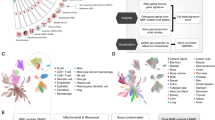
Abstract
Many cancers originate from stem or progenitor cells hijacked by somatic mutations that drive replication, exemplified by adenomatous transformation of pulmonary alveolar epithelial type II (AT2) cells1. Here we demonstrate a different scenario: expression of KRAS(G12D) in differentiated AT1 cells reprograms them slowly and asynchronously back into AT2 stem cells that go on to generate indolent tumours. Like human lepidic adenocarcinoma, the tumour cells slowly spread along alveolar walls in a non-destructive manner and have low ERK activity. We find that AT1 and AT2 cells act as distinct cells of origin and manifest divergent responses to concomitant WNT activation and KRAS(G12D) induction, which accelerates AT2-derived but inhibits AT1-derived adenoma proliferation. Augmentation of ERK activity in KRAS(G12D)-induced AT1 cells increases transformation efficiency, proliferation and progression from lepidic to mixed tumour histology. Overall, we have identified a new cell of origin for lung adenocarcinoma, the AT1 cell, which recapitulates features of human lepidic cancer. In so doing, we also uncover a capacity for oncogenic KRAS to reprogram a differentiated and quiescent cell back into its parent stem cell en route to adenomatous transformation. Our work further reveals that irrespective of a given cancer’s current molecular profile and driver oncogene, the cell of origin exerts a pervasive and perduring influence on its subsequent behaviour.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
Tissue microarray data from ref. 31 reanalysed and presented in Fig. 4 were obtained from the NCBI’s Gene Expression Omnibus under accession number GSE58772. Histologically annotated mutation data from ref. 32 reanalysed and presented in Fig. 4 were obtained from cBioPortal (www.cbioportal.org; ‘Lung Adenocarcinoma (MSK, J Thorac Oncol 2020)’). The mouse reference genome used for mapping reads for scRNA-seq, Gencode version GRCm39, is accessible at https://www.gencodegenes.org/mouse. scRNA-seq data have been deposited as a BioProject with accession number PRJNA975973. Source data are provided with this paper.
Code availability
Code and processed data matrix files used for the scRNA-seq data are accessible at https://github.com/transcriptomics/AT1-Lung-Cancer.
References
Desai, T. J., Brownfield, D. G. & Krasnow, M. A. Alveolar progenitor and stem cells in lung development, renewal and cancer. Nature 507, 190–194 (2014).
Siegel, R. L., Miller, K. D., Fuchs, H. E. & Jemal, A. Cancer statistics, 2022. CA Cancer J. Clin. 72, 7–33 (2022).
Desai, T. J. Developmental insights into lung cancer. Annu. Rev. Cancer Biol. 5, 351–369 (2021).
Logan, C. Y. & Desai, T. J. Keeping it together: pulmonary alveoli are maintained by a hierarchy of cellular programs. Bioessays 37, 1028–1037 (2015).
Lin, C. et al. Alveolar type II cells possess the capability of initiating lung tumor development. PLoS ONE 7, e53817 (2012).
Xu, X. et al. Evidence for type II cells as cells of origin of K-Ras-induced distal lung adenocarcinoma. Proc. Natl Acad. Sci. USA 109, 4910–4915 (2012).
McFadden, D. G. et al. Mutational landscape of EGFR-, MYC-, and Kras-driven genetically engineered mouse models of lung adenocarcinoma. Proc. Natl Acad. Sci. USA 113, E6409–E6417 (2016).
Okubo, K. et al. Bronchoalveolar carcinoma: clinical, radiologic, and pathologic factors and survival. J. Thorac. Cardiovasc. Surg. 118, 702–709 (1999).
Travis, W. D. et al. International Association for the Study of Lung Cancer/American Thoracic Society/European Respiratory Society international multidisciplinary classification of lung adenocarcinoma. J. Thorac. Oncol. 6, 244–285 (2011).
Jones, K. D. Whence lepidic?: the history of a Canadian neologism. Arch. Pathol. Lab. Med. 137, 1822–1824 (2013).
Juul, N. H., Stockman, C. A. & Desai, T. J. Niche cells and signals that regulate lung alveolar stem cells in vivo. Cold Spring Harb. Perspect. Biol. https://doi.org/10.1101/cshperspect.a035717 (2020).
Wolf, F. A. et al. PAGA: graph abstraction reconciles clustering with trajectory inference through a topology preserving map of single cells. Genome Biol. 20, 59 (2019).
