Abstract
A distinctive feature of prokaryotic gene expression is the absence of 5′-capped RNA. In eukaryotes, 5′,5′-triphosphate-linked 7-methylguanosine protects messenger RNA from degradation and modulates maturation, localization and translation1. Recently, the cofactor nicotinamide adenine dinucleotide (NAD) was reported as a covalent modification of bacterial RNA2. Given the central role of NAD in redox biochemistry, posttranslational protein modification and signalling3,4, its attachment to RNA indicates that there are unknown functions of RNA in these processes and undiscovered pathways in RNA metabolism and regulation. The unknown identity of NAD-modified RNAs has so far precluded functional analyses. Here we identify NAD-linked RNAs from bacteria by chemo-enzymatic capture and next-generation sequencing (NAD captureSeq). Among those identified, specific regulatory small RNAs (sRNAs) and sRNA-like 5′-terminal fragments of certain mRNAs are particularly abundant. Analogous to a eukaryotic cap, 5′-NAD modification is shown in vitro to stabilize RNA against 5′-processing by the RNA-pyrophosphohydrolase RppH5 and against endonucleolytic cleavage by ribonuclease (RNase) E6. The nudix phosphohydrolase NudC7 decaps NAD-RNA and thereby triggers RNase-E-mediated RNA decay, while being inactive against triphosphate-RNA. In vivo, ∼13% of the abundant sRNA RNAI is NAD-capped in the presence, and ∼26% in the absence, of functional NudC. To our knowledge, this is the first description of a cap-like structure and a decapping machinery in bacteria.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Accession codes
Data deposits
NGS data have been deposited at http://www.geneprof.org under accession numbers gpXP_001108, gpXP_001123 and gpXP_001153.
References
Topisirovic, I., Svitkin, Y. V., Sonenberg, N. & Shatkin, A. J. Cap and cap-binding proteins in the control of gene expression. RNA 2, 277–298 (2011)
Chen, Y. G., Kowtoniuk, W. E., Agarwal, I., Shen, Y. & Liu, D. R. LC/MS analysis of cellular RNA reveals NAD-linked RNA. Nature Chem. Biol. 5, 879–881 (2009)
Houtkooper, R. H., Cantó, C., Wanders, R. J. & Auwerx, J. The secret life of NAD+: an old metabolite controlling new metabolic signaling pathways. Endocr. Rev. 31, 194–223 (2010)
Oppenheimer, N. J. NAD+ and NADP+ as prosthetic groups for enzymes. eLS http://dx.doi/org/10.1002/9780470015902.a0000637.pub2 (2010)
Deana, A., Celesnik, H. & Belasco, J. G. The bacterial enzyme RppH triggers messenger RNA degradation by 5′ pyrophosphate removal. Nature 451, 355–358 (2008)
Mackie, G. A. Ribonuclease E is a 5′-end-dependent endonuclease. Nature 395, 720–724 (1998)
Frick, D. N. & Bessman, M. J. Cloning, purification, and properties of a novel NADH pyrophosphatase. Evidence for a nucleotide pyrophosphatase catalytic domain in MutT-like enzymes. J. Biol. Chem. 270, 1529–1534 (1995)
Walseth, T. F. & Lee, H. C. Synthesis and characterization of antagonists of cyclic-ADP-ribose-induced Ca2+ release. Biochim. Biophys. Acta 1178, 235–242 (1993)
Preugschat, F., Tomberlin, G. H. & Porter, D. J. The base exchange reaction of NAD+ glycohydrolase: identification of novel heterocyclic alternative substrates. Arch. Biochem. Biophys. 479, 114–120 (2008)
Migaud, M. E., Pederick, R. L., Bailey, V. C. & Potter, B. V. Probing Aplysia californica adenosine 5′-diphosphate ribosyl cyclase for substrate binding requirements: design of potent inhibitors. Biochemistry 38, 9105–9114 (1999)
Rostovtsev, V. V., Green, L. G., Fokin, V. V. & Sharpless, K. B. A stepwise Huisgen cycloaddition process: copper(I)-catalyzed regioselective “ligation” of azides and terminal alkynes. Angew. Chem. Int. Edn Engl. 41, 2596–2599 (2002)
Tornøe, C. W., Christensen, C. & Meldal, M. Peptidotriazoles on solid phase: [1,2,3]-triazoles by regiospecific copper(I)-catalyzed 1,3-dipolar cycloadditions of terminal alkynes to azides. J. Org. Chem. 67, 3057–3064 (2002)
Huang, F. Efficient incorporation of CoA, NAD and FAD into RNA by in vitro transcription. Nucleic Acids Res. 31, e8 (2003)
Opdyke, J. A., Fozo, E. M., Hemm, M. R. & Storz, G. RNase III participates in GadY-dependent cleavage of the gadX-gadW mRNA. J. Mol. Biol. 406, 29–43 (2011)
Opdyke, J. A., Kang, J. G. & Storz, G. GadY, a small-RNA regulator of acid response genes in Escherichia coli. J. Bacteriol. 186, 6698–6705 (2004)
