Abstract
Lysosomal transmembrane acetylation of heparan sulfates (HS) is catalyzed by HS acetyl-CoA:α-glucosaminide N-acetyltransferase (HGSNAT), whose dysfunction leads to lysosomal storage diseases. The mechanism by which HGSNAT, the sole non-hydrolase enzyme in HS degradation, brings cytosolic acetyl-coenzyme A (Ac-CoA) and lysosomal HS together for N-acyltransferase reactions remains unclear. Here, we present cryogenic-electron microscopy structures of HGSNAT alone, complexed with Ac-CoA and with acetylated products. These structures explain that Ac-CoA binding from the cytosolic side causes dimeric HGSNAT to form a transmembrane tunnel. Within this tunnel, catalytic histidine and asparagine approach the lumen and instigate the transfer of the acetyl group from Ac-CoA to the glucosamine group of HS. Our study unveils a transmembrane acetylation mechanism that may help advance therapeutic strategies targeting lysosomal storage diseases.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Data availability
The sequence of human HGSNAT is available at the following link: https://www.uniprot.org/uniprotkb/Q68CP4/entry#sequences. The cryo-EM maps and coordinates of apo-HGSNAT, Ac-CoA-HGSNAT, CoA/NAG-HGSNAT and CoA/4MU-NAG-HGSNAT complexes have been deposited in the Electron Microscopy Data Bank (EMDB) under accession numbers EMD-36376, EMD-36386, EMD-36387 and EMD-37264 and in the Protein Data Bank (PDB) under accession codes 8JKV, 8JL1, 8JL3 and 8W4A, respectively. Source data are provided with this paper.
References
Parish, C. R. The role of heparan sulphate in inflammation. Nat. Rev. Immunol. 6, 633–643 (2006).
Sasisekharan, R., Shriver, Z., Venkataraman, G. & Narayanasami, U. Roles of heparan-sulphate glycosaminoglycans in cancer. Nat. Rev. Cancer 2, 521–528 (2002).
Simon Davis, D. A. & Parish, C. R. Heparan sulfate: a ubiquitous glycosaminoglycan with multiple roles in immunity. Front Immunol. 4, 470 (2013).
Long, K. R. & Huttner, W. B. How the extracellular matrix shapes neural development. Open Biol. 9, 180216 (2019).
Freeman, C. & Hopwood, J. Lysosomal degradation of heparin and heparan sulphate. Adv. Exp. Med. Biol. 313, 121–134 (1992).
Griffin, L. S. & Gloster, T. M. The enzymatic degradation of heparan sulfate. Protein Pept. Lett. 24, 710–722 (2017).
Andrade, F., Aldamiz-Echevarria, L., Llarena, M. & Couce, M. L. Sanfilippo syndrome: overall review. Pediatr. Int. 57, 331–338 (2015).
Bame, K. J. & Rome, L. H. Acetyl coenzyme A:alpha-glucosaminide N-acetyltransferase. Evidence for a transmembrane acetylation mechanism. J. Biol. Chem. 260, 11293–11299 (1985).
Bame, K. J. & Rome, L. H. Acetyl-coenzyme A:alpha-glucosaminide N-acetyltransferase. Evidence for an active site histidine residue. J. Biol. Chem. 261, 10127–10132 (1986).
Pshezhetsky, A. V. Lysosomal storage of heparan sulfate causes mitochondrial defects, altered autophagy, and neuronal death in the mouse model of mucopolysaccharidosis III type C. Autophagy 12, 1059–1060 (2016).
Hrebicek, M. et al. Mutations in TMEM76* cause mucopolysaccharidosis IIIC (Sanfilippo C syndrome). Am. J. Hum. Genet. 79, 807–819 (2006).
Meikle, P. J., Whittle, A. M. & Hopwood, J. J. Human acetyl-coenzyme A:alpha-glucosaminide N-acetyltransferase. Kinetic characterization and mechanistic interpretation. Biochem. J. 308, 327–333 (1995).
Fan, X. et al. Identification of the gene encoding the enzyme deficient in mucopolysaccharidosis IIIC (Sanfilippo disease type C). Am. J. Hum. Genet. 79, 738–744 (2006).
Fedele, A. O. Sanfilippo syndrome: causes, consequences, and treatments. Appl Clin. Genet. 8, 269–281 (2015).
