Abstract
Polycystic ovary syndrome (PCOS), an endocrine disorder afflicting 6–20% of women of reproductive age globally, has been linked to alterations in the gut microbiome. We previously showed that in PCOS, elevation of Bacteroides vulgatus in the gut microbiome was associated with altered bile acid metabolism. Here we show that B. vulgatus also induces a PCOS-like phenotype in female mice via an alternate mechanism independent of bile acids. We find that B. vulgatus contributes to PCOS-like symptoms through its metabolite agmatine, which is derived from arginine by arginine decarboxylase. Mechanistically, agmatine activates the farnesoid X receptor (FXR) pathway to subsequently inhibit glucagon-like peptide-1 (GLP-1) secretion by L cells, which leads to insulin resistance and ovarian dysfunction. Critically, the GLP-1 receptor agonist liraglutide and the arginine decarboxylase inhibitor difluoromethylarginine ameliorate ovarian dysfunction in a PCOS-like mouse model. These findings reveal that agmatine–FXR–GLP-1 signalling contributes to ovarian dysfunction, presenting a potential therapeutic target for PCOS management.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Data availability
All the data relating to the metagenomic sequencing and RNA sequencing of this study have been uploaded to the Sequence Read Archive database and are available for download via accession numbers PRJNA1080049 and PRJNA1080099. Source data are provided with this paper.
Code availability
No custom code was used for this study.
References
Norman, R. J., Dewailly, D., Legro, R. S. & Hickey, T. E. Polycystic ovary syndrome. Lancet 370, 685–697 (2007).
Escobar-Morreale, H. F. Polycystic ovary syndrome: definition, aetiology, diagnosis and treatment. Nat. Rev. Endocrinol. 14, 270–284 (2018).
Azziz, R. et al. Polycystic ovary syndrome. Nat. Rev. Dis. Prim. 2, 16057 (2016).
Hoeger, K. M., Dokras, A. & Piltonen, T. Update on PCOS: consequences, challenges and guiding treatment. J. Clin. Endocrinol. Metab. 106, e1071–e1083 (2020).
Martel, J. et al. Gut barrier disruption and chronic disease. Trends Endocrinol. Metab. 33, 247–265 (2022).
Postler, T. S. & Ghosh, S. understanding the holobiont: how microbial metabolites affect human health and shape the immune system. Cell Metab. 26, 110–130 (2017).
Han, Q., Wang, J., Li, W., Chen, Z.-J. & Du, Y. Androgen-induced gut dysbiosis disrupts glucolipid metabolism and endocrinal functions in polycystic ovary syndrome. Microbiome 9, 101 (2021).
Sun, Y., Gao, S., Ye, C. & Zhao, W. Gut microbiota dysbiosis in polycystic ovary syndrome: mechanisms of progression and clinical applications. Front. Cell. Infect. Microbiol. 13, 1142041 (2023).
Lindheim, L. et al. Alterations in gut microbiome composition and barrier function are associated with reproductive and metabolic defects in women with polycystic ovary syndrome (PCOS): a pilot study. PLoS ONE 12, e0168390 (2017).
Suturina, L. et al. Polycystic ovary syndrome and gut microbiota: phenotype matters. Life 13, 7 (2023).
Chu, W. et al. Metagenomic analysis identified microbiome alterations and pathological association between intestinal microbiota and polycystic ovary syndrome. Fertil. Steril. 113, 1286 (2020).
Zeng, B. et al. Structural and functional profiles of the gut microbial community in polycystic ovary syndrome with insulin resistance (IR-PCOS): a pilot study. Res. Microbiol. 170, 43–52 (2019).
Liu, R. et al. Dysbiosis of gut microbiota associated with clinical parameters in polycystic ovary syndrome. Front. Microbiol. 8, 324 (2017).
Qi, X. et al. Gut microbiota-bile acid-interleukin-22 axis orchestrates polycystic ovary syndrome. Nat. Med. 25, 1225–1233 (2019).