Gerhart E, H. & Nordström, K. Structural analysis of an RNA molecule involved in replication control of plasmid R1. Nucleic Acids Res. 14, 2523–2538 (1986)
Lacatena, R. M. & Cesareni, G. Base pairing of RNA I with its complementary sequence in the primer precursor inhibits ColE1 replication. Nature 294, 623–626 (1981)
D’Alessio, G. & Josse, J. Glyceraldehyde phosphate dehydrogenase, phosphoglycerate kinase, and phosphoglyceromutase of Escherichia coli. Simultaneous purification and physical properties. J. Biol. Chem. 246, 4319–4325 (1971)
Masuda, N. & Church, G. M. Regulatory network of acid resistance genes in Escherichia coli. Mol. Microbiol. 48, 699–712 (2003)
Jin, Y., Watt, R. M., Danchin, A. & Huang, J. D. Small noncoding RNA GcvB is a novel regulator of acid resistance in Escherichia coli. BMC Genomics 10, 165 (2009)
Hayes, E. T. et al. Oxygen limitation modulates pH regulation of catabolism and hydrogenases, multidrug transporters, and envelope composition in Escherichia coli K-12. BMC Microbiol. 6, 89 (2006)
Gustavsson, N., Diez, A. & Nyström, T. The universal stress protein paralogues of Escherichia coli are co-ordinately regulated and co-operate in the defence against DNA damage. Mol. Microbiol. 43, 107–117 (2002)
Eguchi, Y. & Tomizawa, J. Complex formed by complementary RNA stem-loops and its stabilization by a protein: function of CoIE1 Rom protein. Cell 60, 199–209 (1990)
Dumelin, C. E., Chen, Y., Leconte, A. M., Chen, Y. G. & Liu, D. R. Discovery and biological characterization of geranylated RNA in bacteria. Nature Chem. Biol. 8, 913–919 (2012)
Bandyra, K. J. et al. The seed region of a small RNA drives the controlled destruction of the target mRNA by the endoribonuclease RNase E. Mol. Cell 47, 943–953 (2012)
McLennan, A. G. Substrate ambiguity among the nudix hydrolases: biologically significant, evolutionary remnant, or both? Cell. Mol. Life Sci. 70, 373–385 (2013)
Song, M. G., Bail, S. & Kiledjian, M. Multiple Nudix family proteins possess mRNA decapping activity. RNA 19, 390–399 (2013)
Gherardini, P. F., Ausiello, G., Russell, R. B. & Helmer-Citterich, M. Modular architecture of nucleotide-binding pockets. Nucleic Acids Res. 38, 3809–3816 (2010)
Bandyra, K. J., Bouvier, M., Carpousis, A. J. & Luisi, B. F. The social fabric of the RNA degradosome. Biochim. Biophys. Acta 1829, 514–522 (2013)
Fjeld, C. C., Birdsong, W. T. & Goodman, R. H. Differential binding of NAD+ and NADH allows the transcriptional corepressor carboxyl-terminal binding protein to serve as a metabolic sensor. Proc. Natl Acad. Sci. USA 100, 9202–9207 (2003)
Baba, T. et al. Construction of Escherichia coli K-12 in-frame, single-gene knockout mutants: the Keio collection. Mol. Syst. Biol. 2, 2006.0008 (2006)
Hafner, M. et al. Identification of microRNAs and other small regulatory RNAs using cDNA library sequencing. Methods 44, 3–12 (2008)
Lohse, M. et al. RobiNA: a user-friendly, integrated software solution for RNA-Seq-based transcriptomics. Nucleic Acids Res. 40, W622–W627 (2012)
Halbritter, F., Vaidya, H. J. & Tomlinson, S. R. GeneProf: analysis of high-throughput sequencing experiments. Nature Methods 9, 7–8 (2012)
Flicek, P. et al. Ensembl 2013. Nucleic Acids Res. 41, D48–D55 (2013)
Langmead, B., Trapnell, C., Pop, M. & Salzberg, S. L. Ultrafast and memory-efficient alignment of short DNA sequences to the human genome. Genome Biol. 10, R25 (2009)
Nicol, J. W., Helt, G. A., Blanchard, S. G., Jr, Raja, A. & Loraine, A. E. The Integrated Genome Browser: free software for distribution and exploration of genome-scale datasets. Bioinformatics 25, 2730–2731 (2009)
Keseler, I. M. et al. EcoCyc: fusing model organism databases with systems biology. Nucleic Acids Res. 41, D605–D612 (2013)
Bernofsky, C. & Gallagher, W. J. Nonenzymic hydrogen transfer between reduced and oxidized pyridine nucleotides. Biochim. Biophys. Acta 659, 1–6 (1981)
Celesnik, H., Deana, A. & Belasco, J. G. PABLO analysis of RNA: 5′-phosphorylation state and 5′-end mapping. Methods Enzymol. 447, 83–98 (2008)
Celesnik, H., Deana, A. & Belasco, J. G. Initiation of RNA decay in Escherichia coli by 5′ pyrophosphate removal. Mol. Cell 27, 79–90 (2007)
