature thanks the anonymous reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 Comparison of the fitness of WT and yeast-locked efg1Δ/Δ cells from different C. albicans strain backgrounds.
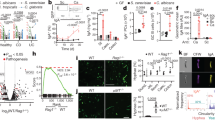
a, Experiments were performed as described in Fig. 1. Mouse GI organs (St; stomach, Si; small intestine, Ce; cecum, Co; colon) were homogenized to identify WT and efg1Δ/Δ cells 14 days post-C. albicans inoculation. WT (CAY2698) versus efg1Δ/Δ (CAY11750) competitions in SD + P/S (P/S; penicillin/streptomycin) model (a), PD + P/S model (b) and PD model (c). dpi; days post-inoculation. d, Schematic of competition between WT and yeast-locked efg1Δ/Δ cells. Mice fed the SD without antibiotics were gavaged with a 1:1 mixture of WT and efg1Δ/Δ cells and C. albicans colonies were examined in fecal pellets at the indicated time points. e,f, Competition outcomes (e) and fecal CFUs (f) of WT versus efg1Δ/Δ in 529L (CAY5016 versus CAY11482) and CHN1 (CAY11170 versus CAY11184). Experiments were performed in BALB/c and C57BL/6 J mice as indicated. Each data point represents an individual mouse, data are mean ± s.e.m in a, b, and f. n = 3 for competitions in BALB/c mice and n = 4 for competitions in C57BL/6 J mice. A paired t-test (two-tailed) was used in e and a Mann-Whitney (two-tailed) test in f.
Extended Data Fig. 2 Comparison of the fitness of WT and flo8Δ/Δ cells in different colonization models.
a, Microscopic images of cells in the colon of BALB/c SD + P/S mice infected with WT (CAY2698) or flo8Δ/Δ (CAY9796) cells at 7 days post-infection. b, Quantitative analysis of yeast and filamentous morphotypes. n = 2. c, Schematic of competition between WT and flo8Δ/Δ cells. n = 2. Each data point represents an individual mouse. A 1:1 mix of WT and flo8Δ/Δ cells were inoculated by oral gavage and fecal samples were collected at the indicated time points. d-f, Results for SC5314 WT (CAY2698) versus flo8Δ/Δ (CAY9796) cells in GF BALB/c mice (d), conventional PD-fed BALB/c mice treated with antibiotics (P/S; penicillin/streptomycin) (e), and conventional PD-fed BALB/c mice (without antibiotics) (f). dpi; days post-inoculation. n = 3 in e and f. g-i, Competition of WT versus flo8Δ/Δ cells in BALB/c mice fed a SD (without antibiotics). Experiments were performed using SC5314 (CAY2698 vs. CAY9796) (g), 529 L (CAY11168 vs. CAY11180) (h), or CHN1 (CAY11170 vs. CAY11186) (i) strain backgrounds. n = 3. Each data point represents an individual mouse. Data are mean ± s.e.m in b. Significance was determined by a Mann-Whitney (two-tailed) test in b and a paired two-tailed t-test in d-i.
Extended Data Fig. 3 Analysis of microbiome composition upon changes in diet, antibiotic treatment, and C. albicans colonization.
a, Schematic of colonization experiments. BALB/c mice were fed the SD or PD, with or without antibiotics (P/S; penicillin/streptomycin). These groups were compared with and without colonization with C. albicans SC5314 cells. b, C. albicans CFUs in fecal pellets collected at the indicated time points. LOD, limit of detection. n = 8. c,d, Total bacterial levels in fecal pellets collected at the indicated time points by quantitative PCR in SD- or PD-fed mice with antibiotics (c) and without antibiotics (d). n = 8. e, Bubble plot depicting the amount of variation in gut microbial composition determined by Permutational Multivariate Analysis of Variance (PERMANOVA) analysis using the adonis function and Bray-Curtis distances of beta diversity. Effect size refers to the magnitude of the differences or dissimilarities between groups. f, Shannon diversity of control mice on SD or PD (n = 8). g-m, Distribution of bacterial phyla in different models as determined by MaAsLin2. PD vs. SD (g), SD vs. SD + P/S (h), PD vs. PD + P/S (i), SD +/- C. albicans (j), PD +/- C. albicans (k), PD + P/S +/- C. albicans (l) and SD + P/S +/- C. albicans (m). Coefficient with standard error shown on x-axis. adj p-val cutoff 0.05. n, Relative abundance of Actinobacteria across different diet fed mice (+/− Candida) on Day 21. n = 8. Each data point represents an individual mouse. Data are mean ± s.e.m in b-m. A Mann-Whitney test was used in b,f,and n.
Extended Data Fig. 4 Linear discriminant analysis Effect Size (LEfSe) analysis to compare the alterations in gut bacterial populations in mice on different diets.
Linear discriminant analysis (LDA) effect size for significant taxa in the microbiome of mice fed the SD vs. PD are plotted onto a cladogram for Day 0 (a), Day 7 (b), Day 14 (c), Day 21 (d). and for mice fed the PD (+ /− C. albicans) for Day 0 (e), Day 7 (f), Day 14 (g) and Day 21 (h). Analysis was performed on mice as shown in Extended Data Fig. 3 (n = 8 per group). Differentially abundant species with LDA score >2 are shown in nodes represented with red (PD) and green (SD) and non-significant species are represented with yellow. A Kruskal-Wallis test was used to compare features between diets (p < 0.05) and the Pairwise Wilcoxon test was used to compare between taxa (p < 0.05).
Extended Data Fig. 5 Analysis of gut bacterial populations in different murine models.