Zheng, L. et al. CRISPR/Cas-based genome editing for human gut commensal bacteroides species. ACS Synth. Biol. 11, 464–472 (2022).
Jensterle, M. et al. The role of glucagon-like peptide-1 in reproduction: from physiology to therapeutic perspective. Hum. Reprod. Update 25, 504–517 (2019).
Gribble, F. M. & Reimann, F. Enteroendocrine cells: chemosensors in the intestinal epithelium. Annu. Rev. Physiol. 78, 277–299 (2016).
Nauck, M. A. & Meier, J. J. Incretin hormones: their role in health and disease. Diabetes Obes. Metab. 20, 5–21 (2018).
Tan, Q. et al. Recent advances in incretin-based pharmacotherapies for the treatment of obesity and diabetes. Front. Endocrinol. 13, 838410 (2022).
Coll, A. P. et al. GDF15 mediates the effects of metformin on body weight and energy balance. Nature 578, 444 (2020).
McLean, B. A. et al. Revisiting the complexity of GLP-1 action from sites of synthesis to receptor activation. Endocr. Rev. 42, 101–132 (2021).
Sandoval, D. A. & D’Alessio, D. A. Physiology of proglucagon peptides: role of glucagon and glp-1 in health and disease. Physiol. Rev. 95, 513–548 (2015).
Trabelsi, M.-S. et al. Farnesoid X receptor inhibits glucagon-like peptide-1 production by enteroendocrine L cells. Nat. Commun. 6, 7629 (2015).
Han, S. et al. A metabolomics pipeline for the mechanistic interrogation of the gut microbiome. Nature 595, 415 (2021).
Sun, X., Song, W. & Liu, L. Enzymatic production of agmatine by recombinant arginine decarboxylase. J. Mol. Catal. B-Enzym. 121, 1–8 (2015).
Burrell, M., Hanfrey, C. C., Murray, E. J., Stanley-Wall, N. R. & Michael, A. J. Evolution and multiplicity of arginine decarboxylases in polyamine biosynthesis and essential role in Bacillus subtilis biofilm formation. J. Biol. Chem. 285, 39224–39238 (2010).
Bitonti, A. J., Casara, P. J., McCann, P. P. & Bey, P. Catalytic irreversible inhibition of bacterial and plant arginine decarboxylase activities by novel substrate and product analogs. Biochem. J. 242, 69–74 (1987).
Pathak, P. et al. Intestine farnesoid X receptor agonist and the gut microbiota activate G-protein bile acid receptor-1 signaling to improve metabolism. Hepatology 68, 1574–1588 (2018).
Yoon, H. S. et al. Akkermansia muciniphila secretes a glucagon-like peptide-1-inducing protein that improves glucose homeostasis and ameliorates metabolic disease in mice. Nat. Microbiol. 6, 563–573 (2021).
Rakipovski, G. et al. The GLP-1 analogs liraglutide and semaglutide reduce atherosclerosis in ApoE(−/−) and LDLr(−/−) mice by a mechanism that includes inflammatory pathways. JACC Basic Transl. Sci. 3, 844–857 (2018).
McCartney, C. R. & Marshall, J. C. Polycystic ovary syndrome. N. Engl. J. Med. 375, 54–64 (2016).
Wahlstrom, A., Sayin, S. I., Marschall, H.-U. & Backhed, F. Intestinal crosstalk between bile acids and microbiota and its impact on host metabolism. Cell Metab. 24, 41–50 (2016).
Teodoro, J. S., Rolo, A. P. & Palmeira, C. M. Hepatic FXR: key regulator of whole-body energy metabolism. Trends Endocrinol. Metab. 22, 458–466 (2011).
Lefebvre, P., Cariou, B., Lien, F., Kuipers, F. & Staels, B. Role of bile acids and bile acid receptors in metabolic regulation. Physiol. Rev. 89, 147–191 (2009).
Jiang, C. et al. Intestine-selective farnesoid X receptor inhibition improves obesity-related metabolic dysfunction. Nat. Commun. 6, 10166 (2015).