Tomcsányi, T. & Apirion, D. Processing enzyme ribonuclease E specifically cleaves RNA I. An inhibitor of primer formation in plasmid DNA synthesis. J. Mol. Biol. 185, 713–720 (1985)
Masukata, H. & Tomizawa, J. Control of primer formation for ColE1 plasmid replication: conformational change of the primer transcript. Cell 44, 125–136 (1986)
Blomberg, P., Wagner, E. G. & Nordström, K. Control of replication of plasmid R1: the duplex between the antisense RNA, CopA, and its target, CopT, is processed specifically in vivo and in vitro by RNase III. EMBO J. 9, 2331–2340 (1990)
Sharma, C. M., Darfeuille, F., Plantinga, T. H. & Vogel, J. A small RNA regulates multiple ABC transporter mRNAs by targeting C/A-rich elements inside and upstream of ribosome-binding sites. Genes Dev. 21, 2804–2817 (2007)
Thomason, M. K., Fontaine, F., De Lay, N. & Storz, G. A small RNA that regulates motility and biofilm formation in response to changes in nutrient availability in Escherichia coli. Mol. Microbiol. 84, 17–35 (2012)
Jørgensen, M. G. et al. Small regulatory RNAs control the multi-cellular adhesive lifestyle of Escherichia coli. Mol. Microbiol. 84, 36–50 (2012)
Jørgensen, M. G., Thomason, M. K., Havelund, J., Valentin-Hansen, P. & Storz, G. Dual function of the McaS small RNA in controlling biofilm formation. Genes Dev. 27, 1132–1145 (2013)
Rasmussen, A. A. et al. A conserved small RNA promotes silencing of the outer membrane protein YbfM. Mol. Microbiol. 72, 566–577 (2009)
Mandin, P. & Gottesman, S. A genetic approach for finding small RNAs regulators of genes of interest identifies RybC as regulating the DpiA/DpiB two-component system. Mol. Microbiol. 72, 551–565 (2009)
Weilbacher, T. et al. A novel sRNA component of the carbon storage regulatory system of Escherichia coli. Mol. Microbiol. 48, 657–670 (2003)
Lease, R. A., Smith, D., McDonough, K. & Belfort, M. The small noncoding DsrA RNA is an acid resistance regulator in Escherichia coli. J. Bacteriol. 186, 6179–6185 (2004)
Sledjeski, D. D., Gupta, A. & Gottesman, S. The small RNA, DsrA, is essential for the low temperature expression of RpoS during exponential growth in Escherichia coli. EMBO J. 15, 3993–4000 (1996)
Lease, R. A. & Belfort, M. A trans-acting RNA as a control switch in Escherichia coli: DsrA modulates function by forming alternative structures. Proc. Natl Acad. Sci. USA 97, 9919–9924 (2000)
Lease, R. A., Cusick, M. E. & Belfort, M. Riboregulation in Escherichia coli: DsrA RNA acts by RNA:RNA interactions at multiple loci. Proc. Natl Acad. Sci. USA 95, 12456–12461 (1998)
Göpel, Y., Papenfort, K., Reichenbach, B., Vogel, J. & Görke, B. Targeted decay of a regulatory small RNA by an adaptor protein for RNase E and counteraction by an anti-adaptor RNA. Genes Dev. 27, 552–564 (2013)
Urban, J. H. & Vogel, J. Two seemingly homologous noncoding RNAs act hierarchically to activate glmS mRNA translation. PLoS Biol. 6, e64 (2008)
Reichenbach, B., Maes, A., Kalamorz, F., Hajnsdorf, E. & Görke, B. The small RNA GlmY acts upstream of the sRNA GlmZ in the activation of glmS expression and is subject to regulation by polyadenylation in Escherichia coli. Nucleic Acids Res. 36, 2570–2580 (2008)
Moon, K. & Gottesman, S. A PhoQ/P-regulated small RNA regulates sensitivity of Escherichia coli to antimicrobial peptides. Mol. Microbiol. 74, 1314–1330 (2009)
Fozo, E. M. et al. Repression of small toxic protein synthesis by the Sib and OhsC small RNAs. Mol. Microbiol. 70, 1076–1093 (2008)
Vogel, J. et al. RNomics in Escherichia coli detects new sRNA species and indicates parallel transcriptional output in bacteria. Nucleic Acids Res. 31, 6435–6443 (2003)
Blasi, F. et al. Inhibition of transcription of the histidine operon in vitro by the first enzyme of the histidine pathway. Proc. Natl Acad. Sci. USA 70, 2692–2696 (1973)
Richards, J., Luciano, D. J. & Belasco, J. G. Influence of translation on RppH-dependent mRNA degradation in Escherichia coli. Mol. Microbiol. 86, 1063–1072 (2012)
Nobelmann, B. & Lengeler, J. W. Molecular analysis of the gat genes from Escherichia coli and of their roles in galactitol transport and metabolism. J. Bacteriol. 178, 6790–6795 (1996)
Tucker, D. L., Tucker, N. & Conway, T. Gene expression profiling of the pH response in Escherichia coli. J. Bacteriol. 184, 6551–6558 (2002)