Platt, F. M. Emptying the stores: lysosomal diseases and therapeutic strategies. Nat. Rev. Drug Discov. 17, 133–150 (2018).
Martins, C. et al. Molecular characterization of a large group of mucopolysaccharidosis type IIIC patients reveals the evolutionary history of the disease. Hum. Mutat. 40, 1084–1100 (2019).
Feldhammer, M., Durand, S. & Pshezhetsky, A. V. Protein misfolding as an underlying molecular defect in mucopolysaccharidosis III type C. PLoS ONE 4, e7434 (2009).
Fedele, A. O. & Hopwood, J. J. Functional analysis of the HGSNAT gene in patients with mucopolysaccharidosis IIIC (Sanfilippo C Syndrome). Hum. Mutat. 31, E1574–E1586 (2010).
Pan, X. et al. Glucosamine amends CNS pathology in mucopolysaccharidosis IIIC mouse expressing misfolded HGSNAT. J. Exp. Med. 219, e20211860 (2022).
Spahiu, L. et al. Mucopolysaccharidosis III: molecular basis and treatment. Pediatr. Endocrinol. Diabetes Metab. 27, 201–208 (2021).
Feldhammer, M. et al. Sanfilippo syndrome type C: mutation spectrum in the heparan sulfate acetyl-CoA:alpha-glucosaminide N-acetyltransferase (HGSNAT) gene. Hum. Mutat. 30, 918–925 (2009).
Durand, S., Feldhammer, M., Bonneil, E., Thibault, P. & Pshezhetsky, A. V. Analysis of the biogenesis of heparan sulfate acetyl-CoA:alpha-glucosaminide N-acetyltransferase provides insights into the mechanism underlying its complete deficiency in mucopolysaccharidosis IIIC. J. Biol. Chem. 285, 31233–31242 (2010).
Marmorstein, R. Structure and function of histone acetyltransferases. Cell. Mol. Life Sci. 58, 693–703 (2001).
Marmorstein, R. & Roth, S. Y. Histone acetyltransferases: function, structure, and catalysis. Curr. Opin. Genet Dev. 11, 155–161 (2001).
Salah Ud-Din, A. I., Tikhomirova, A. & Roujeinikova, A. Structure and functional diversity of GCN5-related N-acetyltransferases (GNAT). Int. J. Mol. Sci. https://doi.org/10.3390/ijms17071018 (2016).
Christensen, D. G. et al. Post-translational protein acetylation: an elegant mechanism for bacteria to dynamically regulate metabolic functions. Front. Microbiol. 10, 1604 (2019).
Navratna, V., Kumar, A. & Mosalaganti, S. Structure of the human heparan-alpha-glucosaminide N-acetyltransferase (HGSNAT). eLife https://doi.org/10.7554/eLife.93510.1 (2024).
Ma, D. et al. Crystal structure of a membrane-bound O-acyltransferase. Nature 562, 286–290 (2018).
Guan, C. et al. Structural insights into the inhibition mechanism of human sterol O-acyltransferase 1 by a competitive inhibitor. Nat. Commun. 11, 2478 (2020).
Jiang, Y., Benz, T. L. & Long, S. B. Substrate and product complexes reveal mechanisms of Hedgehog acylation by HHAT. Science 372, 1215–1219 (2021).
Liu, Y. et al. Mechanisms and inhibition of Porcupine-mediated Wnt acylation. Nature 607, 816–822 (2022).
Qian, H. et al. Structural basis for catalysis and substrate specificity of human ACAT1. Nature 581, 333–338 (2020).
Wang, L. et al. Structure and mechanism of human diacylglycerol O-acyltransferase 1. Nature 581, 329–332 (2020).
Dutnall, R. N., Tafrov, S. T., Sternglanz, R. & Ramakrishnan, V. Structure of the histone acetyltransferase Hat1: a paradigm for the GCN5-related N-acetyltransferase superfamily. Cell 94, 427–438 (1998).
Chen, J. Y. et al. Structure and function of human Naa60 (NatF), a Golgi-localized bi-functional acetyltransferase. Sci. Rep. 6, 31425 (2016).
Liszczak, G., Arnesen, T. & Marmorstein, R. Structure of a ternary Naa50p (NAT5/SAN) N-terminal acetyltransferase complex reveals the molecular basis for substrate-specific acetylation. J. Biol. Chem. 286, 37002–37010 (2011).