Nagendran, M., Andruska, A. M., Harbury, P. B. & Desai, T. J. Advances in proximity ligation in situ hybridization (PLISH). Bio-protocol 10, e3808 (2020).
Nabhan, A., Brownfield, D. G., Harbury, P. B., Krasnow, M. A. & Desai, T. J. Single-cell Wnt signaling niches maintain stemness of alveolar type 2 cells. Science https://doi.org/10.1126/science.aam6603 (2018).
Zacharias, W. J. et al. Regeneration of the lung alveolus by an evolutionarily conserved epithelial progenitor. Nature 555, 251–255 (2018).
Tammela, T. et al. A Wnt-producing niche drives proliferative potential and progression in lung adenocarcinoma. Nature 545, 355–359 (2017).
Pacheco-Pinedo, E. C. et al. Wnt/beta-catenin signaling accelerates mouse lung tumorigenesis by imposing an embryonic distal progenitor phenotype on lung epithelium. J. Clin. Invest. 121, 1935–1945 (2011).
Juan, J., Muraguchi, T., Iezza, G., Sears, R. C. & McMahon, M. Diminished WNT -> beta-catenin -> c-MYC signaling is a barrier for malignant progression of BRAFV600E-induced lung tumors. Genes Dev. 28, 561–575 (2014).
Uhlitz, F. et al. An immediate–late gene expression module decodes ERK signal duration. Mol. Syst. Biol. 13, 928 (2017).
Cicchini, M. et al. Context-dependent effects of amplified MAPK signaling during lung adenocarcinoma initiation and progression. Cell Rep. 18, 1958–1969 (2017).
Snyder, E. L. et al. Nkx2-1 represses a latent gastric differentiation program in lung adenocarcinoma. Mol. Cell 50, 185–199 (2013).
Marjanovic, N. D. et al. Emergence of a high-plasticity cell state during lung cancer evolution. Cancer Cell 38, 229–246 (2020).
Tata, P. R. et al. Developmental history provides a roadmap for the emergence of tumor plasticity. Dev. Cell 44, 679–693 (2018).
Rawlins, E. L., Clark, C. P., Xue, Y. & Hogan, B. L. The Id2+ distal tip lung epithelium contains individual multipotent embryonic progenitor cells. Development 136, 3741–3745 (2009).
Nichane, M. et al. Isolation and 3D expansion of multipotent Sox9+ mouse lung progenitors. Nat. Methods 14, 1205–1212 (2017).
Strunz, M. et al. Alveolar regeneration through a Krt8+ transitional stem cell state that persists in human lung fibrosis. Nat. Commun. 11, 3559 (2020).
Choi, J. et al. Inflammatory signals induce AT2 cell-derived damage-associated transient progenitors that mediate alveolar regeneration. Cell Stem Cell 27, 366–382 (2020).
Kobayashi, Y. et al. Persistence of a regeneration-associated, transitional alveolar epithelial cell state in pulmonary fibrosis. Nat. Cell Biol. 22, 934–946 (2020).
Yang, D. et al. Lineage tracing reveals the phylodynamics, plasticity, and paths of tumor evolution. Cell 185, 1905–1923 (2022).
Choi, Y. et al. Rethinking a non-predominant pattern in invasive lung adenocarcinoma: prognostic dissection focusing on a high-grade pattern. Cancers https://doi.org/10.3390/cancers13112785 (2021).
Zabeck, H. et al. Molecular signatures in IASLC/ATS/ERS classified growth patterns of lung adenocarcinoma. PLoS ONE 13, e0206132 (2018).
Caso, R. et al. The underlying tumor genomics of predominant histologic subtypes in lung adenocarcinoma. J. Thorac. Oncol. 15, 1844–1856 (2020).
Maeda, Y. et al. Kras(G12D) and Nkx2-1 haploinsufficiency induce mucinous adenocarcinoma of the lung. J. Clin. Invest. 122, 4388–4400 (2012).
Yun, C. H. et al. Structures of lung cancer-derived EGFR mutants and inhibitor complexes: mechanism of activation and insights into differential inhibitor sensitivity. Cancer Cell 11, 217–227 (2007).
Hayakawa, Y., Nakagawa, H., Rustgi, A. K., Que, J. & Wang, T. C. Stem cells and origins of cancer in the upper gastrointestinal tract. Cell Stem Cell 28, 1343–1361 (2021).
