Abstract
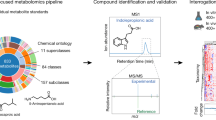
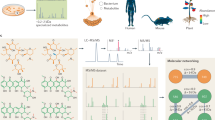
A significant hurdle that has limited progress in microbiome science has been identifying and studying the diverse set of metabolites produced by gut microbes. Gut microbial metabolism produces thousands of difficult-to-identify metabolites, which present a challenge to study their roles in host biology. In recent years, mass spectrometry-based metabolomics has become one of the core technologies for identifying small metabolites. However, metabolomics expertise, ranging from sample preparation to instrument use and data analysis, is often lacking in academic labs. Most targeted metabolomics methods provide high levels of sensitivity and quantification, while they are limited to a panel of predefined molecules that may not be informative to microbiome-focused studies. Here we have developed a gut microbe-focused and wide-spectrum metabolomic protocol using liquid chromatography–mass spectrometry and bioinformatic analysis. This protocol enables users to carry out experiments from sample collection to data analysis, only requiring access to a liquid chromatography–mass spectrometry instrument, which is often available at local core facilities. By applying this protocol to samples containing human gut microbial metabolites, spanning from culture supernatant to human biospecimens, our approach enables high-confidence identification of >800 metabolites that can serve as candidate mediators of microbe–host interactions. We expect this protocol will lower the barrier to tracking gut bacterial metabolism in vitro and in mammalian hosts, propelling hypothesis-driven mechanistic studies and accelerating our understanding of the gut microbiome at the chemical level.
Key points
-
This protocol describes a method for detecting and measuring metabolites derived from the gut microbiota in diverse biological samples.
-
The protocol combines untargeted data collection methods with a targeted data analysis pipeline, allowing characterization of a broad range of microbial metabolites.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
The sample metabolomics data required for bioinformatics analysis are available on GitHub (https://github.com/the-han-lab/Han_et_al_Metabolomics_Protocol_2022). We have deposited our MS2 spectra data on the community-accepted MoNA public database, but the MoNA search algorithm is not optimized to return only the relevant data submitted by our lab; to enable quick user access, we have also deposited the full MS2 data in our lab’s GitHub repository (given above). All metabolomics raw data associated with our primary publication15 are publicly accessible on the Metabolomics Workbench under study number ST001683 for all in vivo data and study number ST001688 for in vitro data. New metabolomics data generated for this protocol are also available on the same platform under study number ST002973.
Code availability
The code required for bioinformatics analysis is available on GitHub (https://github.com/the-han-lab/Han_et_al_Metabolomics_Protocol_2022).
References
Rooks, M. G. & Garrett, W. S. Gut microbiota, metabolites and host immunity. Nat. Rev. Immunol. 16, 341–352 (2016).
Cani, P. D. Microbiota and metabolites in metabolic diseases. Nat. Rev. Endocrinol. 15, 69–70 (2019).
Sonnenburg, J. L. & Backhed, F. Diet–microbiota interactions as moderators of human metabolism. Nature 535, 56–64 (2016).
Kasahara, K. & Rey, F. E. The emerging role of gut microbial metabolism on cardiovascular disease. Curr. Opin. Microbiol. 50, 64–70 (2019).
Lynch, J. B. & Hsiao, E. Y. Microbiomes as sources of emergent host phenotypes. Science 365, 1405–1409 (2019).
Koppel, N., Maini Rekdal, V. & Balskus, E.P. Chemical transformation of xenobiotics by the human gut microbiota. Science https://doi.org/10.1126/science.aag2770 (2017).
Koh, A., De Vadder, F., Kovatcheva-Datchary, P. & Backhed, F. From dietary fiber to host physiology: short-chain fatty acids as key bacterial metabolites. Cell 165, 1332–1345 (2016).
Donia, M. S. & Fischbach, M. A. Human microbiota. Small molecules from the human microbiota. Science 349, 1254766 (2015).
Husted, A. S., Trauelsen, M., Rudenko, O., Hjorth, S. A. & Schwartz, T. W. GPCR-mediated signaling of metabolites. Cell Metab. 25, 777–796 (2017).
Krautkramer, K. A., Fan, J. & Backhed, F. Gut microbial metabolites as multi-kingdom intermediates. Nat. Rev. Microbiol. 19, 77–94 (2021).
Bauermeister, A., Mannochio-Russo, H., Costa-Lotufo, L. V., Jarmusch, A. K. & Dorrestein, P. C. Mass spectrometry-based metabolomics in microbiome investigations. Nat. Rev. Microbiol. 20, 143–160 (2022).
Guijas, C. et al. METLIN: a technology platform for identifying knowns and unknowns. Anal. Chem. 90, 3156–3164 (2018).
Wang, M. et al. Sharing and community curation of mass spectrometry data with Global Natural Products Social Molecular Networking. Nat. Biotechnol. 34, 828–837 (2016).