Peng, C. A., Oliver, M. J. & Wood, A. J. Is the Rehydrin TrDr3 from Tortula ruralis associated with tolerance to cold, salinity, and reduced pH? Physiological evaluation of the TrDr3-orthologue, HdeD from Escherichia coli in response to abiotic stress. Plant Biol. 7, 315–320 (2005)
Lawther, R. P. & Hatfield, G. W. Multivalent translational control of transcription termination at attenuator of ilvGEDA operon of Escherichia coli K-12. Proc. Natl Acad. Sci. USA 77, 1862–1866 (1980)
Purta, E. et al. The yfhQ gene of Escherichia coli encodes a tRNA:Cm32/Um32 methyltransferase. BMC Mol. Biol. 7, 23 (2006)
Elovson, J. & Vagelos, P. R. Acyl carrier protein. X. Acyl carrier protein synthetase. J. Biol. Chem. 243, 3603–3611 (1968)
Lu, P., Vogel, C., Wang, R., Yao, X. & Marcotte, E. M. Absolute protein expression profiling estimates the relative contributions of transcriptional and translational regulation. Nature Biotechnol. 25, 117–124 (2007)
Rudolph, F. B. & Fromm, H. J. The purification and properties of aspartase from Escherichia coli. Arch. Biochem. Biophys. 147, 92–98 (1971)
Karsten, W. E. & Viola, R. E. Kinetic studies of L-aspartase from Escherichia coli: pH-dependent activity changes. Arch. Biochem. Biophys. 287, 60–67 (1991)
Maki, Y., Yoshida, H. & Wada, A. Two proteins, YfiA and YhbH, associated with resting ribosomes in stationary phase Escherichia coli. Genes Cells 5, 965–974 (2000)
Agafonov, D. E., Kolb, V. A., Nazimov, I. V. & Spirin, A. S. A protein residing at the subunit interface of the bacterial ribosome. Proc. Natl Acad. Sci. USA 96, 12345–12349 (1999)
Vila-Sanjurjo, A., Schuwirth, B. S., Hau, C. W. & Cate, J. H. Structural basis for the control of translation initiation during stress. Nature Struct. Mol. Biol. 11, 1054–1059 (2004)
Agafonov, D. E., Kolb, V. A. & Spirin, A. S. Ribosome-associated protein that inhibits translation at the aminoacyl-tRNA binding stage. EMBO Rep. 2, 399–402 (2001)
Kim, K. S. & Lee, Y. Regulation of 6S RNA biogenesis by switching utilization of both sigma factors and endoribonucleases. Nucleic Acids Res. 32, 6057–6068 (2004)
Weber, M. M., French, C. L., Barnes, M. B., Siegele, D. A. & McLean, R. J. A previously uncharacterized gene, yjfO (bsmA), influences Escherichia coli biofilm formation and stress response. Microbiology 156, 139–147 (2010)
Robbins, J. C. & Oxender, D. L. Transport systems for alanine, serine, and glycine in Escherichia coli K-12. J. Bacteriol. 116, 12–18 (1973)
Ghrist, A. C. & Stauffer, G. V. The Escherichia coli glycine transport system and its role in the regulation of the glycine cleavage enzyme system. Microbiology 141, 133–140 (1995)
Pulvermacher, S. C., Stauffer, L. T. & Stauffer, G. V. Role of the sRNA GcvB in regulation of cycA in Escherichia coli. Microbiology 155, 106–114 (2009)
Hugovieux-Cotte-Pattat, N. & Robert-Baudouy, J. Regulation and transcription direction of exuR, a self-regulated repressor in Escherichia coli K-12. J. Mol. Biol. 156, 221–228 (1982)
Rodionov, D. A., Mironov, A. A., Rakhmaninova, A. B. & Gelfand, M. S. Transcriptional regulation of transport and utilization systems for hexuronides, hexuronates and hexonates in gamma purple bacteria. Mol. Microbiol. 38, 673–683 (2000)
Cole, S. T. et al. Nucleotide sequence and gene-polypeptide relationships of the glpABC operon encoding the anaerobic sn-glycerol-3-phosphate dehydrogenase of Escherichia coli K-12. J. Bacteriol. 170, 2448–2456 (1988)
Schryvers, A., Lohmeier, E. & Weiner, J. H. Chemical and functional properties of the native and reconstituted forms of the membrane-bound, aerobic glycerol-3-phosphate dehydrogenase of Escherichia coli. J. Biol. Chem. 253, 783–788 (1978)
Summers, M. C. & Rose, I. A. Proton transfer reactions of methylglyoxal synthase. J. Am. Chem. Soc. 99, 4475–4478 (1977)
Plumbridge, J. A. Repression and induction of the nag regulon of Escherichia coli K-12: the roles of nagC and nagA in maintenance of the uninduced state. Mol. Microbiol. 5, 2053–2062 (1991)
White, R. J. Control of amino sugar metabolism in Escherichia coli and isolation of mutants unable to degrade amino sugars. Biochem. J. 106, 847–858 (1968)
Freundlich, M., Ramani, N., Mathew, E., Sirko, A. & Tsui, P. The role of integration host factor in gene expression in Escherichia coli. Mol. Microbiol. 6, 2557–2563 (1992)
