Abstract
A challenge during development is to ensure lineage segregation while preserving plasticity. Using pluripotency progression as a paradigm, we review how developmental transitions are coordinated by redeployments, rather than global resettings, of cellular components. We highlight how changes in response to extrinsic cues (FGF, WNT, Activin/Nodal, Netrin-1), context- and stoichiometry-dependent action of transcription factors (Oct4, Nanog) and reconfigurations of epigenetic regulators (enhancers, promoters, TrxG, PRC) may confer robustness to naïve to primed pluripotency transition. We propose the notion of Molecular Versatility to regroup mechanisms by which molecules are repurposed to exert different, sometimes opposite, functions in close stem cell configurations.
Similar content being viewed by others
Introduction
In contrast to the all-or-none view of the unidirectional differentiation of a stem cell into terminally specialised derivatives, the development of multicellular organisms involves developmental transitions that often generate plastic and reversible cellular intermediate states. To achieve these transitions, embryonic cells need to integrate complex sets of external cues and respond by precise epigenetic/transcriptomic changes that ensure the stepwise and coordinated lineage segregation1. Mouse peri-implantation development constitutes a prototypical example of such dynamic transitions during which the naïve pluripotent cells of the early epiblast transit to formative and primed pluripotent identities prior to differentiating. During this process, the reconfigurations of the epigenetic/transcriptional programs permit the emergence of distinct molecular features, including the transient competence to form primordial germ cells (PGCs)2. However, the embryonic cells also maintain their ability to self-renew and sustain the expression of certain core pluripotency transcription factors (TFs)2,3. Therefore, a critical challenge for peri-implantation cells is to robustly acquire novel features while preserving their stemness in constrained developmental space and time. In this Perspective, we exploited recent findings in the pluripotency field to question whether these cellular transitions involve exclusively complete molecular resettings (–e.g. extinction of a full transcriptional/epigenetic program and induction of a non-overlapping new one), as traditionally proposed for differentiation processes4, or also alternative mechanisms. Hence, we herein compiled convergent findings depicting how a large battery of molecules are efficiently repurposed to exert strikingly different, sometimes opposite, functions in these closely-related stem cell configurations. We thus propose more broadly that pluripotency progression is also ensured by the fine-tuned redeployment, complementary to the resetting, of a large repertoire of intrinsic components of the cell, ranging from the plasma membrane to the chromatin. Far from a semantic argument, we propose the notion of Molecular Versatility (see Box 1) to regroup these mechanisms that, we believe, might have been evolutionary co-opted to confer robustness to developmental transitions.
During peri-implantation development, pluripotency does not correspond to a unique and fixed cellular state but rather encompasses multiple evolving cellular configurations (Fig. 1)5. These closely-related cellular states share the ability to self-renew and the expression of a set of core TFs, as reviewed elsewhere3,6. Yet, they undergo profound changes in their ability to respond to extrinsic cues, that allow the establishment of the pluripotent compartment of the early and late epiblasts7. Before implantation (embryonic day E4.5), mouse epiblast cells reside in a naïve state, which progressively transitions to formative (E5.5–E6.5) and primed (E6.5–E7.5) configurations upon implantation (Fig. 1)8. These transitions, which are crucial for germ layer formation, prepare embryonic cells for lineage specification while maintaining stemness. The regulatory mechanisms controlling these transitions are complex to decipher in vivo due to the limited access to and quantity of biological material at these embryonic stages. Yet they can be dissected using tractable and faithful in vitro models of embryo-derived stem cells. These models were initially classified dichotomously with mouse embryonic stem cells (ESCs) reflecting the naïve epiblast9, and epiblast stem cells (EpiSCs) resembling the primed epiblast of the anterior primitive streak10,11. Intermediate cellular states were captured, such as the transient Epiblast-like cells (EpiLCs)12. More recently, Rosette-like stem cells (RSCs)13 and pluripotent stem cells harbouring features of formative pluripotency were captured using various culture conditions14,15,16 (Fig. 1). Like their in vivo counterparts, these closely related cell types are characterised by distinct developmental properties, signalling requirements, transcriptomes and epigenomes. Notably, naïve ESCs are unresponsive to germ cell inductive stimuli, in contrast to EpiLCs and formative stem (FS) cells that acquired such competence. Yet, EpiSCs already lost responsiveness to PGC induction (Fig. 1). Of note, the transition from naïve to formative pluripotency can also be explored in vitro using fluorescent reporters of the naïve marker Rex1/Zfp42 in ESCs17.
The schematic diagram depicts the embryonic stages and the corresponding embryo-derived cell types. It also summarises the signalling pathways that sustain the embryo-derived stem cells in vitro and their ability to respond to germ cell inductive signals. ESC: Embryonic Stem Cell, RSC: Rosette Stem Cells13, XPSC: X Pluripotent Stem Cells15, EpiLC: Epiblast-like Cells12, FS: Formative Stem cells137, fPSC: Formative Pluripotent Stem Cells16, EpiSC: Epiblast Stem Cells.
Therefore, during pluripotency progression, an apparently contradictory challenge for embryonic cells is to retain stemness while learning to respond accurately to lineage specifying cues3. A critical underlying question is how such challenge is addressed rapidly (in few days) and precisely at the molecular level. Recent attempts combining CRISPR/Cas9 gene disruption, large-scale transcriptomics and computational systems biology revealed that the transition from naïve to formative pluripotency is not based on the action of few master TFs but rather the product of multiple crosstalking components linking at least four signalling inputs with epigenetic and TF networks to rewire embryonic cells18,19. Here, we selected in the recent literature a battery of molecular mechanisms by which signalling pathways, TFs and epigenetic regulators are repurposed to reach this objective. In addition to the established notion that molecules can exert strikingly different functions in unrelated cell types, we propose the notion of Molecular Versatility to regroup these molecular mechanisms that specifically take place in similar or closely related stem cell configurations during peri-implantation development (Box 1).
Redeploy signalling pathways
The maintenance of the naïve, formative and primed pluripotent states in vitro heavily relies on signalling pathways20. It has to be noted that some of these signals may be dispensable for embryo development, except under certain conditions (diapause), as reported for LIF21. Naïve ESCs self-renew in a metastable state in the presence of LIF and BMP422 whereas the activation of the WNT pathway prevents their transition towards primed EpiSC23. The combined use of chemical inhibitors (2i) to suppress FGF/MAPK (via MEK1/2) and activate WNT (via GSK3 α/β) signalling instructs a ground state of pluripotency24 that can be partially mimicked by the Netrin-1 signalling pathway25. In contrast, the combined inhibition of FGF/MAPK and WNT signalling results in the capture and maintenance of RSCs in a self-renewing intermediate naïve-primed state13. Moreover, EpiLCs, FS and EpiSCs rely on Activin/Nodal and FGF/MAPK to maintain their undifferentiated state10,12 (Fig. 1). In line with the Molecular Versatility notion, we selected below key mechanisms that allow closely related embryonic cells to integrate differently these signalling molecules.
The FGF/MAPK signalling pathway: a tight control of ERK1/2 activity
A first prototypical illustration of Molecular Versatility is associated with the FGF/MAPK pathway, and in particular with its receptors FGFR1/FGFR2 and its effectors ERK1/2. The activation of the pathway is initially instrumental for the emergence of the extraembryonic trophectoderm (TE) and primitive endoderm (PrE) lineages26. Of interest for the Perspective, the pathway is required for naïve ESCs to acquire competence to form somatic cell types12,27,28 but, in striking contrast, the pathway then sustains pluripotency in EpiLCs/FS/EpiSCs10,29. Therefore, the response to FGF must be rapidly and efficiently modified in these closely-related stem cell configurations to ensure those strikingly different effects (Fig. 2a). Even if the precise molecular mechanisms are not known, we describe below some related findings that might contribute to the phenomenon.