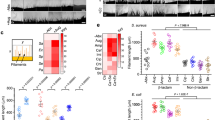
16S rRNA sequencing data was used to determine the relative abundance of bacterial phyla in BALB/c colonization experiments using mice fed the SD or PD, and supplemented or not supplemented with antibiotics (P/S; penicillin/streptomycin). Each group of mice were also compared +/− inoculation with C. albicans SC5314 cells, as shown in Extended Data Fig. 3. a,b, Shannon diversity for bacterial populations from fecal pellets for the shown experiments. n = 8. c, Relative abundance of bacteria shown for fecal pellets for days 0, 10, 14, and 21, and for the small intestine at day 21. n = 8. Data are mean ± s.e.m in a-c.
Extended Data Fig. 6 C. albicans WT cells outcompete efg1Δ/Δ cells in gnotobiotic mice harboring different bacterial populations.
a, Schematic of competition between WT and yeast-locked efg1Δ/Δ cells in gnotobiotic colonization models. Mice were gavaged with a 1:1 mixture of WT SC5314 (CAY2698) and efg1Δ/Δ (CAY11750) cells. b-d, C. albicans cells were tested in GF BALB/c mice (b), GF NMRI mice (c), or GF C57BL/6 mice colonized with E. coli prior to inoculation with C. albicans (d). n = 4 in b, 8 in c, and 5 in d. e, C. albicans CFUs in fecal pellets collected at the indicated time points in GF NMRI mice (e) and GF C57BL/6 mice (f). n = 8 in e, n = 3–5 in f. LOD, limit of detection. g, WT v. efg1Δ/Δ competition in Amp-treated mice gavaged with heat-killed AmpR E. coli. n = 4. h,i, Competition between CHN1 WT (CAY11170) and efg1∆/∆ (CAY11184) cells in WT mice (h) or Rag1-/- mice (i) on the SD (no antibiotics). n = 4. j, C. albicans CFU levels in the fecal pellets from WT and Rag1-/- mice. n = 4. Each data point represents an individual mouse. Data are mean ± s.e.m in j. A paired t-test (two-tailed) was used in b-d and g-i, and a Mann-Whitney (two-tailed) test in e,f and j.
Extended Data Fig. 7 Examining the role of ALS3 and SOD5 in GI colonization fitness.
a, Schematic of competition between WT C. albicans and sod5Δ/Δ or als3Δ/Δ cells (SC5314 background). b-e, BALB/c mice fed a PD without antibiotics or a SD with antibiotics (P/S; penicillin/streptomycin). Experiments used WT (CAY8785) and sod5Δ/Δ (CAY14738) in b and c. n = 4. WT (CAY8785) and als3Δ/Δ (CAY14696) in d and e. n = 8. Each data point represents an individual mouse. A paired t-test (two-tailed) was used for statistical significance.
Extended Data Fig. 8 Colonization fitness of WT and ece1Δ/Δ mutants.
Experiments were performed as described in Fig. 3 Mouse GI organs (St; stomach, Si; small intestine, Duo; duodenum, Je; jejunum, Ile; ileum, Ce; cecum, Co; colon) were homogenized to identify the ratio of WT and ece1Δ/Δ cells 14 days post C. albicans infection. b, Competition between CAY11533 (WT) versus CAY11507 (ece1Δ/Δ) cells in BALB/c fed a PD. n = 5. c, Competition between WT (CAY11168) and ece1Δ/Δ cells (CAY12441) in the 529L background BALB/c mice fed a PD. n = 4. d, Competition between WT (CAY11170) and ece1Δ/Δ cells (CAY12446) in the CHN1 background in SD-fed BALB/c mice. n = 4. e-g, Competition between CAY12202 (WT) versus CAY8578 (ece1Δ/Δ) BALB/c mice fed a SD plus antibiotics (P/S; penicillin/streptomycin) (e), GF C57BL/6 mice (f), and GF C57BL/6 mice colonized with E. coli prior to C. albicans inoculation (g). n = 4 in e, and 5 in f and g. Competition outcomes of WT (CAY12202) versus ece1Δ/Δ+ECE1ΔpIII (CAY8580) in GF C57BL/6 mice (h) and in BALB/c mice fed a PD (i). n = 4 in h and i. Each data point represents an individual mouse. Data are mean ± s.e.m in c–e, h, i. A paired t-test (two-tailed) was used to determine the significance between two populations (b-h).
Extended Data Fig. 9 Comparison of C. albicans WT and ece1Δ/Δ cells in vitro.
a, Heat map of transcriptome analysis representing the fold change (FC) in expression of fungal genes when strains were grown under hyphal-inducing conditions (3 h, 37 °C, 5% CO2). b-h, Competitive fitness of SC5314 WT and ece1Δ/Δ cells in RPMI (37 °C, 5% CO2) (b), acidic pH (pH 4.7) (c), high salt (1 M NaCl) (d), hypoxia (1% O2) (e), oxidative stress (2 mM H2O2) (f), in the presence of 2% acetate (g), and during incubation with TR146 epithelial cells (h). Competitive index was calculated using the formula: log2 [(MUTcompetition/WTcompetition)/ (MUTsingle/WTsingle)]. Fitness of the ece1Δ/Δ relative to the WT is represented as log2 competitive index. Data are representative of three biological replicates (n = 3). Graphs show the mean ± sem in b - h. Statistical analysis was performed using 1-way ANOVA with Bonferroni post hoc test to detect significance.
Supplementary information
Supplementary Tables
Supplementary Tables 1–3
Source data
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Liang, SH., Sircaik, S., Dainis, J. et al. The hyphal-specific toxin candidalysin promotes fungal gut commensalism. Nature 627, 620–627 (2024). https://doi.org/10.1038/s41586-024-07142-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-024-07142-4
This article is cited by
-
Hyphae promote Candida albicans fitness and commensalism in the gut
Nature Reviews Microbiology (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.