Jiang, C. et al. Intestinal farnesoid X receptor signaling promotes nonalcoholic fatty liver disease. J. Clin. Invest. 125, 386–402 (2015).
Fiorucci, S., Distrutti, E., Carino, A., Zampella, A. & Biagioli, M. Bile acids and their receptors in metabolic disorders. Prog. Lipid Res. 82, 101094 (2021).
Liu, H., Hu, C., Zhang, X. & Jia, W. Role of gut microbiota, bile acids and their cross-talk in the effects of bariatric surgery on obesity and type 2 diabetes. J. Diabetes Investig. 9, 13–20 (2018).
Pyke, C. et al. GLP-1 receptor localization in monkey and human tissue: novel distribution revealed with extensively validated monoclonal antibody. Endocrinology 155, 1280–1290 (2014).
Drucker, D. J. The cardiovascular biology of glucagon-like peptide-1. Cell Metab. 24, 15–30 (2016).
Glintborg, D., Mumm, H., Holst, J. J. & Andersen, M. Effect of oral contraceptives and/or metformin on GLP-1 secretion and reactive hypoglycaemia in polycystic ovary syndrome. Endocr. Connect. 6, 267–277 (2017).
Vrbikova, J. et al. Incretin levels in polycystic ovary syndrome. Eur. J. Endocrinol. 159, 121–127 (2008).
Gama, R. et al. The entero-insular axis in polycystic ovarian syndrome. Ann. Clin. Biochem. 33, 190–195 (1996).
Pontikis, C. et al. The incretin effect and secretion in obese and lean women with polycystic ovary syndrome: a pilot study. J. Women’s Health 20, 971–976 (2011).
Cassar, S. et al. Biomarkers and insulin sensitivity in women with polycystic ovary syndrome: characteristics and predictive capacity. Clin. Endocrinol. 83, 50–58 (2015).
Abdalla, M. A., Deshmukh, H., Atkin, S. & Sathyapalan, T. The potential role of incretin-based therapies for polycystic ovary syndrome: a narrative review of the current evidence. Ther. Adv. Endocrinol. Metab. 12, 2042018821989238 (2021).
Baranowska-Bik, A. Therapy of obesity in women with PCOS using GLP-1-benefits and limitations. Endokrynol. Pol. 73, 627–643 (2022).
Helvaci, N. & Yildiz, B. O. Current and emerging drug treatment strategies for polycystic ovary syndrome. Expert Opin. Pharmacother. 24, 105–120 (2023).
Morris, S. M. Jr. Arginine metabolism revisited. J. Nutr. 146, 2579–2586 (2016).
Yu, J., Lee, J., Lee, S.-H., Cho, J.-H. & Kim, H.-S. A study on weight loss cause as per the side effect of liraglutide. Cardiovasc. Ther. 2022, 5201684 (2022).
Baekdal, T. A. et al. Effect of various dosing conditions on the pharmacokinetics of oral semaglutide, a human glucagon-like peptide-1 analogue in a tablet formulation. Diabetes Ther. 12, 1915–1927 (2021).
Saxena, A. R. et al. Efficacy and safety of oral small molecule glucagon-like peptide 1 receptor agonist danuglipron for glycemic control among patients with type 2 diabetes: a randomized clinical trial. JAMA Netw. Open 6, e2314493 (2023).
Zheng, X. et al. Hyocholic acid species improve glucose homeostasis through a distinct TGR5 and FXR signaling mechanism. Cell Metab. 33, 791–803.e7 (2020).
Liu, C. et al. Gut commensal Christensenella minuta modulates host metabolism via acylated secondary bile acids. Nat. Microbiol 9, 434–450 (2024).
Li, T. et al. A gut microbiota-bile acid axis promotes intestinal homeostasis upon aspirin-mediated damage. Cell Host Microbe 32, 191–208.e9 (2024).
Yildiz, M. & Ofori-Mensah, S. The effects of different commercial feeds and seasonal variation on fillet amino acid profile of sea bream (Sparus aurata) and sea bass (Dicentrarchus labrax). Turkish J. Fish. Aquat. Sci. 17, 1297–1307 (2017).