Nelson, W. C., Howard, M. T., Sherman, J. A. & Matson, S. W. The traY gene product and integration host factor stimulate Escherichia coli DNA helicase I-catalyzed nicking at the F plasmid oriT. J. Biol. Chem. 270, 28374–28380 (1995)
Dhavan, G. M., Crothers, D. M., Chance, M. R. & Brenowitz, M. Concerted binding and bending of DNA by Escherichia coli integration host factor. J. Mol. Biol. 315, 1027–1037 (2002)
Yanagisawa, T., Sumida, T., Ishii, R., Takemoto, C. & Yokoyama, S. A paralog of lysyl-tRNA synthetase aminoacylates a conserved lysine residue in translation elongation factor P. Nature Struct. Mol. Biol. 17, 1136–1143 (2010)
Kaiser, K. & Murray, N. E. Physical characterisation of the “Rac prophage” in E. coli K12. Mol. Gen. Genet. 175, 159–174 (1979)
Caldas, T. et al. The FtsJ/RrmJ heat shock protein of Escherichia coli is a 23 S ribosomal RNA methyltransferase. J. Biol. Chem. 275, 16414–16419 (2000)
Theobald, A., Springer, M., Grunberg-Manago, M., Ebel, J. P. & Giege, R. Tertiary structure of Escherichia coli tRNAThr3 in solution and interaction of this tRNA with the cognate threonyl-tRNA synthetase. Eur. J. Biochem. 175, 511–524 (1988)
Serres, M. H. et al. A functional update of the Escherichia coli K-12 genome. Genome Biol. 2, RESEARCH0035 (2001)
Tschowri, N., Busse, S. & Hengge, R. The BLUF-EAL protein YcgF acts as a direct anti-repressor in a blue-light response of Escherichia coli. Genes Dev. 23, 522–534 (2009)
Daley, D. O. et al. Global topology analysis of the Escherichia coli inner membrane proteome. Science 308, 1321–1323 (2005)
Lhoest, J. & Colson, C. Cold-sensitive ribosome assembly in an Escherichia coli mutant lacking a single methyl group in ribosomal protein L3. Eur. J. Biochem. 121, 33–37 (1981)
Heurgué-Hamard, V., Champ, S., Engström, A., Ehrenberg, M. & Buckingham, R. H. The hemK gene in Escherichia coli encodes the N5-glutamine methyltransferase that modifies peptide release factors. EMBO J. 21, 769–778 (2002)
White, P. J., Millar, G. & Coggins, J. R. The overexpression, purification and complete amino acid sequence of chorismate synthase from Escherichia coli K12 and its comparison with the enzyme from Neurospora crassa. Biochem. J. 251, 313–322 (1988)
Imamura, R. et al. Identification of the cpdA gene encoding cyclic 3′,5′-adenosine monophosphate phosphodiesterase in Escherichia coli. J. Biol. Chem. 271, 25423–25429 (1996)
Long, C. W. & Pardee, A. B. Cytidine triphosphate synthetase of Escherichia coli B. I. Purification and kinetics. J. Biol. Chem. 242, 4715–4721 (1967)
Py, B., Higgins, C. F., Krisch, H. M. & Carpousis, A. J. A DEAD-box RNA helicase in the Escherichia coli RNA degradosome. Nature 381, 169–172 (1996)
Jeanningros, R., Creuzet-Sigal, N., Frixon, C. & Cattaneo, J. Purification and properties of a debranching enzyme from Escherichia coli. Biochim. Biophys. Acta 438, 186–199 (1976)
Ballicora, M. A., Iglesias, A. A. & Preiss, J. ADP-glucose pyrophosphorylase, a regulatory enzyme for bacterial glycogen synthesis. Microbiol. Mol. Biol. Rev. 67, 213–225 (2003)
Joly, J. C. & Wickner, W. The SecA and SecY subunits of translocase are the nearest neighbors of a translocating preprotein, shielding it from phospholipids. EMBO J. 12, 255–263 (1993)
Wada, A. & Sako, T. Primary structures of and genes for new ribosomal proteins A and B in Escherichia coli. J. Biochem. 101, 817–820 (1987)
Sparrow, C. P. & Raetz, C. R. Purification and properties of the membrane-bound CDP-diglyceride synthetase from Escherichia coli. J. Biol. Chem. 260, 12084–12091 (1985)
Langley, K. E. & Kennedy, E. P. Partial purification and properties of CTP:phosphatidic acid cytidylyltransferase from membranes of Escherichia coli. J. Bacteriol. 136, 85–95 (1978)
Kanehara, K., Ito, K. & Akiyama, Y. YaeL (EcfE) activates the sigma(E) pathway of stress response through a site-2 cleavage of anti-sigma(E), RseA. Genes Dev. 16, 2147–2155 (2002)
Alba, B. M., Leeds, J. A., Onufryk, C., Lu, C. Z. & Gross, C. A. DegS and YaeL participate sequentially in the cleavage of RseA to activate the sigma(E)-dependent extracytoplasmic stress response. Genes Dev. 16, 2156–2168 (2002)
Takeda, K., Akimoto, C. & Kawamukai, M. Effects of the Escherichia coli sfsA gene on mal genes expression and a DNA binding activity of SfsA. Biosci. Biotechnol. Biochem. 65, 213–217 (2001)