Wolf, E. et al. Crystal structure of a GCN5-related N-acetyltransferase: Serratia marcescens aminoglycoside 3-N-acetyltransferase. Cell 94, 439–449 (1998).
Leaback, D. H. & Walker, P. G. Studies on glucosaminidase. 4. The fluorimetric assay of N-acetyl-beta-glucosaminidase. Biochem. J. 78, 151–156 (1961).
Voznyi, Ya,V. et al. A fluorimetric enzyme assay for the diagnosis of Sanfilippo disease C (MPS III C). J. Inherit. Metab. Dis. 16, 465–472 (1993).
Goehring, A. et al. Screening and large-scale expression of membrane proteins in mammalian cells for structural studies. Nat. Protoc. 9, 2574–2585 (2014).
Kawate, T. & Gouaux, E. Fluorescence-detection size-exclusion chromatography for precrystallization screening of integral membrane proteins. Structure 14, 673–681 (2006).
Wang, R. et al. Global stable-isotope tracing metabolomics reveals system-wide metabolic alternations in aging Drosophila. Nat. Commun. 13, 3518 (2022).
Punjani, A., Rubinstein, J. L., Fleet, D. J. & Brubaker, M. A. cryoSPARC: algorithms for rapid unsupervised cryo-EM structure determination. Nat. Methods 14, 290–296 (2017).
Goddard, T. D. et al. UCSF ChimeraX: meeting modern challenges in visualization and analysis. Protein Sci. 27, 14–25 (2018).
Punjani, A., Zhang, H. & Fleet, D. J. Non-uniform refinement: adaptive regularization improves single-particle cryo-EM reconstruction. Nat. Methods 17, 1214–1221 (2020).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D. Biol. Crystallogr. 60, 2126–2132 (2004).
Afonine, P. V. et al. Real-space refinement in PHENIX for cryo-EM and crystallography. Acta Crystallogr. D. Struct. Biol. 74, 531–544 (2018).
Chen, V. B. et al. MolProbity: all-atom structure validation for macromolecular crystallography. Acta Crystallogr. D. Biol. Crystallogr. 66, 12–21 (2010).
The PyMOL Molecular Graphics System. v.2.3.5 (Schrodinger, 2020).
Smart, O. S., Goodfellow, J. M. & Wallace, B. A. The pore dimensions of gramicidin A. Biophys. J. 65, 2455–2460 (1993).
Acknowledgements
We thank the Cryo-Electron Microscopy Center at the Interdisciplinary Research Center on Biology and Chemistry, Shanghai Institute of Organic Chemistry for the help with data collection. We thank ShanghaiTech Cryo-EM Imaging Facility for its assistance in data collection. We thank the Molecular and Cell Biology Core Facility at the School of Life Science and Technology, ShanghaiTech University, for providing technical support. This work was supported by Shanghai Lingang Laboratory (grant nos. LG-QS-202203-03 to J.Y. and LG-QS-202203-05 to J.G.) and Shanghai Pujiang Program (grant no. 22PJ1410300 to J.G.).
Author information
Authors and Affiliations
Contributions
J.Y. and J.G. designed the project. R.X. and Y.N. performed the sample preparation for cryo-EM. R.X., Y.N. and C.G. performed the biochemistry studies. R.X. performed the fluorescence-based acetylation assay. F.R. and Z.Z. carried out MS analysis. J.Y. and J.G. performed the cryo-EM data collection, data analysis and model building. J.Y., J.G., A.V.P. and X.P. wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Structural & Molecular Biology thanks the anonymous reviewers for their contribution to the peer review of this work. Peer reviewer reports are available. Primary Handling Editor: Katarzyna Ciazynska, in collaboration with the Nature Structural & Molecular Biology team.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
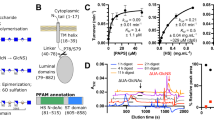
Extended Data Fig. 1 Biochemical characterization and cryo-EM analysis of human HGSNAT.
a. Topology of HGSNAT. Unresolved regions are marked with dashed lines. b. Representative size exclusion chromatography (SEC) profile (left) of HGSNAT and SDS-PAGE gel (right) of HGSNAT in the presence or absence of β-ME. The trace and gel image are representative of 4 experimental replicates. c. A representative cryo-EM micrograph of HGSNAT. (out of 30,545 similar micrographs). d. Selected representative 2D class averages.