Gorska-Warsewicz, H. et al. Food products as sources of protein and amino acidsthe case of poland. Nutrients 10, 1977 (2018).
Guan, F. et al. Amino acids and lipids associated with long-term and short-term red meat consumption in the chinese population: an untargeted metabolomics study. Nutrients 13, 4567 (2021).
Rossocha, M., Schultz-Heienbrok, R., von Moeller, H., Coleman, J. P. & Saenger, W. Conjugated bile acid hydrolase is a tetrameric N-terminal thiol hydrolase with specific recognition of its cholyl but not of its tauryl product. Biochemistry 44, 5739–5748 (2005).
Yao, L. et al. A selective gut bacterial bile salt hydrolase alters host metabolism. eLife 7, e37182 (2018).
Arifuzzaman, M. et al. Inulin fibre promotes microbiota-derived bile acids and type 2 inflammation. Nature 611, 578–584 (2022).
Caldwell, A. S. L. et al. Neuroendocrine androgen action is a key extraovarian mediator in the development of polycystic ovary syndrome. Proc. Natl Acad. Sci. USA 114, E3334–E3343 (2017).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–U354 (2012).
Mortazavi, A., Williams, B. A., McCue, K., Schaeffer, L. & Wold, B. Mapping and quantifying mammalian transcriptomes by RNA-Seq. Nat. Methods 5, 621–628 (2008).
Robinson, M. D., McCarthy, D. J. & Smyth, G. K. edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26, 139–140 (2010).
Acknowledgements
This work was supported by the National Natural Science Foundation of the Peoples’ Republic of China (grant nos. 31925021, 82130022, 92357305 and 82341226 to C.J.), the National Key Research and Development Program of China (grant nos. 2022YFA0806400 to C.J. and 2022YFC2702500 to R.L.), the National Natural Science Foundation of the Peoples’ Republic of China (grant nos. 82022028 and 82171627 to Y.P., 82288102 to J.Q., 82001506 to X.Q., 92149306 and 81921001 to C.J., 82301841 to S.Y.). C.J. acknowledges the support from the Tencent Foundation through the Xplorer Prize. We thank L. Dai (Shenzhen Institute of Synthetic Biology, Institute of Advanced Technology) for providing the Bacteroides tool plasmid pB041; D. L. Gumucio (University of Michigan) for providing the villin Cre mice; G. L. Guo (Rutgers University) for providing the PGL4-Shp-TK firefly luciferase construct and human FXR expression plasmid; P. A. Dawson (Emory University) for providing the human ASBT expression plasmid and D. Drucker for providing the GLUTag cell line.
Author information
Authors and Affiliations
Contributions
Y.P., C.J., J.Q. and R.L. designed the study. C. Yun and X.Q. enroled the patients and collected the patient samples. C. Yun, S.Y., B.L. and M.Z. performed the animal experiments and analysed the data. Y.D. and K.W. guided the microbial genetic operations. S.Y. and Q.N. performed the LC–MS analysis. Y.Z., C. Ye, P.X. and M.M. analysed the metagenomic and RNA sequencing data. S.Y., C. Yun, C.J., J.Q. and Y.P. wrote the paper with input from all authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Metabolism thanks John Chiang, Wendong Huang and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Primary Handling Editor: Yanina-Yasmin Pesch, in collaboration with the Nature Metabolism team.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Bv-Δbsh colonization leads to a PCOS-like phenotype in mice without affecting the bile acid profile.