Concepcion, C. P. et al. Smarca4 inactivation promotes lineage-specific transformation and early metastatic features in the lung. Cancer Discov. 12, 562–585 (2022).
Sutherland, K. D. et al. Multiple cells-of-origin of mutant K-Ras-induced mouse lung adenocarcinoma. Proc. Natl Acad. Sci. USA 111, 4952–4957 (2014).
Brownfield, D. G. et al. Alveolar cell fate selection and lifelong maintenance of AT2 cells by FGF signaling. Nat. Commun. 13, 7137 (2022).
Liu, Z. et al. MAPK-mediated YAP activation controls mechanical-tension-induced pulmonary alveolar regeneration. Cell Rep. 16, 1810–1819 (2016).
Treutlein, B. et al. Reconstructing lineage hierarchies of the distal lung epithelium using single-cell RNA-seq. Nature 509, 371–375 (2014).
van Veen, J. E. et al. Mutationally-activated PI3′-kinase-alpha promotes de-differentiation of lung tumors initiated by the BRAF(V600E) oncoprotein kinase. Elife https://doi.org/10.7554/eLife.43668 (2019).
Weibel, E. R. On the tricks alveolar epithelial cells play to make a good lung. Am. J. Respir. Crit. Care Med. 191, 504–513 (2015).
Nicholson, A. G. et al. The 2021 WHO classification of lung tumors: impact of advances since 2015. J. Thorac. Oncol. 17, 362–387 (2022).
Travis, W. D. et al. The IASLC Lung Cancer Staging Project: proposals for coding T categories for subsolid nodules and assessment of tumor size in part-solid tumors in the forthcoming eighth edition of the TNM classification of lung cancer. J. Thorac. Oncol. 11, 1204–1223 (2016).
Berry, M. F. et al. Presence of even a small ground-glass component in lung adenocarcinoma predicts better survival. Clin. Lung Cancer 19, e47–e51 (2018).
Ye, T. et al. Lung adenocarcinomas manifesting as radiological part-solid nodules define a special clinical subtype. J. Thorac. Oncol. 14, 617–627 (2019).
Hattori, A. et al. Distinct clinicopathologic characteristics and prognosis based on the presence of ground glass opacity component in clinical stage IA lung adenocarcinoma. J. Thorac. Oncol. 14, 265–275 (2019).
Hou, Y. et al. The presence of lepidic and micropapillary/solid pathological patterns as minor components has prognostic value in patients with intermediate-grade invasive lung adenocarcinoma. Transl. Lung Cancer Res. 11, 64–74 (2022).
Moreira, A. L. et al. A grading system for invasive pulmonary adenocarcinoma: a proposal from the International Association for the Study of Lung Cancer Pathology Committee. J. Thorac. Oncol. 15, 1599–1610 (2020).
Joubert, P. & Travis, W. D. Prognostic impact of ground-glass opacity/lepidic component in pulmonary adenocarcinoma: a hazy staging dilemma. J. Thorac. Oncol. 17, 19–21 (2022).
Takeda, N. et al. Interconversion between intestinal stem cell populations in distinct niches. Science 334, 1420–1424 (2011).
Rock, J. R. et al. Multiple stromal populations contribute to pulmonary fibrosis without evidence for epithelial to mesenchymal transition. Proc. Natl Acad. Sci. USA 108, E1475–1483 (2011).
Chapman, H. A. et al. Integrin alpha6beta4 identifies an adult distal lung epithelial population with regenerative potential in mice. J. Clin. Invest. 121, 2855–2862 (2011).
Jackson, E. L. et al. Analysis of lung tumor initiation and progression using conditional expression of oncogenic K-ras. Genes Dev. 15, 3243–3248 (2001).
Harada, N. et al. Intestinal polyposis in mice with a dominant stable mutation of the beta-catenin gene. EMBO J. 18, 5931–5942 (1999).
Muzumdar, M. D., Tasic, B., Miyamichi, K., Li, L. & Luo, L. A global double-fluorescent Cre reporter mouse. Genesis 45, 593–605 (2007).
Hama, H. et al. Scale: a chemical approach for fluorescence imaging and reconstruction of transparent mouse brain. Nat. Neurosci. https://doi.org/10.1038/nn.2928 (2011).
Nagendran, M., Riordan, D. P., Harbury, P. B. & Desai, T. J. Automated cell type classification in intact tissues by single-cell molecular profiling. Elife https://doi.org/10.7554/eLife.30510 (2018).