Kawamukai, M. et al. Nucleotide sequence and characterization of the sfs1 gene: sfs1 is involved in CRP*-dependent mal gene expression in Escherichia coli. J. Bacteriol. 173, 2644–2648 (1991)
Paul, B. J. et al. DksA: a critical component of the transcription initiation machinery that potentiates the regulation of rRNA promoters by ppGpp and the initiating NTP. Cell 118, 311–322 (2004)
Clark, R. B. & Ogilvie, J. W. Aspartokinase I-homoserine dehydrogenase I of Escherichia coli K12. Subunit molecular weight and nicotinamide-adenine dinucleotide phosphate binding. Biochemistry 11, 1278–1282 (1972)
Starnes, W. L. et al. Threonine-sensitive aspartokinase-homoserine dehydrogenase complex, amino acid composition, molecular weight, and subunit composition of the complex. Biochemistry 11, 677–687 (1972)
Burr, B., Walker, J., Truffa-Bachi, P. & Cohen, G. N. Homoserine kinase from Escherichia coli K12. Eur. J. Biochem. 62, 519–526 (1976)
Johansen, J., Eriksen, M., Kallipolitis, B. & Valentin-Hansen, P. Down-regulation of outer membrane proteins by noncoding RNAs: unraveling the cAMP-CRP- and σE-dependent CyaR-ompX regulatory case. J. Mol. Biol. 383, 1–9 (2008)
De Lay, N. & Gottesman, S. The Crp-activated small noncoding regulatory RNA CyaR (RyeE) links nutritional status to group behavior. J. Bacteriol. 191, 461–476 (2009)
Andersen, J., Forst, S. A., Zhao, K., Inouye, M. & Delihas, N. The function of micF RNA. micF RNA is a major factor in the thermal regulation of OmpF protein in Escherichia coli. J. Biol. Chem. 264, 17961–17970 (1989)
Deighan, P., Free, A. & Dorman, C. J. A role for the Escherichia coli H-NS-like protein StpA in OmpF porin expression through modulation of micF RNA stability. Mol. Microbiol. 38, 126–139 (2000)
Trotochaud, A. E. & Wassarman, K. M. 6S RNA function enhances long-term cell survival. J. Bacteriol. 186, 4978–4985 (2004)
Wassarman, K. M. & Storz, G. 6S RNA regulates E. coli RNA polymerase activity. Cell 101, 613–623 (2000)
Kim, E. Y., Shin, M. S., Rhee, J. H. & Choy, H. E. Factors influencing preferential utilization of RNA polymerase containing sigma-38 in stationary-phase gene expression in Escherichia coli. J. Microbiol. 42, 103–110 (2004)
Muto, A. et al. Structure and function of 10Sa RNA: trans-translation system. Biochimie 78, 985–991 (1996)
Roche, E. D. & Sauer, R. T. SsrA-mediated peptide tagging caused by rare codons and tRNA scarcity. EMBO J. 18, 4579–4589 (1999)
Kawano, M., Reynolds, A. A., Miranda-Rios, J. & Storz, G. Detection of 5′- and 3′-UTR-derived small RNAs and cis-encoded antisense RNAs in Escherichia coli. Nucleic Acids Res. 33, 1040–1050 (2005)
Acknowledgements
We thank the CellNetworks Deep Sequencing Core Facility for Solexa sequencing, M. Brunner for access to the light cycler, M. Helm for advice on RNA mass spectrometry and B. Luisi for the RNase E expression vector, as well as A. Krause, J. Becker, A. Samanta, M. Tesch, F. Siebert, L. Obenauer and other members of the Jäschke laboratory for help and discussions. H.C. was supported by a postdoctoral fellowship from the Alexander-von-Humboldt Foundation. M.-L.W. acknowledges a PhD fellowship from the Hartmut Hoffmann-Berling International Graduate School of Molecular & Cellular Biology. A.J. is supported by the Deutsche Forschungsgemeinschaft, SFB 623, the Federal Ministry of Education and Research (BMBF), and the Helmholtz Initiative on Synthetic Biology.
Author information
Authors and Affiliations
Contributions
All authors designed the experiments, analysed and interpreted results and wrote the paper. H.C., M.-L.W., K.H. and G.N. performed the experiments.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Extended data figures and tables
Extended Data Figure 1 Development of the NAD capture protocol.
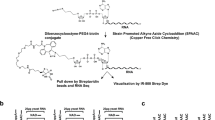
a, General scheme: ADPRC-catalysed transglycosylation with different alcohols, followed by ‘click’ biotinylation with biotin-PEG3-azide. b, Isolated yields and HPLC retention times. c, HPLC chromatogram of the ADPRC-catalysed reaction of NAD with 4-pentyn-1-ol. d, High-resolution ESI mass spectrum of the transglycosylation product. e, HPLC chromatogram of the copper-catalysed azide-alkyne cycloaddition (CuAAC) reaction mixture with the product from c. f, High-resolution ESI mass spectrum of the CuAAC reaction product.