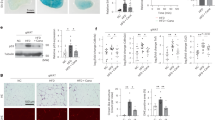
a, DNA gel electrophoresis experiment to verify the knockout of Bv-Δ1032, Bv-Δ2699, and Bv-Δ3993 in B. vulgatus. b, Growth curve of wild type B. vulgatus (Bv) and Bv-Δ2699 (Bv-Δbsh) in BHI medium. c-l, SPF mice were gavaged with wild type B. vulgatus (Bv), BVU2699-knockout B. vulgatus (Bv-Δbsh), or PBS (Vehicle) for three weeks. c, Colonization of Bv and Bv-Δbsh in mice. d, The total bile acid profile of serum. e, The content of UDCA in feces. f, The content of UDCA in serum. g, The levels of IL-22 in serum. h, Quantitative analysis of cystic follicles in the ovaries. i, Area under the curve (AUC) of GTT. j, The levels of fasting glucose in serum. k, The levels of insulin in serum. l, HOMA-IR. All data are presented as the mean ± s.e.m. (b-g, i-l) n = 6/group; h, n = 5/group. In c, the P values were determined by Kruskal–Wallis test followed by Dunn’s post hoc test. The P values were determined by one-way ANOVA with Tukey’s multiple comparison post hoc test in e, i, j, and one-way ANOVA with LSD test in f-h, k, l.
Extended Data Fig. 2 Cholestyramine cannot reverse B. vulgatus-induced PCOS-like phenotype.
a and b, Cholestyramine reduces the content of bile acids in mouse intestinal epithelium (a) and serum (b). (a) CA, P = 2.32 × 10−7; α-MCA, P = 0.000008; TCA, P = 2.85 × 10−7. (b) TUDCA, P = 0.000016. c-j, Cholestyramine treatment mice were gavaged with wild type B. vulgatus (Bv) or PBS (Vehicle) for three weeks. c, Quantitative analysis of cystic follicles in the ovaries. d, Levels of testosterone in serum. e, Levels of LH in serum. f, Glucose tolerance test (GTT) (15min: ***P = 0.0007; 30min: *P = 0.0249; *P = 0.0183; *compared with the vehicle + Cholestyramine). g, Area under the curve (AUC) of GTT. h, Insulin tolerance test (ITT) (15min: *P = 0.0152; 30min: **P = 0.0087; *compared with the vehicle + Cholestyramine). i, The levels of fasting glucose in serum. j, The level of insulin in serum. All data are presented as the mean ± s.e.m. (a, b, c) n = 5/group. (d-j) n = 6/group. In a, b, d-g, i, j, the P values were determined by two-tailed Student’s t-test. In c, h, the P values were determined by two-tailed Mann–Whitney U-test.
Extended Data Fig. 3 GLP-1R agonist liraglutide improves B. vulgatus-induced PCOS-like phenotype.
a, The levels of GIP in patients with PCOS and healthy control serum. b, The levels of GDF15 in patients with PCOS and healthy control serum. c, The correlation between the total GLP-1 level in the serum and LH/FSH ratio. d, The correlation between the total GLP-1 level in the serum and HOMA-IR. e, Quantitative analysis of cystic follicles in the ovaries. f, Area under the curve (AUC) of GTT. g, The levels of fasting glucose in serum. h, The level of insulin in serum. All data are presented as the mean ± s.e.m. (a-d) n = 20 in the control group and n = 20 in the PCOS group. (e) n = 5/group. (f-h) n = 6/group. In a, the P values were determined by two-tailed Mann–Whitney U-test. In b, the P values were determined by two-tailed Student’s t-test. In c, d, correlation analysis was determined by one tailed Pearson r-test. In e, the P values were determined by Kruskal–Wallis test followed by Dunn’s post hoc test. In f-h, the P values were determined by one-way ANOVA with Tukey’s multiple comparison post hoc test.
Extended Data Fig. 4 B. vulgatus inhibits GLP-1 secretion and induces PCOS-like phenotype depended on intestinal epithelial FXR.