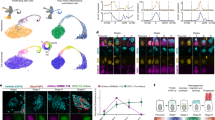
a FGF/MAPK signalling. The integration of the FGF signal is modulated by the differential expression of the FGFR1 and FGFR2 receptors. Such integration will lead to various degrees of ERK1/2 activation that will subsequently activate different sets of target genes. b Wnt signalling. The activation of the pathway will promote stem cell self-renewal or lineage commitment. These versatile effects are related to the bifunctional activity of β-catenin in the nucleus and in the cytoplasm. It has to be noted that TCF1 and TCF3 also behave antagonistically on pluripotency genes and ESC self-renewal. c Activin/Nodal signalling. Activin/Nodal signalling leads to a various degree of Smad2 activation. This precise degree triggers the regulation of different sets of genes and therefore of different cell fate decisions in ESCs. d Netrin-1 signalling. The Netrin-1 molecule has opposite effects on ESC fate by promoting self-renewal or lineage commitment. This Molecular Versatility is controlled by the stoichiometry of its receptors Neo1 and Unc5b that leads to ERK1/2 activity induction or repression.
A first element of response lies in the combined use of the FGF receptors on epiblast (Epi) and PrE progenitors. FGFR2 was long considered to be the main receptor actively promoting PrE emergence as the onset of its expression was detected by E3.5 in PrE-biased cells30,31. However, this sole FGF4/FGFR2 axis was not sufficient to explain the different functions exerted by the pathway in PrE emergence and Epi maturation. Quantitative single-cell-resolution imaging, combined with gene expression profiling of FGFR mutant/knock-in embryos, elegantly revealed how the stoichiometry of the receptors FGFR1 and FGFR2 is at least partially responsible for these differential actions28,32. While Epi-biased cells express solely FGFR1, PrE-biased blastomeres co-express FGFR1 and FGFR2. This variation ensures that apparently similar cells respond differently to the same ligand FGF4. The autocrine FGF4/FGFR1 axis is required for epiblast cells to mature towards the primed state while the paracrine FGF4/FGFR1/FGFR2 signalling stabilises the fate of PrE cells. In line with this, the activation of FGF/MAPK signalling in PrE- and Epi-biased cells was described to trigger the activation of different sets of target genes28. Accordingly, the recent development of ERK-kinase translocation reporter, enabling live quantification of ERK1/2 activity at the single-cell resolution in vivo, revealed that, prior to lineage specification, PrE progenitors already harbour higher ERK1/2 activity than Epi progenitors33. Of note, the sensitivity of embryonic cells to FGFs can also be mediated by the differential expression of FGF/MAPK inhibitors such as Dusp4, Dusp6, and Spred1, and the induction of negative feedback loops34 (Fig. 2a). The modulation of FGF availability and engagement on its receptors via heparan sulphate proteoglycans35 constitute other regulatory mechanisms, as well as the distinct usage of intracellular pathways (Jak/Stat, PLCγ, PI3K and MAPK), as reviewed elsewhere36,37.
Another interesting example of Molecular Versatility comes from works on the MEK/ERK factors that were found to exert dual functions in naïve ESCs, respectively promoting (i) self-renewal and (ii) lineage commitment. On the one hand, the chemical inhibition of MEK/ERK is known to constrain ESCs differentiation and to be instrumental for sustaining the ground state of naïve pluripotency24. However, on the other hand, Chen and colleagues demonstrated, using an inducible KO of both ERK1 and ERK2, that such complete depletion led to rapid genomic instability and compromised ESC self-renewal38. In line with this, the prolonged MEK inhibition was also shown to alter ground state ESC genomic stability39. Therefore, a precise dosage of ERK1/2 activity (intensity, duration) seems to dictate very different outcomes in naïve cells. An alternative way to tightly control the effect of ERK1/2 activity in ESCs came from an elegant study from the Brickman lab. The authors reported that the pro-lineage commitment activity of ERK1/2 is reversible, to a certain extent. Following stimulation, ERK1/2 was found to regulate first the transcription and enhancer activity of genes without inducing any changes in the binding of pluripotency TFs but rather by modifying the association of RNA polymerase II and cofactors such as Mediator components with the chromatin. Later on, if the stimulation persists, ERK1/2 finally dismantles pluripotency by directly phosphorylating/destabilising TFs such as Klf4/Nanog and RNA polymerase II at Ser5 residue. In contrast, if stimulation stops, the reduced ERK1/2 activity restores the normal expression of pluripotency proteins. This mode of regulation indicates how the activation of ERK1/2 activity induces a transient window of lineage commitment during which reversion to naïve pluripotency is preserved40. Altogether, these examples underline various ways by which embryonic cells may modulate their response to FGF signals.
The WNT signalling pathway: bi-functional properties of β-catenin
The WNT signalling pathway also exerts strikingly different functions in naïve and primed pluripotent stem cells. While its activation contributes to maintaining naïve pluripotency in ESCs41, it subsequently triggers exit from primed pluripotency and mesendoderm differentiation in EpiSCs42. We highlight below some molecular mechanisms that may allow embryonic cells to precisely modulate their responsiveness to Wnt during pluripotency progression.
First, the bi-functional nature of the effector β-catenin per se allows it to exert dual functions in promoting (i) self-renewal but also (ii) priming for differentiation (Fig. 2b). β-catenin harbours transactivating functions in the nucleus but it is also a component of adherens junctions in the cytoplasm. In the nucleus, β-catenin contributes to sustaining naïve pluripotency by triggering TCF3 inactivation by a plethora of mechanisms, ranging from the control of TCF3 protein binding/stability to the epigenetic regulation of its promoter or transcript by miRNA43,44. More recently, nuclear β-catenin was also found to strengthen the robustness of the transcriptional apparatus by supplying transcriptional co-regulators (such as BRD4, CDK9, Mediator, Cohesin and p300) at pluripotency loci45. In contrast in the cytoplasm, β-catenin contributes to controlling cell adhesion by connecting cadherins through α-catenin to the actin cytoskeleton46,47. The use of a TCF-signalling-defective-β-catenin variant demonstrated that its cell-adhesion functions are critical for endodermal and neuronal differentiation46.
Additional layers of regulation contribute to the Molecular Versatility of the Wnt pathway, such as the repertoire of the TCF TFs. In ESCs, TCF1 and TCF3 were shown to exert opposite functions, respectively promoting and destabilising the pluripotent network by competing for DNA binding through virtually identical HMG domains43,48,49. In addition, the noncanonical Wnt ligand Wnt5a was shown to exert opposite actions, namely activating or repressing β-catenin/TCF signalling, depending on the receptor it bound to (Ror2 or Frizzled4), indicating that the repertoire of receptors controls the response to WNT pathway50. This phenomenon will be discussed in more detail in the Netrin-1 signalling pathway section. Finally, the secretion of WNT antagonists, such as Dkk1 by the PrE13, and the existence of auto-activating loops, constitute alternative strategies to modulate response to WNT signalling51.
The Activin/Nodal signalling pathway: a dose-dependent switch of Smad binding
Activin and Nodal are morphogens of the TGFβ superfamily that direct differential stem cell fate decisions in a dose- and distance-dependent manner52. During early embryonic development, the pathway is responsible for the specification of mesoderm, endoderm, node, and mesendoderm. In contradiction to this function in differentiation, the pathway also plays important roles in ESCs and EpiSCs (Fig. 2c). Indeed, recombinant Activin A is used in vitro for the continued propagation and expansion of EpiSC10,29 and Activin/Nodal were found to promote ESC proliferation53,54. In that context, Lee and Colleagues proposed a fascinating model of how ESCs interpret and respond differentially to Activin/Nodal dosage55. They reported that, whereas the endogenous level of Activin/Nodal signaling is required for ESC self-renewal, its perturbation leads to an exit from pluripotency towards different paths: mesendoderm induction at high signalling level and TE differentiation at low signalling level. Mechanistically, phospho-Smad2, the primary downstream transcriptional effector of the Activin/Nodal pathway, was found to bind and regulate in a dose-dependent manner distinct subsets of target genes, including the Oct4 promoter. This dose-dependent switch of binding by phospho-Smad2 provides a molecular basis for the versatile functions of Activin/Nodal in ESCs55.
The Netrin-1 signalling pathway: modulating ligand and receptors stoichiometry
The Netrin-1 signalling pathway also permits to highlight Molecular Versatility during embryonic development. Netrins are a conserved family of laminin-related molecules that act chiefly through receptors of two distinct families: the deleted in colorectal cancer and UNC-5 families56. We recently revealed that the ligand Netrin-1, initially described as an axon guidance cue, exerts opposite effects on naïve ESC self-renewal, by activating or repressing ERK1/2, when the relative stoichiometry of its receptors Neogenin1 (Neo1) and Unc5b is experimentally modulated (Fig. 2d)25. Therefore, the repertoire of receptors allows the same ligand to trigger different responses in ESCs. Interestingly, the same receptor can also exert alternative functions depending on the ligand it binds to. Indeed, in the same settings, the Neo1 receptor was found to induce chemoattraction via Netrin-1 but chemorepulsion via the ligand Repulsive guidance molecule a (Rgma)57.