Melnik, A. V. et al. Coupling targeted and untargeted mass spectrometry for metabolome–microbiome-wide association studies of human fecal samples. Anal. Chem. 89, 7549–7559 (2017).
Han, S. et al. A metabolomics pipeline for the mechanistic interrogation of the gut microbiome. Nature 595, 415–420 (2021).
Showalter, M. R. et al. Obesogenic diets alter metabolism in mice. PLoS ONE 13, e0190632 (2018).
Evans, A. M. et al. High resolution mass spectrometry improves data quantity and quality as compared to unit mass resolution mass spectrometry in high-throughput profiling metabolomics. Metabolomics https://doi.org/10.4172/2153-0769.1000132 (2014).
Mars, R. A. T. et al. Longitudinal multi-omics reveals subset-specific mechanisms underlying irritable bowel syndrome. Cell 183, 1137–1140 (2020).
Holmes, E., Wijeyesekera, A., Taylor-Robinson, S. D. & Nicholson, J. K. The promise of metabolic phenotyping in gastroenterology and hepatology. Nat. Rev. Gastroenterol. Hepatol. 12, 458–471 (2015).
Fiehn, O. Metabolomics by gas chromatography–mass spectrometry: combined targeted and untargeted profiling. Curr. Protoc. Mol. Biol. 114, 303431–303432 (2016).
Rinschen, M. M., Ivanisevic, J., Giera, M. & Siuzdak, G. Identification of bioactive metabolites using activity metabolomics. Nat. Rev. Mol. Cell Biol. 20, 353–367 (2019).
Liebisch, G. et al. Quantification of fecal short chain fatty acids by liquid chromatography tandem mass spectrometry-investigation of pre-analytic stability. Biomolecules https://doi.org/10.3390/biom9040121 (2019).
Boulet, L., Faure, P., Flore, P., Monteremal, J. & Ducros, V. Simultaneous determination of tryptophan and 8 metabolites in human plasma by liquid chromatography/tandem mass spectrometry. J. Chromatogr. B 1054, 36–43 (2017).
Wegner, K. et al. Rapid analysis of bile acids in different biological matrices using LC–ESI–MS/MS for the investigation of bile acid transformation by mammalian gut bacteria. Anal. Bioanal. Chem. 409, 1231–1245 (2017).
Ayon, N. J., Sharma, A. D. & Gutheil, W. G. LC–MS/MS-based separation and quantification of Marfey’s reagent derivatized proteinogenic amino acid dl-stereoisomers. J. Am. Soc. Mass Spectrom. 30, 448–458 (2019).
Pismennoi, D., Kiritsenko, V., Marhivka, J., Kutt, M.L. & Vilu, R. Development and Optimisation of HILIC–LC–MS method for determination of carbohydrates in fermentation samples. Molecules https://doi.org/10.3390/molecules26123669 (2021).
Kind, T. et al. Identification of small molecules using accurate mass MS/MS search. Mass Spectrom. Rev. 37, 513–532 (2018).
Schymanski, E. L. et al. Identifying small molecules via high resolution mass spectrometry: communicating confidence. Environ. Sci. Technol. 48, 2097–2098 (2014).
Pandher, R., Naegele, E., Fischer, S.M. & Raynaud, F.I. in Metabolomics (eds Roessner, U.) 318–328 (InTech, 2012).
Sousa, M. B. & Ziegler, T. E. Diurnal variation on the excretion patterns of fecal steroids in common marmoset (Callithrix jacchus) females. Am. J. Primatol. 46, 105–117 (1998).
Tsugawa, H. et al. MS-DIAL: data-independent MS/MS deconvolution for comprehensive metabolome analysis. Nat. Methods 12, 523–526 (2015).
Cajka, T., Smilowitz, J. T. & Fiehn, O. Validating quantitative untargeted lipidomics across nine liquid chromatography–high-resolution mass spectrometry platforms. Anal. Chem. 89, 12360–12368 (2017).
Wastyk, H. C. et al. Gut-microbiota-targeted diets modulate human immune status. Cell 184, 4137–4153 e4114 (2021).
Kotagale, N. R., Taksande, B. G. & Inamdar, N. N. Neuroprotective offerings by agmatine. Neurotoxicology 73, 228–245 (2019).
Chin, R. M. et al. The metabolite alpha-ketoglutarate extends lifespan by inhibiting ATP synthase and TOR. Nature 510, 397–401 (2014).
Acknowledgements
We thank Stanford ChEM-H for use of the Agilent 6545 LC/Q-TOF instrument, and W. Van Treuren, D. Dodd, C. R. Fischer, B. C. DeFelice, J. M. Sanchez, M. A. Fischbach and A. Stewart for helpful input on method development and optimization. This work is supported by R01-DK085025, DP1-AT009892, R01-DK101674 and a Stanford Discovery Innovation Fund Award (J.L.S.); Mallinckrodt Foundation Grant, Glenn Foundation for Medical Research and AFAR Grants for Junior Faculty, Duke Science and Technology Initiative and Whitehead Scholar (S.H.); 5T32AI007328-35 (E.R.G.); and Duke University School of Medicine Biomedical PhD Student Research Pilot Grant (Y.L.). J.L.S. is a Chan-Zuckerberg Biohub Investigator.