a, The expression levels of Fgf15 and Shp in intestinal epithelium. b, The levels of C4 in serum. c, FXR expression in siFXR STC-1 cells. d, The levels of total GLP-1 in the cellular supernatant. e, The relative expression of Gcg. f, The levels of total GLP-1 in the cellular supernatant. g, Luciferase activity. h, The expression levels of Tgr5 in GLUTag cells. i, The levels of total GLP-1 in the cellular supernatant. j, Relative fold change of total GLP-1 levels to vehicle. k, The expression levels of Fxr and downstream genes in the intestinal epithelium of Fxrfl/fl and FxrΔIE mice. l-q, FxrΔIE or Fxrfl/fl mice were gavaged with Bv or PBS for three weeks. l, Quantitative analysis of corpora lutea. m, Quantitative analysis of cystic follicles. n, AUC of GTT. o, The levels of fasting glucose in serum. p, The level of insulin in serum. q, HOMA-IR. All data are presented as the mean ± s.e.m. (a, k-q) n = 5/group. (b) n = 20 in both control and PCOS group. (c, h) n = 3/group. (d-g, i, j) n = 4/group. In a, the P values were determined by two-tailed Mann–Whitney U-test. In b, d, e, k, the P values were determined by two-tailed Student’s t-test. The P values were determined by one-way ANOVA with Tukey’s multiple comparison post hoc test in c, f, h, m, o-q; one-way ANOVA with Dunnett’s T3 test in g and one-way ANOVA with LSD test in n. In i, j, l, the P values were determined by Kruskal–Wallis test followed by Dunn’s post hoc test.
Extended Data Fig. 5 GLP-1 receptor inhibitor reversed the improvement of B. vulgatus-induced PCOS-like phenotype in intestinal Fxr knockout mice.
a, Quantitative analysis of corpora lutea in the ovaries. b, Quantitative analysis of cystic follicles in the ovaries. c, Area under the curve (AUC) of GTT. d, The levels of fasting glucose in serum. e, The level of insulin in serum. f, HOMA-IR. All data are presented as the mean ± s.e.m. (a-f) n = 5/group. In a, b, d, the P values were determined by Kruskal–Wallis test followed by Dunn’s post hoc test. In c, e, f, the P values were determined by one-way ANOVA with LSD test.
Extended Data Fig. 6 Agmatine induces PCOS-like phenotype via a FXR signaling pathway.
a, The content of cytidine 5′-diphosphocholine in the feces of B. vulgatus model mice. b-h, FxrΔIE or Fxrfl/fl mice were gavaged with agmatine or PBS (Vehicle) for three weeks. b, Quantitative analysis of cystic follicles in the ovaries. c, Levels of testosterone in serum. d, Levels of LH in serum. e, Area under the curve (AUC) of GTT. f, The levels of fasting glucose in serum. g, The level of insulin in serum. h, HOMA-IR. All data are presented as the mean ± s.e.m. (a) n = 6/group. (b-h) n = 5/group. In a, the P values were determined by two-tailed Student’s t-test. In b, the P values were determined by Kruskal–Wallis test followed by Dunn’s post hoc test. The P values were determined by one-way ANOVA with Tukey’s multiple comparison post hoc test in c-f, and one-way ANOVA with LSD test in g, h.
Extended Data Fig. 7 The ability of Bv-ΔspeA to induce PCOS-like phenotype is significantly reduced compared to B. vulgatus.
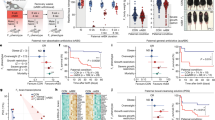
a, The percentage Bacteroides vulgatus strains with or without speA. b, The relative abundance of speA gene in the metagenomic dataset from PMID: 31332392. n = 25 in Control (BMI < 25) group; n = 25 in PCOS (BMI < 25) group; n = 18 in Control (BMI ≥ 25) group; n = 25 in PCOS (BMI ≥ 25) group. For box plots, the midline represents the median; box represents the interquartile range (IQR) between the first and third quartiles, and whiskers represent the minimum to maximum values. Bacteroides vulgatus: Control (BMI < 25) vs PCOS (BMI < 25) P = 2.56 × 10−7, Control (BMI ≥ 25) vs PCOS (BMI ≥ 25) P = 0.000015; Bifidobacterium longum: Control (BMI ≥ 25) vs PCOS (BMI ≥ 25) P = 0.000025. c, DNA gel electrophoresis experiment to verify the knockout of Bv-ΔspeA in B. vulgatus. d, Growth curve of wild type B. vulgatus (Bv) and Bv-ΔspeA in BHI medium. e-m, SPF mice were gavaged with Bv or Bv-ΔspeA for three weeks. e, Colonization of Bv and Bv-ΔspeA in mice. f, Quantitative analysis of corpora lutea in the ovaries. g, Quantitative analysis of cystic follicles in the ovaries. h, Levels of testosterone in serum. i, Levels of LH in serum. j, Area under the curve (AUC) of GTT. k, The levels of fasting glucose in serum. l, The level of insulin in serum. m, HOMA-IR. All data are presented as the mean ± s.e.m. (d) n = 3/group. (e, h-m) n = 6/group. (f, g) n = 5/group. In b, the P values were determined by Kruskal–Wallis test followed by Dunn’s post hoc test. In d, i-m, the P values were determined by two-tailed Student’s t-test. In e-h, the P values were determined by two-tailed Mann–Whitney U-test.