More generally, these results questioned why, rather than using single ligand and receptor couples to dictate different outcomes, signalling pathways comprise multiple ligand and receptor variants that interact promiscuously to generate large sets of distinct signalling complexes. The use of redundant ligands and receptors was proposed to confer regulatory flexibility58 or robustness to biological processes59 but this apparent redundancy recently appeared as a strong way to generate diverse and specific cellular responses. As an example, in an attempt to understand how the ligands Netrin-1 and Rgma cross-talk to transduce their signals, Robinson and colleagues reported that their simultaneous binding to Neo1 does not lead to competition for binding but to the formation of a ternary super-complex that globally diminishes their functional outputs60. Moreover, by combining theoretical and experimental approaches, the Elowitz laboratory showed how cells perceive absolute but also relative concentrations of multiple ligands to compute multi-dimensional response profiles. In this model, by modifying the stoichiometry of receptors, cells modulate the computations to perform, and therefore the responses to provide to the same combination of ligands61. Mathematical modelling even revealed that such multi ligand/receptor system outperforms seemingly more specific one-to-one signalling architectures in terms of diversity and specificity of responses62. In line with this, this section highlighted different mechanisms by which embryonic cells modulate their response to their surrounding signals.
Redirect transcription factors
Naïve, formative and primed stem cells are characterised by the shared expression of TFs such as Oct4 and Sox210,12,13, raising fascinating questions about how distinctive properties can be established in the presence of an overlapping set of master TFs. In line with the Molecular Versatility notion, a large fraction of pluripotency TFs were initially found to exert opposite functions in ESCs by promoting (i) self-renewal at endogenous levels but (ii) germ layer formation when ectopically expressed, including Sox2 (neurectoderm)63 and Esrrb/Tbx3 (endoderm)64,65. A limitation of these studies is that overexpression may show non-physiological activity of TFs as a result of non-specific binding and/or sequestration of factors. However, in the case of Oct4, its endogenous level was also reported to destabilise self-renewal, as illustrated by the fact that Oct4± heterozygous ESCs harbour a more robust pluripotent state than WT ESCs66. Therefore, based on these findings, Loh and Lim proposed to consider naïve pluripotency TFs as “pure lineage specifiers” and pluripotency as an inherently precarious condition in which these TFs compete to generate mutually exclusive lineages. This intriguing concept was slightly contrasting with the model proposed by A. Smith of a “ground” state of pluripotency in which pluripotency TFs rather cooperate to constrain the fluctuations provided by opposing differentiation programs24,67,68,69. Of note, this notion of TF battles was initially described in the hematopoietic system where single multipotent cells activate distinct lineage-affiliated gene expression programs prior to exclusive lineage commitment70.
Recently, computational systems biology approaches conducted on the transition from naïve to formative pluripotency rather proposed that pluripotency progression does not rely solely upon instructions delivered by few master TFs. It rather appears that a cloud of activity involving multiple coordinated and partially redundant inputs operates to destabilise the existing and self-reinforced naïve gene regulatory network19. In that context, the notion of “Molecular Versatility” presented herein might also provide a basis for unifying those principles. We propose that, during pluripotency transition, the stepwise dismantling of the robust naïve TF network, combined with epigenetic remodeling, allows pluripotency TFs to start exerting alternative functions. Therefore, rather than genuine lineage specifiers, pluripotency TFs may be considered as Molecular Versatile actors, and this intrinsic property leads them to exert both pro-self-renewal and pro-lineage commitment functions in closely-related cellular configurations. We detailed this notion below for Oct4 and Nanog.
The TF Oct4 (octamer-binding transcription factor 4), also known as POU5F1, promotes (i) self-renewal but also (ii) exit from pluripotency and (iii) endoderm/mesoderm formation during early development66,71,72,73. The comparative analysis of Oct4 genome-wide binding revealed its redistribution on distinct enhancers with and by specific cofactors in ESCs (Klf4, Nanog, Ctcf) and EpiLCs (Otx2, Zic3) but also during endodermal specification (Sox17), indicating how cofactors modulate Oct4 binding and function74,75,76,77. In addition, Oct4 levels need to be tightly controlled, at both transcriptional and post-transcriptional levels, because slight differences can profoundly alter its function78. Reduced Oct4 expression in ESCs was unexpectedly shown to trigger the establishment of a robust self-renewing state in which the Oct4 protein is redistributed on the genome and increasingly bound to enhancer regulatory regions with Nanog66,71. However, in a primed context in EpiSCs, reduced Oct4 levels destabilises the pluripotent state and promotes the reversion to a naïve state, indicating differential functions for the same Oct4 dosage66. In contrast, at physiological levels, Oct4 triggers the generation of a Nanog-low subpopulation of ESCs that is highly responsive to differentiation cues66, in agreement with findings of exogenous Oct4 expression triggering differentiation79. The Crispr/Cas9 technology, combined with the recent advances of single-cell approaches, will offer a unique opportunity to revisit these concepts by generating knock-in reporter ESCs for multiple pluripotency TFs and analysing the behaviour of single cells harbouring differential endogenous levels of them. In that sense, a recent report indicated that endogenous elevated OCT4 levels strongly biased ESCs towards both neuroectodermal and mesendodermal fates by increasing chromatin accessibility at differentiation-associated enhancers80. Of note, recent evidence also demonstrated the versatility of Oct4 during reprogramming to pluripotency. Silva and colleagues showed first that distinct molecular trajectories to pluripotency required to reach a very precise Oct4 level to be successful81. In line with this view, Schöler’s lab refuted the long-standing dogma of a uniquely essential role of Oct4 during reprogramming. Oct4 overexpression was indeed shown to divert cells from a direct route to pluripotency by triggering aberrant gene activation and also, more importantly, to reduce significantly the developmental potential of iPS cells (Fig. 3)82.
The schematic diagram depicts the different functions exerted by the TF Oct4 depending on its expression level. The expression of Oct4 is tightly regulated because gain and loss-of expression respectively triggering differentiation into primitive endoderm/mesoderm or trophectoderm derivatives in ESCs. Unexpectedly, its endogenous level sustains self-renewal but also primes embryonic stem cells for differentiation while a reduced Oct4 level ensures a robust self-renewal state. A precise Oct4 level is also crucial during reprogramming. Its overexpression triggers aberrant gene inductions while iPS cells generated in absence of Oct4 present an unexpected enhanced developmental potential. PrE: Primitive endoderm, WT level: Wild-type level.
The TF Nanog also harbours strikingly different functions during peri-implantation development. It is well-characterised for its role in the formation of the naïve pluripotent epiblast and the repression of the PrE fate83,84. However, it also represses anterior epiblast identity during mouse gastrulation85. In addition, Nanog is re-expressed in PGCs and required for germline development in vivo86. An elegant study from Surani and colleagues dissected the Molecular Versatility of Nanog. While its exogenous expression triggers naïve ESC self-renewal and resistance to differentiation, it was shown to trigger PGC specification when conducted in primed EpiLCs. Molecularly, the epigenetic resetting occurring in EpiLCs combined with changes in cofactor composition (reduced Sox2 expression), were proposed to allow Nanog to bind and activate enhancers of different sets of genes including the germ cell genes Prdm1 and Prdm1487.
Of note, alternative splicing (AS) constitutes another mechanism that allows to redirect TFs during pluripotency progression. A large program of ESC specific-AS events, controlled by the RNA-binding proteins MBNL1 and MBNL2, was described to orchestrate pluripotency progression88. Blencowe and colleagues identified in particular a conserved AS event in the Foxp1 transcripts, specifically activated in pluripotent cells, that modifies critical amino acid residues within the forkhead domain and alters its DNA-binding specificity. The two resulting FoxP1 isoforms exert opposite effects by activating pluripotency or differentiation genes, respectively88,89,90.
Altogether, these data indicate a battery of mechanisms by which TFs exert different functions during pluripotency progression.