Bankhead, P. et al. QuPath: open source software for digital pathology image analysis. Sci. Rep. 7, 16878 (2017).
Raudvere, U. et al. g:Profiler: a web server for functional enrichment analysis and conversions of gene lists (2019 update). Nucleic Acids Res. 47, W191–W198 (2019).
Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl Acad. Sci. USA 102, 15545–15550 (2005).
Robinson, J. T. et al. Integrative genomics viewer. Nat. Biotechnol. 29, 24–26 (2011).
Acknowledgements
We thank R. Nusse, M. Winslow, L. Attardi, S. Plevritis and members of the laboratories of T.J.D. and R. Nusse for valuable discussions, the CZI Biohub, P. Chen, D. Wu, M. Eckart, C. Carswell Crumpton, C. Pan and J. Perrino. This research was supported in part by the Virginia and D.K. Ludwig Fund for Cancer Research. The project described was supported, in part, by NIH S10 Award Number 1S10OD028536-01, titled “OneView 4kX4k sCMOS camera for transmission electron microscopy applications” from the Office of Research Infrastructure Programs. Its contents are solely the responsibility of the authors and do not necessarily represent the official views of the National Center for Research Resources or the National Institutes of Health. The following are acknowledged for funding: NHLBI 5R01HL14254902 and NIH 5UG3HL14562302 (T.J.D.); Ludwig Cancer Institute (T.J.D., N.H.J. and R.S.); NIH 1T32HL129970-01A1, NIH 5T32HL129970-03, NIH Loan Repayment Program and NHLBI 1 F32 HL147417-01 (N.H.J.); Stanford Medical Scholars Fellowship Program (M.C.M.); Woods Family Endowed Faculty Scholar in Pediatric Translational Medicine of the Stanford Child Health Research Institute and Robert A. and Gertrude T. Hudson Endowed Professorship (T.J.D.).
Author information
Authors and Affiliations
Contributions
N.H.J. designed and conducted immunohistochemistry, in situ, scRNA-seq and transgenic mouse experiments; conducted the bulk of the statistical analyses; conducted the scRNA-seq analysis; created the figures; and contributed to the writing of the manuscript. J.-K.Y. conducted the tissue microarray and WNT mutation analysis, and assisted with scRNA-seq analysis and generation of associated figures. M.C.M. and N.R. conducted immunohistochemistry and imaging on human tissue and assisted in analysis of immunohistochemistry data. Y.I.K. conducted immunohistochemistry and imaging of mouse tissue. M.M. and N.F.F. facilitated scRNA-seq. W.L.T. and J.B.S. assisted in obtaining appropriate human tissue. R.S. assisted in conception of the scRNA-seq experiment and guided its analysis. T.J.D. conceived the original hypothesis and subsequent experiments, supervised the project and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature thanks the anonymous reviewers for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 A subset of AT1 cells is unresponsive to KrasG12D, and pJUN expression in AT1-phenotype cells in Hopx-CreER>KrasG12D lungs is not increased.
(a) PAGA of Hopx-CreER>KrasG12D mTmG mouse highlighting 13 AT1-phenotype cells expressing KrasG12D at 10m after tamoxifen. (b) (left) Immunostaining analysis of pJUN in AT1-phenotype cells (NKX2-1+/LAMP3−) in Hopx-CreER>KrasG12D and Hopx-CreER>KrasWT mice reveals pJUN+ (filled caret) and pJUN− (open caret) cells with (right) quantitation (p = 0.4601) (n = 3 WT mice and 4 KrasG12D mice, 3 25x fields per mouse). Scale bar, 10 μm m, months; PAGA, Partition-Based Graph Abstraction; pJUN, phosphorylated c-JUN and JUND; WT, wild type. Data in bar graphs b are mean±s.e.m. P value calculated by unpaired two-tailed Student’s t test.
Extended Data Fig. 2 Molecular comparison of AT2 and iAT2 cells and their corresponding early-stage tumors.