Extended Data Fig. 8 The arginine decarboxylase (ADC) inhibitor DFMA improves the B. vulgatus-induced PCOS-like phenotype.
a, Inhibition rate of agmatine production by adding DFMA to BHI medium. b, Growth curve of Bv and Bv + DFMA in BHI medium. c-l, SPF mice were gavaged with wild type B. vulgatus (Bv), and treated with PBS or arginine decarboxylase inhibitor DFMA for three weeks. c, Quantitative analysis of corpora lutea in the ovaries. d, Quantitative analysis of cystic follicles in the ovaries. e, Levels of testosterone in serum. f, Levels of LH in serum. g, Glucose tolerance test (GTT) (15min: *P = 0.0401; 30min:*P = 0.0203; * compared with the Bv group). h, Area under the curve (AUC) of GTT. i, Insulin tolerance test (ITT) (15min: ***P = 0.000003; 30min: ***P = 0.0006; 60min: *P = 0.0342; 90min: *P =0.0181; * compared with the Bv group). j, The levels of fasting glucose in serum. k, The level of insulin in serum. l, HOMA-IR. All data are presented as the mean ± s.e.m. (a, b) n = 3/group. (c and d) n = 5/group. (e-l) n = 6/group. In b, e-l, the P values were determined by two-tailed Student’s t-test. In c, d, the P values were determined by two-tailed Mann–Whitney U-test.
Extended Data Fig. 9 The arginine decarboxylase (ADC) inhibitor DFMA improves the DHEA-induced hyperandrogenic PCOS-like phenotype.
a, The relative abundance of B. vulgatus in the DHEA model. b, The concentration of agmatine in feces. c, The concentration of agmatine in intestinal epithelium. d, Expression levels of Fgf15 and Shp in intestinal epithelium. e, Levels of total GLP-1 in serum. f, Quantitative analysis of estrous cycles. g, Hematoxylin and eosin staining of representative ovaries. The corpora lutea are indicated by * and the cystic follicle is indicated by #. Scale bar, 200 μm. h, Quantitative analysis of corpora lutea. i, Quantitative analysis of cystic follicles. j, Levels of testosterone in serum. k, Levels of LH in serum. l, Glucose tolerance test (GTT) (15min: ***P = 0.000026, ##P = 0.0034; 30min: ***P = 0.0002, #P = 0.0103; 60min: *P = 0.0297; * compared with the vehicle group, # compared with the DHEA group.). m, Area under the curve (AUC) of GTT. n, Insulin tolerance test (ITT) (15min: **P = 0.0048; 30min: *P = 0.0214; * compared with the vehicle group). o, The levels of fasting glucose in serum. p, The level of insulin in serum. q, HOMA-IR. r, Expression levels of Ucp1, Pgc1a, Cited1, Cox8b, Nr2f6, Prdm16 in the subcutaneous fat. s, Expression levels of Ucp1, Pgc1a, Cited1, Cox8b in the brown adipose tissue. t, Expression levels of Il-8, Il-6, Il-1β, Il-18, Ifng, Ccl20, Ccl2, in the ovaries. All data are presented as the mean ± s.e.m. (a-f, j-t) n = 6/group. (h, i) n = 5/group. In a, the P values were determined by two-tailed Student’s t-test. The P values were determined by one-way ANOVA with Tukey’s multiple comparison post hoc test in c, h-j, l, m, o, q, one-way ANOVA with Dunnett’s T3 test in b and one-way ANOVA with LSD test in k, n. In d-f, p, r-t the P values were determined by Kruskal–Wallis test followed by Dunn’s post hoc test.