Extended Data Figure 2 NAD capture protocol applied to RNA and comparison of the enrichment in NGS data sets from three different NAD captureSeq experiments.
a–c, Agarose gels of the PCR amplification products of cDNA obtained from the NAD capture protocol, applied to in vitro transcribed unmodified RNA and in vitro transcribed model NAD-RNA after 5 (left), 10 (middle), and 15 (right) cycles (a); and total RNA from E. coli JM109 (b) or K-12 (c). The region (120–160 bp) around the band present in the JM109 sample (∼150 bp, arrow) that occurred only in the fully treated sample but not in the controls was chosen for NGS libraries of minus ADPRC control A1 and fully treated sample B1, in agreement with the expected size (<200 nucleotides) of NAD-modified RNA according to previous observations2. The intense band of ∼150 bp size was later attributed to the most abundant enriched RNA, RNAI, which was absent in the K-12 sample. For this reason, the K-12 sample did not contain such a prominent additional band. For comparability, the same region (120–160 bp) was excised for library preparation for K-12 control A2 and sample B2. To achieve a more comprehensive picture and study the influence of sample preparation on data, a third library was prepared from JM109 (control A3, sample B3) choosing a broader size range (50–300 bp). d–i, Abundance of RNAs in fully treated sample (B) versus minus ADPRC control (A) in data set 1 (JM109, 120–160 bp; d, e), data set 2 (K-12, 120–160 bp; f, g), and data set 3 (JM109, 50–300 bp; h, i) found by strand-specific NGS analysis (RPM, average of forward and reverse read counts; Supplementary Data 1–3). Triangles, enriched; circles, non-enriched.
Extended Data Figure 3 NGS analysis of specific sequences found in control A1 and sample B1 in forward and reverse reads.
Analysis of sRNAs, sRNA-like 5′-terminal fragments of mRNAs, and non-enriched tRNASer. Normalized nucleotide counts derived from NGS reads (JM109, 120–160 bp) of control A1 (red) and fully treated sample B1 (green) are plotted for the regions on the E. coli K-12 genome or the respective plasmid in which the sequences are situated. A black bar indicates the protein-coding sequence of mRNAs, while non-coding RNAs are indicated by a thin black line. Plus and minus signs designate the orientation of genes in the genome/plasmid.
Extended Data Figure 4 NGS, real-time PCR, and northern blot analysis of RNAs enriched in NAD captureSeq.
a, Enrichment of selected sequences calculated from NGS read numbers or real-time PCR data. Numbers in parentheses show enrichment of genes below abundance threshold T in the respective data set. N.D., not determined. b, Real-time PCR analysis of selected mRNAs for 5′-termini (regions found to be enriched) and regions in the middle of the mRNA. Data show that the 5′-termini are highly enriched, while middle regions are not, or are considerably less enriched. All real-time PCR experiments were performed in duplicate for the same cDNA, and no-template controls were used to assess primer dimerization. c, Northern blot analysis. The expected size range (∼100 to 340 nucleotides for gatY, ∼100 to 240 nucleotides for hdeD) is indicated and for hdeD the fragment of expected size is indicated by an arrow, d, Example of an enriched 5′-terminus, gatY. The 5′-untranslated region is indicated in grey, the coding sequence in black. Minus sign indicates that the gene is encoded on the minus-strand of the genome, thus in reverse orientation. The position of the promoter (gatYp) is indicated by an arrow. Regions amplified in real-time PCR are indicated by opposing arrows.
Extended Data Figure 5 Mass-spectrometric analysis of the ADPRC-catalysed transglycosylation reaction of NAD-RNAI from E. coli total RNA and 4-pentyn-1-ol, and analysis of the 5′-modification status of GcvB.
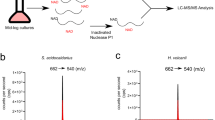
a, b, Disappearance of the ADP-ribose peak ([M–H]− m/z = 540.05, elution at 5–6 min) (a), and appearance of a peak at m/z = 624.10 (elution time 2.0–2.5 min) upon ADPRC reaction (b). Samples were digested with nuclease P1 before mass spectrometry. c, Simulated signal pattern of the expected reaction product (isotopic distribution). d–f, Observed mass spectrum in the chromatography fractions eluting at 2–2.5 min of untreated RNAI (d), RNAI treated with ADPRC and 4-pentyn-1-ol and then digested by nuclease P1 (e) and 5S rRNA treated with ADPRC and 4-pentyn-1-ol and then digested by nuclease P1 (f). g, HPLC-MS extracted ion chromatograms of the peak [M]+ m/z = 664.116 of free NAD, and nuclease-P1-digested 5S rRNA and GcvB RNA. h, High-resolution mass spectra of the HPLC fractions eluting at 2.2–2.4 min.