(a) Co-immunostaining of early-stage AT1 tumors shows absence of AT1 (LEL, AGER) and presence of AT2 (SFTPB, SFTPC) markers in hyperplastic mGFP+ cells with near-cuboidal morphologies (n = 5 Hopx-CreER>KrasG12D mice). (b) (top) Feature plots for AT2 markers (Sftpa1, Lyz2) and Axin2 and indicated cell classes shows co-clustering with intermixing of individual Axin2+ AT2 (blue) and iAT2 (red) cells. (bottom) Heatmap of the most highly differentially expressed genes between these two populations highlights their remarkably similar transcriptional profiles. (c) (top) Feature plots highlighting AT2>Kras cells 18d after tamoxifen and Early tumor stage AT1>Kras cells and (bottom) heatmap of their most differentially expressed genes. Note that the early-stage AT1 and AT2 tumor populations are non-overlapping and initiate distinct trajectories as visualized on PAGA. Scale bars, 10 μm. LEL, Lycopersicon esculentum lectin; PAGA, Partition-Based Graph Abstraction.
Extended Data Fig. 3 Histological features and classification of AT1- and AT2-derived lung adenomas.
(a) H&E of Hopx-CreER>KRASG12D and Sftpc-CreER-rtTA>KRASG12D lungs showing examples of the predominant (top) lepidic histology of AT1-derived and (bottom) non-lepidic histology of AT2-derived adenomas. Note immune cells largely restrict to tumor foci in lepidic AT1 but infiltrate multiple uninvolved alveoli in solid AT2 tumors (n = 6 mice of each genotype). (b) H&E of human lepidic (25% lepidic non-mucinous (green arrows) with 75% mucinous papillary) and non-lepidic (mixed mucinous and pleomorphic spindle cell carcinoma) KRAS-driven LuAds (n = 3 lepidic and 3 non-lepidic LuAds). (c) Exemplary H&E stains of histological classifications of lung tumors applied in the manuscript, with (right) high-magnification image of mixed histology AT1 tumor (n = 6 mice of each genotype). Scale bars, 200 μm (a, c) and 400 μm (b).
Extended Data Fig. 4 Advanced AT1-derived adenomas are highly proliferative and enriched for gene scores of a high plasticity tumor and AT2-to-AT1 molecular intermediate states.
(a) Feature plot shows enrichment of Mki67 at Late tumor stage. (b) Gene scores derived from marker genes for high plasticity state in advanced AT2-derived Kras LuAd and for AT2-to-AT1 transitional intermediates in lung injury show enrichment in cells bridging Early and Late AT1 tumor stages. ADI, alveolar differentiation intermediate; DATP, damage-associated transient progenitor; PATS, pre-alveolar type 1 transitional cell state.
Extended Data Fig. 5 Diverse molecular states in human LuAd tumors, including lung/gastric hybrid cells.
(a) Tumor cells co-expressing lung (NKX2-1) and intestinal markers are found in human lepidic mucinous (top, CTSE, left side of image) and non-mucinous (bottom HNF4A, open caret) LuAds. The non-mucinous lepidic tumor also contains NKX2-1+HNF4A− (arrowhead) and NKX2-1−HNF4A+ (arrow) cells. The mucinous (MUC5AC+) region is NKX2-1Lo (asterisk) and distinct from the CTSE+ region. (b) Immunostaining of human invasive mucinous LuAd reveals co-expression of MUC5AC and CTSE by NKX2-1− tumor cells (arrowheads). Non-mucinous tumor regions contain cells simultaneously expressing lung and gastric markers (NKX2-1+CTSE+, open carets), as well as NKX2-1−CTSE+, and NKX2-1+CTSE− cells. Note some regions are combined mucinous and gastric (MUC5AC+CTSE+) while others are gastric-only (MUC5AC−CTSE+) (n = 2 individuals). Scale bars, 20 μm.
Extended Data Fig. 6 Squamous cells co-expressing AT1 and AT2 markers and also carrying the driver mutation reside at the periphery of human lepidic tumors.
(a-b) Co-staining of AT1 (AGER), AT2 (MUC1) and driver mutation (EGFR L858R) antibodies in human lepidic LuAd shows expected oncogene and AT2 marker co-staining of the tumor but also flat cells in the periphery positive for the oncogene that co-express both AT1 and AT2 markers (LuAd1) or AT1 marker (LuAd2) (arrowheads and inset) (n = 3 individuals). Scale bars, 20 μm (a, b lower rows), 200 μm (a, b upper rows).
Supplementary information
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Juul, N.H., Yoon, JK., Martinez, M.C. et al. KRAS(G12D) drives lepidic adenocarcinoma through stem-cell reprogramming. Nature 619, 860–867 (2023). https://doi.org/10.1038/s41586-023-06324-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-023-06324-w
This article is cited by
-
AT1 cells appear centre stage
Nature Reviews Cancer (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.