Repurpose epigenetic determinants
Epigenetic determinants play a fundamental role during early development and pluripotency progression, as reviewed elsewhere91. Recent multiomics profiling of the transition from naïve ESCs to EpiLCs revealed for example major waves of protein phosphorylations but also epigenetic modifications92. Moreover, the 3D organization of the chromatin is largely modified during the naïve to primed transition, with the appearance of long-range intra- and inter-chromosomal interactions between H3K27me3 marked genes93. Here we present more specifically how epigenetic regulators are repurposed during pluripotency progression. We focus in particular on enhancer and promoter rewiring as well as Trithorax and Polycomb complexes.
Enhancer reconfigurations
One of the crucial effectors of cell fate transitions is the sequence-dependent binding of TFs to enhancers to regulate gene expression in a cell type-specific manner94,95. This is particularly true in the context of pluripotency progression, where the differences in transcriptional output between naïve ESCs and primed EpiSCs are less dramatic than the alterations observed in chromatin profiles at enhancers74,96,97. In line with this, different classes of enhancers have been defined in ESCs and EpiSCs, based on varying levels of H3K4me1, H3K27ac, H3K36me3 and pSer2/5 forms of RNA polymerase II98. Therefore, a critical challenge for embryonic cells is to ensure a coordinated rewiring of enhancers. We detailed below how Molecular Versatility applies to this process.
As a first example, the FoxD3 TF has emerged as a versatile regulator that orchestrates enhancer reconfigurations during exit from naïve pluripotency and progression to the EpiLC state99. Foxd3 was initially reported as a repressor that decommissions active enhancers of naïve pluripotency genes via the recruitment of Lsd1 and the reduced binding of co-activators like p300 (Fig. 4a). However, in addition to its repressive role on naïve enhancers, FoxD3 was found to exert other dual functions. It was indeed reported to (i) initiate the activity of newly bound EpiLC-specific enhancers while simultaneously (ii) repress their maximal activation. Molecularly, this is achieved by the co-recruitment of the SWI/SNF chromatin remodeling complex ATPase Brg1 to promote nucleosome removal on enhancers but also of histone deacetylases 1/2 to inhibit their optimal activation97 (Fig. 4a).
a The activity of enhancers is controlled by the versatile actions of the FoxD3 transcription factor. In naïve cells, FoxD3 act as a repressor that decommissions active enhancers of naïve pluripotency genes. In primed cells, FoxD3 exerts dual functions by initiating enhancer activity by recruiting Brg1 while simultaneously repressing their maximal activation by recruiting histone deacetylases, illustrating its molecular versatility. b The TF Grhl2 orchestrates an enhancer switching program during pluripotency progression. Grhl2 partitions the pluripotency network and sustain the expression of a subnetwork of epithelial genes during naïve-primed transition. c Promoter switching for the SET gene in ESCs. This switch leads to the generation of two protein isoforms, SETα and SETβ, that play different functions in ESCs. d The subunits of PRC2 dictate different functions for the complex during exit from naïve pluripotency. MTF2 and JARID2 depletion leads to different outcomes during exit from naïve pluripotency. e PRC1 has versatile functions depending on the class of genes it binds to. PRC1-bound active genes showed greater cell-to-cell variation when compared with globally active genes.
Interestingly, a complex enhancer switching program was also evidenced during pluripotency progression for genes that are expressed at identical levels in ESCs and EpiSCs, such as Oct410,100. This type of genes was shown to harbour at least 2 classes of enhancers driving their expression in naïve and primed cellular configurations, respectively. Molecularly, this enhancer switching program is controlled by the redistribution of core (Oct4 and Sox2) and stage-specific (Klf4, Prdm14, and Otx2) pluripotency TFs101. In more detail, an elegant study by Blelloch and colleagues reported how the TF Grhl2 orchestrates such enhancer switching to partition the pluripotency network and sustain the expression of a subnetwork of epithelial genes during pluripotency progression, notably by regulating Cohesin localisation101. As depicted in Fig. 4b, the expression level of the corresponding genes are similar in naïve and primed stem cells but the robustness of the activation is reduced in primed cells to ensure a rapid and efficient silencing later on.
Even if still restricted, these examples highlight how enhancers are rewired during pluripotency progression.
Promoter reconfigurations
Promoter switching constitutes an additional way to generate Molecular Versatility during pluripotency progression. The use of two alternative promoters led for example to the production of two isoforms of the protein SET (SETα and SETβ) that were found to exert differential effects on stem cell self-renewal and commitment. SETα has beneficial effects on proliferation and linker chaperone activities without interfering with differentiation, while SETβ is essential for proper differentiation (Fig. 4c)102. A complex switch between the distal (pLiz) and proximal (pZdbf2) promoters of the Zdbf2 gene has also been described during the naïve-primed transition103. The Zdbf2 gene is expressed transiently in the early embryo (and in naïve ESCs) via its pLiz promoter and then from pZdbf2 in somatic cells. Even if the precise function of the protein remains largely unknown, this transient transcription was found to set up an epigenetic state that is required later on and programs postnatal growth. Indeed, mouse embryos deficient for the distal pLiz promoter develop normally but completely fail to activate Zdbf2 in the postnatal brain and show reduced growth. Therefore, even if this promoter switch appears dispensable for embryogenesis, it signals essential regulatory information in the adult104. At the molecular level, the transient transcription from pLiz promotes de novo DNA methylation upstream of the pZdbf2 promoter, which antagonises Polycomb-mediated repression. Interestingly, the two promoters use similar enhancers but their respective activity in ESCs and EpiLCs is controlled by contacts mediated by the CTCF protein. The alternative usage of enhancers/promoters represents an elegant example of Molecular Versatility that contributes to sustain stemness but prepares the genome for a robust and coordinated progression towards gastrulation.
Trithorax (TrxG) complexes
TrxG complexes mediate the deposition of the active histone mark H3K4 methylation via their catalytic subunits, the histone methyltransferases SET/MLL105. TrxG complexes display versatile functions in ESCs, namely coordinating (i) self-renewal or (ii) lineage commitment, depending on the composition of subunits. The core member subunit Wdr5 was described to promote naïve ESCs self-renewal by maintaining global H3K4me3 levels and by cooperating with the Oct4/Nanog/Sox2 circuitry, sharing target genes and regulatory functions106. Ash2l, another core subunit, is also required for pluripotency: Ash2l knockdown in ESCs leads to differentiation and derepression of mesodermal markers107. Conversely, the depletion of the subunit Dpy-30 does not impact ESCs self-renewal but rather impairs their differentiation, particularly along the neural lineage. This phenotype is largely mediated by the erasure of the H3K4me3 mark at bivalent promoters that induces the dissolution of their poised state108. Interestingly, the MLL1 subunit was on the contrary reported to maintain the identity of primed EpiSCs. MLL1 inhibition leads to a global redistribution of the H3K4me1 mark and therefore to a profound rewiring of the enhancer landscape, leading to the spontaneous reversion of primed EpiSCs into naïve ESCs109.
Polycomb repressive complexes
The PRC complexes (PRC2 and PRC1), formed by Polycomb group proteins, play essential roles in pluripotency and development by mediating chromatin modifications. PRC2 is recruited to specific genomic locations and catalyzes deposition of H3K27me3, which in turn recruits PRC1, resulting in the generation of H2AK119ub1 and transcriptional repression. In ESCs, PRC1/2 co-occupy indeed regions marked with H3K27me3, with a large proportion of these sites being proximal to genes of lineage commitment110,111. Accordingly, PRC2- (Eed KO) and PRC1- (Ring1A/B KO) deficient ESCs showed increased expression of differentiation genes110,112. However, PRC function is far more broad and versatile during pluripotency progression than initially established113.
First, recent integrative proteomic profiling of ground state ESCs revealed a critical function for PRC2 in globally shaping the pluripotent epigenome to avoid the acquisition of primed features such as DNA methylation114. In ground state ESCs, PRC2 and H3K27me3 are indeed widely distributed on both euchromatin and heterochromatin, when compared with Serum/LIF ESCs.
Second, a recent body of work demonstrated the versatile functions exerted by the different subunits of PRC2. By combining single-cell transcriptomics and embryoid body formation, Loh et al. showed that the depletion of the two distinct PRC2 subunits MTF2 and JARID2 led to different phenotypes during exit to naïve pluripotency. MTF2 absence enhanced and accelerated differentiation towards the three lineages, as expected, but JARID2 null ESCs struggled to turn off pluripotency genes and were predominantly converted into early differentiating precursors, with reduced efficiency towards the mesendoderm lineages (Fig. 4d)115.