Extended Data Figure 6 NudC processing, 5′-dependent sensitization to RNase E processing by RppH or NudC, and investigation of RNA stability in vivo depending on NudC expression.
a, b, Decapping activity on in vitro transcribed NAD-RNAI of wild-type NudC enzyme and the E178Q point mutant. Gel-electrophoretic assay, monitoring the disappearance of a site-specifically installed radiolabel (see Fig. 3c). Mean of a technical duplicate ± s.d. is shown. c, d, Successive decapping and digestion of NAD-RNAI and triphosphate-RNAI. Time course of RppH/RNase E or NudC/RNase E decapping/digests of triphosphate-RNAI (c) or NAD-RNAI (d). e–h, Determination of RNAI (mean of biological duplicate ± s.d.) (e, f) and GcvB (g, h) cellular half-life depending on NudC expression by rifampicin stop assay. Detection by northern blotting, tRNASer as loading control.
Extended Data Figure 7 Determination of the 5′-modification status of cellular RNAs by a modified PABLO assay5,40.
a, Schematic overview of NAD quantification by dephosphorylation, NudC cleavage and splinted ligation. Total RNA is isolated from different E. coli strains: wild type + RNAI (1); ΔnudC + RNAI (2); ΔnudC + NudC + RNAI (3); or ΔnudC + NudC(E178Q) + RNAI (4) (for strain details, see i). To remove all 5′-monophosphate and 5′-triphosphate termini, total RNA samples are subjected to alkaline phosphatase treatment. Then, to remove 5′-NAD and produce 5′-monophosphate termini, the dephosphorylated samples are treated with NudC (or NudC(E178Q) as a negative control). Finally, the resulting 5′-monophosphate RNA is ligated specifically to an adaptor by splinted ligation. Ligated fractions are determined by northern blotting. b, c, Validation of time-dependent in vitro NudC processing of cellular sRNAs (RNAI, 6S RNA) in total RNA from E. coli wild type + RNAI (1), using the assay described in a. Aliquots were taken after different times of NudC treatment and subjected to ligation as shown. tRNASer is blotted as loading control. d, e, Quantification of the efficiency of NudC treatment and splinted ligation (processing efficiency) for NAD-full-length RNAI (NAD-FL-RNAI) and NAD-6S RNA. To calculate the percentage of NAD modification (shown in Fig. 4), it is necessary to normalize to the cumulative processing efficiency of NudC treatment and splinted ligation of NAD-RNA, which is determined experimentally for each total RNA preparation. To this end, in vitro transcribed radiolabelled NAD32P-FL-RNAI (d) or NAD32P-6S RNA (e) is spiked into total RNAs isolated from the four E. coli strains mentioned above. The mixture is then subjected to the treatment described in a. Processing efficiencies of radiolabelled RNAs are quantified ratiometrically after gel electrophoresis and electroblotting. f, g, Quantification of the percentage of RNAI−5. Cellular RNAI consists of a mixture of full-length and processed species, among them RNAI−5 (ref. 42), which arises from RNase E cleavage, thus contains a 5′-monophosphate, and lacks only five 5′-terminal nucleotides. Both species exhibit very similar electrophoretic mobility. Here, we were interested only in the percentage of NAD-modified FL-RNAI. Thus, we quantified the percentage of RNAI−5 by PABLO analysis (splinted ligation of non-treated total RNA with RNAI−5-specific adaptor). Carrying out the respective control (in vitro transcribed monophosphate32P-RNAI−5, spiked into total RNA samples) (f), we determined the percentage of ligation product, and normalized to ligation efficiencies to obtain the percentage of RNAI−5 (g). With the help of the values determined in d–g, we then determined the percentage of FL-RNAI and the percentage of NAD-FL-RNAI (shown in Fig. 4a), using the formulas shown in h. Values shown in this figure represent replicate 1. (Expected) ligation products are marked with arrows (b–g). i, Strains used in this study.
Supplementary information
Supplementary Information
This file contains Supplementary Table 1. (PDF 178 kb)
Supplementary Data 1
This file contains strand-specific and non-strand-specific sequence count analysis for data set 1. (XLSX 1937 kb)
Supplementary Data 2
This file contains strand-specific and non-strand-specific sequence count analysis for data set 2. (XLSX 3915 kb)
Supplementary Data 3
This file contains strand-specific and non-strand-specific sequence count analysis for data set 3. (XLSX 1782 kb)
Rights and permissions
About this article
Cite this article
Cahová, H., Winz, ML., Höfer, K. et al. NAD captureSeq indicates NAD as a bacterial cap for a subset of regulatory RNAs. Nature 519, 374–377 (2015). https://doi.org/10.1038/nature14020
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature14020
This article is cited by
-
Recent advances in the plant epitranscriptome
Genome Biology (2023)
-
NADcapPro and circNC: methods for accurate profiling of NAD and non-canonical RNA caps in eukaryotes
Communications Biology (2023)
-
Arabidopsis DXO1 activates RNMT1 to methylate the mRNA guanosine cap
Nature Communications (2023)
-
Hepatitis C virus RNA is 5′-capped with flavin adenine dinucleotide
Nature (2023)
-
Xrn1 is a deNADding enzyme modulating mitochondrial NAD-capped RNA
Nature Communications (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.