Author information
Authors and Affiliations
Contributions
S.H., E.R.G. and J.L.S. conceived the study and wrote the paper. S.H., E.R.G. and Y.L. generated the figures. S.H. developed methods for building the MS1 and MS2 mass spectrometry reference libraries, metabolite extraction and data analysis via MS-DIAL, conducted bioinformatics analyses and built the bioinformatics pipeline integrating data from diverse sample types. Y.L. optimized and validated the metabolomics analysis workflow using an independent orbitrap instrument, conducted metabolomic experiments and performed data analyses.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Protocols thanks Thomas Horvath and Martin Giera for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Related links
Key references in the development of the protocol
Han, S. et al. Nature 595, 415–420 (2021): https://doi.org/10.1038/s41586-021-03707-9
Mars, R. A. T. et al. Cell 182, 1460–1473.e17 (2020): https://doi.org/10.1016/j.cell.2020.08.007
Wastyk, H. C. et al. Cell 184, 4137–4153.e14 (2021): https://doi.org/10.1016/j.cell.2021.06.019
Extended data
Extended Data Fig. 1 Example images of extracted ion chromatograms generated in the Agilent MassHunter Qualitative Data Analysis software.
a, Example of a single peak identified from extracting a chromatogram from a single m/z value. b, Example of multiple peaks identified from extracting a chromatogram based on a single m/z value. c, Example of no peak identified from extracting a chromatogram based on a single m/z value.
Extended Data Fig. 2 Pre- and post-filtering comparisons in peak areas.
Pairwise molecular feature comparisons before and after the mega media being passed through a 0.2 μm membrane filter. a, b, Two independent batches of mega media have been tested (n = 5 for each of pre- and post-filtered conditions for each batch) using the C18 positive method. 1882 unique molecular features are reported for both batches following the background noise elimination and peak quality checks. Linear regression and r2 values of pairwise molecular feature comparisons, performed on log10 transformed peak area data: pre-filtered versus post-filtered in batch 1 (r2 = 0.977, Post-filtered peak area = 1 × Pre-filtered peak area - 0.0109, a) and batch 2 (r2 = 0.994, Post-filtered peak area = 0.9974 × Pre-filtered peak area + 0.0206, b).
Extended Data Fig. 3 Agilent qTOF mass calibration settings on Agilent MassHunter.
Example calibration shown is in the negative ionization mode (e.g., C18 negative method). For the positive ionization mode (C18 positive, HILIC positive), these calibration settings remain identical with the exception of shifting ion polarity from “negative” to “positive” in both “Ion polarity” and “qTOF” sections. a, Tune and calibration. b, Instrument state. Red boxes highlight each tab in the software that corresponds to each type of settings.
Extended Data Fig. 4 Project settings using the MS-DIAL software.
a, General project information. b, Project file information. Red boxes and associated arrows highlight explanations for specific input.
Extended Data Fig. 5 MS-DIAL analysis parameters and alignment result export settings.
a, Data collection. b, Peak detection. c, MS2 deconvolution (not used in this protocol). d, Identification. e, Adduct (not modified here, but information is provided in the reference library input file in “Identification.”) f, Alignment. g, Isotope tracking (not used in this protocol). h, Alignment result export. Red boxes and associated arrows highlight explanations for specific input.
Extended Data Fig. 6 Residual plots for RT, precursor ion m/z, and peak area of a panel of internal standards detected in individual biological samples from a representative mouse serum metabolomic study.
a, b, Residual plot for linear regression model constructed using RTs (a) or precursor ion m/z values (b). Residuals are calculated based on values from individual biological samples vs. reference library. c, Residual plot for linear regression model constructed on peak areas of extracted ion chromatograms. Residuals are calculated based on detected values from individual biological samples (log10-transformed) vs. QC samples (log10-transformed). A single dot represents individual biological replicate per condition (e.g., germ-free or mono-association). Overlapping dots, which represent identical values, are shown in darker shades than the non-overlapping dots.
Extended Data Fig. 7 MS-DIAL user interface for checking the quality of annotated peaks.
a, Example of visualizing an analyzed dataset on MS-DIAL. b, Example of a low-quality peak to be eliminated from the output result file. Orange boxes highlight specific items to select for visualization. Red arrows highlight specific explanations or instructions.
Supplementary information
Supplementary Information
Supplementary guide for Supplementary Tables 1 and 2.
Supplementary Table 1
Bacterial metabolites.
Supplementary Table 2
Co-eluting metabolites.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Han, S., Guiberson, E.R., Li, Y. et al. High-throughput identification of gut microbiome-dependent metabolites. Nat Protoc (2024). https://doi.org/10.1038/s41596-024-00980-6
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41596-024-00980-6
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.