Moreover, another study highlighted the critical role of PRC2 in promoting the specification of the ectoderm in rodent and human primed stem cells116. Targeting PRC2 genes in human ESCs led to pluripotency loss and to spontaneous differentiation towards a meso-endoderm fate. The attempts to convert mouse PRC2-deficient naïve ESCs into primed EpiSCs failed and led to similar differentiation defects as human cells. Even if the precise molecular mechanisms remain to be elucidated, these versatile functions of PRC2 might be attributable to (i) the coordinated relocation of various PRC2 complexes and to (ii) different thresholds for activation of various classes of genes.
PRC1 was also reported to promote both (i) self-renewal and (ii) lineage commitment in ESCs, with reports highlighting the importance of the composition of its subunits in these versatile functions117,118. In naïve ESC, Cbx7, which is the main PRC1-associated expressed Cbx protein, is required to maintain the undifferentiated state in a robust manner. This Cbx7-containing PRC1 complex represses the transcription of differentiation genes and of the other subunits Cbx2 and Cbx4. Once induced during differentiation, Cbx2- and Cbx4-containing complexes are redistributed to repress a large set of genes, including pluripotency-related loci. In addition, Cbx2/4 deficiency was demonstrated to skew in vivo teratoma differentiation towards the endodermal fate, indicating a specific function in germ layer formation, as also revealed for the other PRC1 subunits Ring1B and Pcgf6119,120. Thus, Cbx proteins confer distinct target selectivity and functions to the canonical PRC1 complex. In addition, variant PRC1 complexes, that are recruited to target genes mainly independently of PRC2 and H3K27me3, were found recently to be mainly responsible for gene repression in ESCs121,122.
Moreover, although PRCs are known to exert a repressive effect, interestingly, the cohort of PRC-bound genes contains not only silent genes, but also genes with intermediate and high expression123. Recent single-cell RNA-seq analyses showed that active genes bound by PRC in ESCs are subjected to a switch between active and repressed states and harbour therefore greater cell-to-cell variation than classical active genes. This phenomenon, by favoring stochastic expression of genes, may play a key role in stem cell physiology (Fig. 4e)124,125. In the same line, PRC1 complexes containing Cbx8 are recruited transiently on differentiation genes to facilitate their activation126. Similarly, the PRC1 subunit Mel18, crucial for the repression of differentiation genes in ESCs, facilitates the expression of TFs essential for cardiac differentiation in progenitors127. Altogether, these findings support the versatile functions and modes of action of PRC during early development.
A tight control of molecular versatility
Altogether, the previous sections highlighted how molecules are accurately redeployed to exert different, sometimes opposite, functions in closely-related stem cell configurations. However, in order for the changes to occur at the correct time and place, Molecular Versatility must be tightly regulated and connected with the microenvironment to ensure channelling and robustness of pluripotency progression4. Signalling pathways are known to be directly connected to pluripotency TFs. Effectors such as Stat3 and Smad2/3 co-occupy the ESC genome with the TFs Oct4, Nanog and Sox2, providing a direct rationale for the connection between extrinsic cues, signalling and the TF network128,129,130. However, certain signalling pathways were recently found to directly control the Molecular Versatility of TFs. For instance, the TF Etv5 was reported to undergo a switch in its function depending on ERK1/2 signalling activity. Indeed, Etv5 supports ESC self-renewal when ERK1/2 is inhibited but promotes exit from naïve pluripotency when ERK1/2 is active17. As another example, the EMT-TF Snail1 was found to control metabolic cascades in ESCs in a WNT-independent manner. However, during ESC commitment, Snail1 is induced by WNT to regulate exit from primed pluripotency and neuroectodermal fate emergence131.
In order to be precisely regulated, the activity of versatile factors was also found to be controlled by multiple signals. As a first example, we reported that the Netrin-1 signalling pathway regulates naïve pluripotency by modulating the responsiveness of embryonic cells to both FGF and WNT signals25,132. This dual function is achieved via the induction of complex signalling cascades downstream of the two Netrin-1 receptors Unc5b and Neo1. However, the expression of the Netrin-1 transcript itself is also directly and antagonistically controlled by the WNT and FGF/MAPK pathways, positioning this molecule as an integrated molecular sensor for the embryonic cell.
The activity of basic helix-loop-helix (bHLH) TFs is also controlled by multiple signals during pluripotency progression. For instance, Tcf15 was described to prime pluripotent cells for differentiation in response to FGF/MAPK signalling133. However, while Tcf15 transcription is directly induced by FGF/MAPK, its biological activity is suppressed by Id proteins that are under the control of BMP signalling22. Therefore, Tcf15 is able to molecularly sense multiple external cues (FGF/BMP) to finely tune its activity and to confer accuracy to the naïve-primed transition. Consistently, Tcf15 expression is transient in the early embryo, readily detectable at E4.5 but turned off by E5.5. The bHLH TF Tfe3, that promotes naïve pluripotency, is rapidly excluded from the nucleus of ESCs by the mTORC1 and lysosome signalling134,135, highlighting an alternative way to precisely integrate signalling cues.
Another elegant study from Lowell et al. deciphered how the BHLH TF Id1 preserves pluripotency and avoids premature differentiation during the naïve-primed transition by being multiply regulated. Id1 expression is induced when embryonic cells lose Nanog expression but have not yet acquired Nodal activity. The Id1 protein suppresses FGF/MAPK signalling in order to protect these Nanog-negative cells from differentiation. Then, once Nodal activity is operating in these transitioning cells, Nodal itself suppresses Id1 expression and leads to a rise in to FGF activity contributing to sustaining pluripotency in primed cells136.
Concluding remarks
This perspective presents converging findings demonstrating that a large repertoire of molecules are repurposed to exert different, sometimes opposite, functions in closely-related stem cell configurations during pluripotency progression. We propose to regroup these multiple molecular mechanisms by which signalling pathways, TFs and epigenetic regulators are rapidly rewired with the notion of Molecular Versatility, as summarised in Fig. 5. We believe that these redeployments, by acting in concert with profound transcriptomic and epigenomic rewirings, contribute to ensure a robust and coordinated programming of embryonic stem cell fate in response to extrinsic signals. In addition, we propose that Molecular Versatility may confer a certain degree of reversibility and flexibility to the developmental transition processes by involving molecules already present within the embryonic cells. Because the examples used in this Perspective are mainly based on the comparison of naïve ESCs and primed EpiLCs/EpiSCs, it remains to be investigated whether and how Molecular Versatility takes place dynamically in truly transitioning cells. The recent capture of intermediate states such as Rosette and Formative stem cells (RSC and FS respectively), combined with the power of computational biology, will certainly help in better defining these relatively underexplored layers of regulation. Moreover, the advances in the development of single cell-based approaches will also permit to address precisely how these mechanisms take place in transitioning cells. In that sense, pluripotency progression provides an amenable experimental system for dissection of stem cell fate transition paradigm, that will surely bring highly valuable concepts for the biology of embryonic but also adult normal and pathological stem cells.
References
Zhu, M. & Zernicka-Goetz, M. Principles of Self-Organization of the Mammalian Embryo. Cell 183, 1467–1478 (2020).
Kurimoto, K. et al. Quantitative Dynamics of Chromatin Remodeling during Germ Cell Specification from Mouse Embryonic Stem Cells. Cell Stem Cell 16, 517–532 (2015).
Kalkan, T. & Smith, A. Mapping the route from naive pluripotency to lineage specification. Philos. Trans. R Soc. Lond. B Biol. Sci. 369, https://doi.org/10.1098/rstb.2013.0540 (2014).
Waddington, C. H. Canalization of development and genetic assimilation of acquired characters. Nature 183, 1654–1655 (1959).
Nichols, J. & Smith, A. Naive and primed pluripotent states. Cell Stem Cell 4, 487–492 (2009).
Pera, M. F. & Rossant, J. The exploration of pluripotency space: Charting cell state transitions in peri-implantation development. Cell Stem Cell 28, 1896–1906 (2021).
Rossant, J. & Tam, P. P. Blastocyst lineage formation, early embryonic asymmetries and axis patterning in the mouse. Development 136, 701–713 (2009).
Smith, A. Formative pluripotency: the executive phase in a developmental continuum. Development 144, 365–373 (2017).