Extended Data Fig. 10 The speA and bsh of B. vulgatus can participate in the development of PCOS-like phenotypes in mice independently.
a, DNA gel electrophoresis experiment to verify the knockout of Bv-ΔspeAΔbsh in B. vulgatus. b, Growth curve of wild type B. vulgatus (Bv) and Bv-ΔspeAΔbsh in BHI medium. c, The content of agmatine after 24 h cultured with B. vulgatus or Bv-ΔspeAΔbsh in BHI medium. d, The ability of hydrolyzing TUDCA (12 h) after knocking out both the speA and bsh in B. vulgatus. e, Colonization of Bv and Bv-ΔspeA, Bv-Δbsh and Bv-ΔspeAΔbsh in cecum of mice. f, Quantitative analysis of cystic follicles in the ovaries. g, Area under the curve (AUC) of GTT. h, The levels of fasting glucose in serum. i, The level of insulin in serum. j, Expression levels of Ucp1, Pgc1a, Cited1, Cox8b, Nr2f6, Prdm16 in the subcutaneous fat. Ucp1: Vehicle versus Bv, P = 8.06 × 10−7; Bv versus Bv-ΔspeAΔbsh, P = 8.09 × 10−7. k, Expression levels of Ucp1, Pgc1a, Cited1, Cox8b in the brown adipose tissue. Ucp1: Vehicle versus Bv, P = 0.000019. l, Expression levels of Il-8, Il-6, Il-1β, Il-18, Ifng, Ccl20, Ccl2, in the ovaries. All data are presented as the mean ± s.e.m. (b, e, g-l) n = 6/group. (c, d) n = 3/group. (f) n = 5/group. The P values were determined by one-way ANOVA with Dunnett’s T3 test in c, d, one-way ANOVA with Tukey’s multiple comparison post hoc test in g, k and one-way ANOVA with LSD test in f, h-j. In e, l, the P values were determined by Kruskal–Wallis test followed by Dunn’s post hoc test.
Supplementary information
Supplementary Information
Supplementary Figs. 1–4 and Tables 1 and 2.
Source data
Source Data Fig. 1
Statistical source data.
Source Data Fig. 2
Statistical source data.
Source Data Fig. 3
Statistical source data.
Source Data Fig. 4
Statistical source data.
Source Data Fig. 5
Statistical source data.
Source Data Fig. 6
Statistical source data.
Source Data Fig. 7
Statistical source data.
Source Data Extended Data Fig. 1
Statistical source data.
Source Data Extended Data Fig. 1
Unprocessed gels.
Source Data Extended Data Fig. 2
Statistical source data.
Source Data Extended Data Fig. 3
Statistical source data.
Source Data Extended Data Fig. 4
Statistical source data.
Source Data Extended Data Fig. 5
Statistical source data.
Source Data Extended Data Fig. 6
Statistical source data.
Source Data Extended Data Fig. 7
Statistical source data.
Source Data Extended Data Fig. 7
Unprocessed gels.
Source Data Extended Data Fig. 8
Statistical source data.
Source Data Extended Data Fig. 9
Statistical source data.
Source Data Extended Data Fig. 10
Statistical source data.
Source Data Extended Data Fig. 10
Unprocessed gels.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Yun, C., Yan, S., Liao, B. et al. The microbial metabolite agmatine acts as an FXR agonist to promote polycystic ovary syndrome in female mice. Nat Metab 6, 947–962 (2024). https://doi.org/10.1038/s42255-024-01041-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s42255-024-01041-8