Evans, M. J. & Kaufman, M. H. Establishment in culture of pluripotential cells from mouse embryos. Nature 292, 154–156 (1981).
Tesar, P. J. et al. New cell lines from mouse epiblast share defining features with human embryonic stem cells. Nature 448, 196–199 (2007).
Kojima, Y. et al. The transcriptional and functional properties of mouse epiblast stem cells resemble the anterior primitive streak. Cell Stem Cell 14, 107–120 (2014).
Hayashi, K., Ohta, H., Kurimoto, K., Aramaki, S. & Saitou, M. Reconstitution of the mouse germ cell specification pathway in culture by pluripotent stem cells. Cell 146, 519–532 (2011).
Neagu, A. et al. In vitro capture and characterization of embryonic rosette-stage pluripotency between naive and primed states. Nat. Cell Biol. 22, 534–545 (2020).
Kinoshita, M. et al. Capture of Mouse and Human Stem Cells with Features of Formative Pluripotency. Cell Stem Cell 28, 453–471 e458 (2021).
Yu, L. et al. Derivation of Intermediate Pluripotent Stem Cells Amenable to Primordial Germ Cell Specification. Cell Stem Cell 28, 550–567.e512 (2021).
Wang, X. et al. Formative pluripotent stem cells show features of epiblast cells poised for gastrulation. Cell Res. 31, 526–541 (2021).
Kalkan, T. et al. Complementary Activity of ETV5, RBPJ, and TCF3 Drives Formative Transition from Naive Pluripotency. Cell Stem Cell 24, 785–801.e787 (2019).
Dunn, S. J., Martello, G., Yordanov, B., Emmott, S. & Smith, A. G. Defining an essential transcription factor program for naive pluripotency. Science 344, 1156–1160 (2014).
Lackner, A. et al. Cooperative genetic networks drive embryonic stem cell transition from naive to formative pluripotency. EMBO J. 40, e105776 (2021).
Weinberger, L., Ayyash, M., Novershtern, N. & Hanna, J. H. Dynamic stem cell states: naive to primed pluripotency in rodents and humans. Nat. Rev. Mol. Cell Biol. 17, 155–169 (2016).
Nichols, J., Chambers, I., Taga, T. & Smith, A. Physiological rationale for responsiveness of mouse embryonic stem cells to gp130 cytokines. Development 128, 2333–2339 (2001).
Ying, Q. L., Nichols, J., Chambers, I. & Smith, A. BMP induction of Id proteins suppresses differentiation and sustains embryonic stem cell self-renewal in collaboration with STAT3. Cell 115, 281–292 (2003).
ten Berge, D. et al. Embryonic stem cells require Wnt proteins to prevent differentiation to epiblast stem cells. Nat. Cell Biol. 13, 1070–1075 (2011).
Ying, Q. L. et al. The ground state of embryonic stem cell self-renewal. Nature 453, 519–523 (2008).
Huyghe, A. et al. Netrin-1 promotes naive pluripotency through Neo1 and Unc5b co-regulation of Wnt and MAPK signalling. Nat. Cell Biol. 22, 389–400 (2020).
Yamanaka, Y., Lanner, F. & Rossant, J. FGF signal-dependent segregation of primitive endoderm and epiblast in the mouse blastocyst. Development 137, 715–724 (2010).
Kunath, T. et al. FGF stimulation of the Erk1/2 signalling cascade triggers transition of pluripotent embryonic stem cells from self-renewal to lineage commitment. Development 134, 2895–2902 (2007).
Kang, M., Garg, V. & Hadjantonakis, A. K. Lineage Establishment and Progression within the Inner Cell Mass of the Mouse Blastocyst Requires FGFR1 and FGFR2. Dev. Cell 41, 496–510 e495 (2017).
Brons, I. G. et al. Derivation of pluripotent epiblast stem cells from mammalian embryos. Nature 448, 191–195 (2007).
Boroviak, T. et al. Lineage-Specific Profiling Delineates the Emergence and Progression of Naive Pluripotency in Mammalian Embryogenesis. Dev. Cell 35, 366–382 (2015).
Ohnishi, Y. et al. Cell-to-cell expression variability followed by signal reinforcement progressively segregates early mouse lineages. Nat. Cell Biol. 16, 27–37 (2014).
Molotkov, A., Mazot, P., Brewer, J. R., Cinalli, R. M. & Soriano, P. Distinct Requirements for FGFR1 and FGFR2 in Primitive Endoderm Development and Exit from Pluripotency. Dev. Cell 41, 511–526 e514 (2017).
Simon, C. S., Rahman, S., Raina, D., Schroter, C. & Hadjantonakis, A. K. Live Visualization of ERK Activity in the Mouse Blastocyst Reveals Lineage-Specific Signaling Dynamics. Dev. Cell 55, 341–353.e345 (2020).
Lanner, F. et al. Heparan sulfation-dependent fibroblast growth factor signaling maintains embryonic stem cells primed for differentiation in a heterogeneous state. Stem Cells 28, 191–200 (2010).
Shimokawa, K. et al. Cell surface heparan sulfate chains regulate local reception of FGF signaling in the mouse embryo. Dev. Cell 21, 257–272 (2011).
Vasudevan, H. N., Mazot, P., He, F. & Soriano, P. Receptor tyrosine kinases modulate distinct transcriptional programs by differential usage of intracellular pathways. Elife 4, https://doi.org/10.7554/eLife.07186 (2015).
Lanner, F. & Rossant, J. The role of FGF/Erk signaling in pluripotent cells. Development 137, 3351–3360 (2010).
Chen, H. et al. Erk signaling is indispensable for genomic stability and self-renewal of mouse embryonic stem cells. Proc. Natl Acad. Sci. USA 112, E5936–E5943 (2015).
Choi, J. et al. Prolonged Mek1/2 suppression impairs the developmental potential of embryonic stem cells. Nature 548, 219–223 (2017).
Hamilton, W. B. et al. Dynamic lineage priming is driven via direct enhancer regulation by ERK. Nature 575, 355–360 (2019).
Sato, N., Meijer, L., Skaltsounis, L., Greengard, P. & Brivanlou, A. H. Maintenance of pluripotency in human and mouse embryonic stem cells through activation of Wnt signaling by a pharmacological GSK-3-specific inhibitor. Nat. Med. 10, 55–63 (2004).
Kurek, D. et al. Endogenous WNT signals mediate BMP-induced and spontaneous differentiation of epiblast stem cells and human embryonic stem cells. Stem Cell Rep. 4, 114–128 (2015).
Atlasi, Y. et al. Wnt signaling regulates the lineage differentiation potential of mouse embryonic stem cells through Tcf3 down-regulation. PLoS Genet. 9, e1003424 (2013).
Shy, B. R. et al. Regulation of Tcf7l1 DNA binding and protein stability as principal mechanisms of Wnt/beta-catenin signaling. Cell Rep. 4, 1–9 (2013).
Zhang, M. et al. beta-Catenin safeguards the ground state of mousepluripotency by strengthening the robustness of the transcriptional apparatus. Sci. Adv. 6, eaba1593 (2020).
Lyashenko, N. et al. Differential requirement for the dual functions of beta-catenin in embryonic stem cell self-renewal and germ layer formation. Nat. Cell Biol. 13, 753–761 (2011).
Singh, A. M. et al. Signaling network crosstalk in human pluripotent cells: a Smad2/3-regulated switch that controls the balance between self-renewal and differentiation. Cell Stem Cell 10, 312–326 (2012).
De Jaime-Soguero, A. et al. Wnt/Tcf1 pathway restricts embryonic stem cell cycle through activation of the Ink4/Arf locus. PLoS Genet. 13, e1006682 (2017).
Chatterjee, S. S. et al. Inhibition of beta-catenin-TCF1 interaction delays differentiation of mouse embryonic stem cells. J. Cell Biol. 211, 39–51 (2015).
van Amerongen, R., Fuerer, C., Mizutani, M. & Nusse, R. Wnt5a can both activate and repress Wnt/beta-catenin signaling during mouse embryonic development. Dev. Biol. 369, 101–114 (2012).
Schuijers, J. et al. Ascl2 acts as an R-spondin/Wnt-responsive switch to control stemness in intestinal crypts. Cell Stem Cell 16, 158–170 (2015).
Pauklin, S. & Vallier, L. Activin/Nodal signalling in stem cells. Development 142, 607–619 (2015).
Ashida, Y., Nakajima-Koyama, M., Hirota, A., Yamamoto, T. & Nishida, E. Activin A in combination with ERK1/2 MAPK pathway inhibition sustains propagation of mouse embryonic stem cells. Genes Cells 22, 189–202 (2017).
Ogawa, K. et al. Activin-Nodal signaling is involved in propagation of mouse embryonic stem cells. J. Cell Sci. 120, 55–65 (2007).
Lee, K. L. et al. Graded Nodal/Activin signaling titrates conversion of quantitative phospho-Smad2 levels into qualitative embryonic stem cell fate decisions. PLoS Genet. 7, e1002130 (2011).
Serafini, T. et al. The netrins define a family of axon outgrowth-promoting proteins homologous to C. elegans UNC-6. Cell 78, 409–424 (1994).
Rajagopalan, S. et al. Neogenin mediates the action of repulsive guidance molecule. Nat. Cell Biol. 6, 756–762 (2004).
Llimargas, M. & Lawrence, P. A. Seven Wnt homologues in Drosophila: a case study of the developing tracheae. Proc. Natl Acad. Sci. USA 98, 14487–14492 (2001).
Edson, M. A. et al. Granulosa cell-expressed BMPR1A and BMPR1B have unique functions in regulating fertility but act redundantly to suppress ovarian tumor development. Mol. Endocrinol. 24, 1251–1266 (2010).
Robinson, R. A. et al. Simultaneous binding of Guidance Cues NET1 and RGM blocks extracellular NEO1 signaling. Cell 184, 2103–2120 e2131 (2021).
Antebi, Y. E. et al. Combinatorial Signal Perception in the BMP Pathway. Cell 170, 1184–1196 e1124 (2017).
Su, C. J. et al. Ligand-receptor promiscuity enables cellular addressing. Cell Syst. 13, 408–425 e412 (2022).
Corsinotti, A. et al. Distinct SoxB1 networks are required for naive and primed pluripotency. Elife 6, https://doi.org/10.7554/eLife.27746 (2017).
Lu, R., Yang, A. & Jin, Y. Dual functions of T-box 3 (Tbx3) in the control of self-renewal and extraembryonic endoderm differentiation in mouse embryonic stem cells. J. Biol. Chem. 286, 8425–8436 (2011).
Herchcovici Levy, S. et al. Esrrb is a cell-cycle-dependent associated factor balancing pluripotency and XEN differentiation. Stem Cell Rep. 17, 1334–1350 (2022).
Karwacki-Neisius, V. et al. Reduced Oct4 expression directs a robust pluripotent state with distinct signaling activity and increased enhancer occupancy by Oct4 and Nanog. Cell Stem Cell 12, 531–545 (2013).
Thomson, M. et al. Pluripotency factors in embryonic stem cells regulate differentiation into germ layers. Cell 145, 875–889 (2011).
Loh, K. M. & Lim, B. A precarious balance: pluripotency factors as lineage specifiers. Cell Stem Cell 8, 363–369 (2011).
Silva, J. & Smith, A. Capturing pluripotency. Cell 132, 532–536 (2008).
Hu, M. et al. Multilineage gene expression precedes commitment in the hemopoietic system. Genes Dev. 11, 774–785 (1997).
Radzisheuskaya, A. et al. A defined Oct4 level governs cell state transitions of pluripotency entry and differentiation into all embryonic lineages. Nat. Cell Biol. 15, 579–590 (2013).
Osorno, R. et al. The developmental dismantling of pluripotency is reversed by ectopic Oct4 expression. Development 139, 2288–2298 (2012).
Frum, T. et al. Oct4 cell-autonomously promotes primitive endoderm development in the mouse blastocyst. Dev. Cell 25, 610–622 (2013).
Buecker, C. et al. Reorganization of enhancer patterns in transition from naive to primed pluripotency. Cell Stem Cell 14, 838–853 (2014).
Yang, S. H. et al. Otx2 and Oct4 drive early enhancer activation during embryonic stem cell transition from naive pluripotency. Cell Rep. 7, 1968–1981 (2014).
Galonska, C., Ziller, M. J., Karnik, R. & Meissner, A. Ground State Conditions Induce Rapid Reorganization of Core Pluripotency Factor Binding before Global Epigenetic Reprogramming. Cell Stem Cell 17, 462–470 (2015).
Aksoy, I. et al. Oct4 switches partnering from Sox2 to Sox17 to reinterpret the enhancer code and specify endoderm. EMBO J. 32, 938–953 (2013).
Sohn, E. J., Moon, H. J., Lim, J. K., Kim, D. S. & Kim, J. H. Regulation of the protein stability and transcriptional activity of OCT4 in stem cells. Adv. Biol. Regul. 79, 100777 (2020).
Niwa, H., Miyazaki, J. & Smith, A. G. Quantitative expression of Oct-3/4 defines differentiation, dedifferentiation or self-renewal of ES cells. Nat. Genet. 24, 372–376 (2000).
Strebinger, D. et al. Endogenous fluctuations of OCT4 and SOX2 bias pluripotent cell fate decisions. Mol. Syst. Biol. 15, e9002 (2019).
Stuart, H. T. et al. Distinct Molecular Trajectories Converge to Induce Naive Pluripotency. Cell Stem Cell 25, 388–406 e388 (2019).
Velychko, S. et al. Excluding Oct4 from Yamanaka Cocktail Unleashes the Developmental Potential of iPSCs. Cell Stem Cell 25, 737–753.e734 (2019).
Silva, J. et al. Nanog is the gateway to the pluripotent ground state. Cell 138, 722–737 (2009).
Singh, A. M., Hamazaki, T., Hankowski, K. E. & Terada, N. A heterogeneous expression pattern for Nanog in embryonic stem cells. Stem Cells 25, 2534–2542 (2007).
Barral, A. et al. Nanog regulates Pou3f1 expression at the exit from pluripotency during gastrulation. Biol. Open 8, https://doi.org/10.1242/bio.046367 (2019).
Chambers, I. et al. Nanog safeguards pluripotency and mediates germline development. Nature 450, 1230–1234 (2007).
Murakami, K. et al. NANOG alone induces germ cells in primed epiblast in vitro by activation of enhancers. Nature 529, 403–407 (2016).
Han, H. et al. MBNL proteins repress ES-cell-specific alternative splicing and reprogramming. Nature 498, 241–245 (2013).
Gabut, M. et al. An alternative splicing switch regulates embryonic stem cell pluripotency and reprogramming. Cell 147, 132–146 (2011).
Cirera-Salinas, D. et al. Noncanonical function of DGCR8 controls mESC exit from pluripotency. J. Cell Biol. 216, 355–366 (2017).
Schlesinger, S. & Meshorer, E. Open Chromatin, Epigenetic Plasticity, and Nuclear Organization in Pluripotency. Dev. Cell 48, 135–150 (2019).
Yang, P. et al. Multi-omic Profiling Reveals Dynamics of the Phased Progression of Pluripotency. Cell Syst. 8, 427–445.e410 (2019).
Joshi, O. et al. Dynamic Reorganization of Extremely Long-Range Promoter-Promoter Interactions between Two States of Pluripotency. Cell Stem Cell 17, 748–757 (2015).
Calo, E. & Wysocka, J. Modification of enhancer chromatin: what, how, and why. Mol. Cell 49, 825–837 (2013).
Shlyueva, D., Stampfel, G. & Stark, A. Transcriptional enhancers: from properties to genome-wide predictions. Nat. Rev. Genet. 15, 272–286 (2014).
Choi, H. W. et al. Distinct Enhancer Activity of Oct4 in Naive and Primed Mouse Pluripotency. Stem Cell Rep. 7, 911–926 (2016).
Krishnakumar, R. et al. FOXD3 Regulates Pluripotent Stem Cell Potential by Simultaneously Initiating and Repressing Enhancer Activity. Cell Stem Cell 18, 104–117 (2016).
Zentner, G. E., Tesar, P. J. & Scacheri, P. C. Epigenetic signatures distinguish multiple classes of enhancers with distinct cellular functions. Genome Res. 21, 1273–1283 (2011).
Respuela, P. et al. Foxd3 Promotes Exit from Naive Pluripotency through Enhancer Decommissioning and Inhibits Germline Specification. Cell Stem Cell 18, 118–133 (2016).
Factor, D. C. et al. Epigenomic comparison reveals activation of “seed” enhancers during transition from naive to primed pluripotency. Cell Stem Cell 14, 854–863 (2014).
Chen, A. F. et al. GRHL2-Dependent Enhancer Switching Maintains a Pluripotent Stem Cell Transcriptional Subnetwork after Exit from Naive Pluripotency. Cell Stem Cell 23, 226–238 e224 (2018).
Edupuganti, R. R. et al. Alternative SET/TAFI Promoters Regulate Embryonic Stem Cell Differentiation. Stem Cell Rep. 9, 1291–1303 (2017).
Greenberg, M., Teissandier, A., Walter, M., Noordermeer, D. & Bourc’his, D. Dynamic enhancer partitioning instructs activation of a growth-related gene during exit from naive pluripotency. Elife 8, https://doi.org/10.7554/eLife.44057 (2019).
Greenberg, M. V. et al. Transient transcription in the early embryo sets an epigenetic state that programs postnatal growth. Nat. Genet. 49, 110–118 (2017).
Schuettengruber, B., Martinez, A. M., Iovino, N. & Cavalli, G. Trithorax group proteins: switching genes on and keeping them active. Nat. Rev. Mol. Cell Biol. 12, 799–814 (2011).
Ang, Y. S. et al. Wdr5 mediates self-renewal and reprogramming via the embryonic stem cell core transcriptional network. Cell 145, 183–197 (2011).
Wan, M. et al. The trithorax group protein Ash2l is essential for pluripotency and maintaining open chromatin in embryonic stem cells. J. Biol. Chem. 288, 5039–5048 (2013).
Jiang, H. et al. Role for Dpy-30 in ES cell-fate specification by regulation of H3K4 methylation within bivalent domains. Cell 144, 513–525 (2011).
Zhang, H. et al. MLL1 Inhibition Reprograms Epiblast Stem Cells to Naive Pluripotency. Cell Stem Cell 18, 481–494 (2016).
Boyer, L. A. et al. Polycomb complexes repress developmental regulators in murine embryonic stem cells. Nature 441, 349–353 (2006).
Ku, M. et al. Genomewide analysis of PRC1 and PRC2 occupancy identifies two classes of bivalent domains. PLoS Genet. 4, e1000242 (2008).
Endoh, M. et al. Polycomb group proteins Ring1A/B are functionally linked to the core transcriptional regulatory circuitry to maintain ES cell identity. Development 135, 1513–1524 (2008).
Camahort, R. & Cowan, C. A. Cbx proteins help ESCs walk the line between self-renewal and differentiation. Cell Stem Cell 10, 4–6 (2012).
van Mierlo, G. et al. Integrative Proteomic Profiling Reveals PRC2-Dependent Epigenetic Crosstalk Maintains Ground-State Pluripotency. Cell Stem Cell 24, 123–137.e128 (2019).
Loh, C. H., van Genesen, S., Perino, M., Bark, M. R. & Veenstra, G. J. C. Loss of PRC2 subunits primes lineage choice during exit of pluripotency. Nat. Commun. 12, 6985 (2021).
Shan, Y. et al. PRC2 specifies ectoderm lineages and maintains pluripotency in primed but not naive ESCs. Nat. Commun. 8, 672 (2017).
O’Loghlen, A. et al. MicroRNA regulation of Cbx7 mediates a switch of Polycomb orthologs during ESC differentiation. Cell Stem Cell 10, 33–46 (2012).
Morey, L. et al. Nonoverlapping functions of the Polycomb group Cbx family of proteins in embryonic stem cells. Cell Stem Cell 10, 47–62 (2012).
Leeb, M. et al. Polycomb complexes act redundantly to repress genomic repeats and genes. Genes Dev. 24, 265–276 (2010).
Zdzieblo, D. et al. Pcgf6, a polycomb group protein, regulates mesodermal lineage differentiation in murine ESCs and functions in iPS reprogramming. Stem Cells 32, 3112–3125 (2014).
Zepeda-Martinez, J. A. et al. Parallel PRC2/cPRC1 and vPRC1 pathways silence lineage-specific genes and maintain self-renewal in mouse embryonic stem cells. Sci. Adv. 6, eaax5692 (2020).
Fursova, N. A. et al. Synergy between Variant PRC1 Complexes Defines Polycomb-Mediated Gene Repression. Mol. Cell 74, 1020–1036.e1028 (2019).
Brookes, E. et al. Polycomb associates genome-wide with a specific RNA polymerase II variant, and regulates metabolic genes in ESCs. Cell Stem Cell 10, 157–170 (2012).
Kar, G. et al. Flipping between Polycomb repressed and active transcriptional states introduces noise in gene expression. Nat. Commun. 8, 36 (2017).
Eldar, A. & Elowitz, M. B. Functional roles for noise in genetic circuits. Nature 467, 167–173 (2010).
Creppe, C., Palau, A., Malinverni, R., Valero, V. & Buschbeck, M. A Cbx8-containing polycomb complex facilitates the transition to gene activation during ES cell differentiation. PLoS Genet. 10, e1004851 (2014).
Morey, L. et al. Polycomb Regulates Mesoderm Cell Fate-Specification in Embryonic Stem Cells through Activation and Repression Mechanisms. Cell Stem Cell 17, 300–315 (2015).
Chen, X. et al. Integration of external signaling pathways with the core transcriptional network in embryonic stem cells. Cell 133, 1106–1117 (2008).
Kagey, M. H. et al. Mediator and cohesin connect gene expression and chromatin architecture. Nature 467, 430–435 (2010).
Senft, A. D. et al. Combinatorial Smad2/3 Activities Downstream of Nodal Signaling Maintain Embryonic/Extra-Embryonic Cell Identities during Lineage Priming. Cell Rep. 24, 1977–1985 e1977 (2018).
Lin, Y. et al. Snail1-dependent control of embryonic stem cell pluripotency and lineage commitment. Nat. Commun. 5, 3070 (2014).
Ozmadenci, D. et al. Netrin-1 regulates somatic cell reprogramming and pluripotency maintenance. Nat. Commun. 6, 7398 (2015).
Davies, O. R. et al. Tcf15 primes pluripotent cells for differentiation. Cell Rep. 3, 472–484 (2013).
Betschinger, J. et al. Exit from pluripotency is gated by intracellular redistribution of the bHLH transcription factor Tfe3. Cell 153, 335–347 (2013).
Villegas, F. et al. Lysosomal Signaling Licenses Embryonic Stem Cell Differentiation via Inactivation of Tfe3. Cell Stem Cell 24, 257–270 e258 (2019).
Malaguti, M., Migueles, R. P., Blin, G., Lin, C. Y. & Lowell, S. Id1 Stabilizes Epiblast Identity by Sensing Delays in Nodal Activation and Adjusting the Timing of Differentiation. Dev. Cell 50, 462–477 e465 (2019).
Kinoshita, M. et al. Capture of Mouse and Human Stem Cells with Features of Formative Pluripotency. Cell Stem Cell https://doi.org/10.1016/j.stem.2020.11.005 (2020).
Mills, J. C., Stanger, B. Z. & Sander, M. Nomenclature for cellular plasticity: are the terms as plastic as the cells themselves? EMBO J. 38, e103148 (2019).
Acknowledgements
We are grateful to Brigitte Manship for proofreading of the paper. This work was supported by institutional grants from INSERM/CNRS, La ligue contre le cancer nationale et régionale (F.L.), Labex Dev2Can (F.L), Fondation Bettencourt Schueller (Impulscience)(F.L.), INCa (F.L.), ANR (F.L.), Labex DEV2CAN (F.L.), Fondation ARC (F.L., A.H.), Centre Léon Bérard (F.L., A.H.), Fondation pour la recherche médicale (N.C., G.F., F.L.) and Institut Convergence PLAsCAN, ANR-17-CONV-0002.
Author information
Authors and Affiliations
Contributions
G.F. and A.H. worked equally on specific paragraphs of the Perspective. N.C. edited the document and generated the Figures. F.L. designed and supervised the work.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks the anonymous reviewers for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Furlan, G., Huyghe, A., Combémorel, N. et al. Molecular versatility during pluripotency progression. Nat Commun 14, 68 (2023). https://doi.org/10.1038/s41467-022-35775-4
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41467-022-35775-4
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.