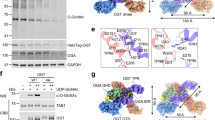
Extended Data Fig. 2 A representative flow chart of data processing and representative densities of the HGSNAT map.
a. Flow chart of data processing for Ac-CoA-bound HGSNAT complexes. Details may be found in Methods. b. Cryo-EM densities of the transmembrane helixes in the Ac-CoA-bound HGSNAT complexes.
Extended Data Fig. 3 3D reconstructions of the apo, Ac-CoA-bound, CoA/NAG-bound, and CoA/4MU-NAG-bound HGSNAT complexes.
a, e, j, o. Local resolution estimates of the apo (a), Ac-CoA-bound (e), CoA/NAG-bound (j), and CoA/4MU-NAG-bound (o) HGSNAT maps. b, f, k, p. FSC curves before and after masking and between the model and the final maps of the apo (b), Ac-CoA-bound (f), CoA/NAG-bound (k), and CoA/4MU-NAG-bound (p) HGSNAT. c, g, l, q. Projection orientation distribution of apo (c), Ac-CoA-bound (g), CoA/NAG-bound (l), and CoA/4MU-NAG-bound (q) HGSNAT maps, generated from cryoSPARC. d, h, m, r. The 3DFSC sphericity of the apo (d), Ac-CoA-bound (h), CoA/NAG-bound (m), and CoA/4MU-NAG-bound (r) HGSNAT maps, analyzed with 3DFSC in cryoSPARC. i, n, s. Densities of the substrates determined in the Ac-CoA-bound (i), CoA/NAG-bound (n), and CoA/4MU-NAG-bound (s) HGSNAT complexes. t. Densities of the Met282, Phe313, Lys341, and Arg345 in the apo and Ac-CoA-bound HGSNAT structures to illustrate the repositioning of these four residues upon the binding of Ac-CoA.
Extended Data Fig. 4 Effects of the indicated mutants on the activity of HGSNAT.
a. The relative enzyme activities of the indicated mutants. Data are shown as mean ± standard deviations, n = 3 biologically independent samples. b–s. Size exclusion chromatography profiles of the HGSNAT mutants.
Extended Data Fig. 5 Thermostability of wild type HGSNAT and HGSNAT mutants.
a–d. Melting curves of wild type HGSNAT, indicated HGSNAT mutants and CFP. The CFP tag, located at the C-terminal of the wild type HGSNAT and mutants, was used for protein detection. Data are shown as mean ± standard deviations, n = 3 biologically independent samples. e. The interactions in the NTD-TMD interface. Possible hydrogen bonds are indicated with dashed lines.
Extended Data Fig. 6 LC-MS identification of the endogenous Ac-CoA in HGSNAT.
a. LC profiles of commercial standard Ac-CoA (top) and purified HGSNAT enzyme from the solubilization of the whole cells without any additional dialysis (bottom). b. MS/MS spectrum of commercial standard Ac-CoA (top) and purified HGSNAT enzyme from the solubilization of the whole cells without any additional dialysis (bottom).
Extended Data Fig. 7 Sequence alignment of the HGSNAT across various species.
Secondary elements are labeled above the alignment. The catalytic residues Asn286 and His297 are highlighted and indicated with red stars.
Extended Data Fig. 8 The comparisons of the overall structure and the positions of catalytic site between HGSNAT with lipid-modifying MBOATs.
a–e. The overall structure of HGSNAT (a) in comparison with HHAT (b), Porcupine (c), ACAT1 (d), and DGAT1 (e). The acyl-CoA and hallmark catalytic histidine in each enzyme are indicated and shown in ball and stick style.
Supplementary information
Supplementary Video
Illustrating the transmembrane acetylation reaction catalyzed by HGSNAT.
Source data
Source Data Fig. 1
Statistical source data.
Source Data Fig. 2
Statistical source data.
Source Data Fig. 3
Statistical source data.
Source Data Extended Data Fig. 1
Unprocessed gels.
Source Data Extended Data Fig. 3
Statistical source data.
Source Data Extended Data Fig. 4
Statistical source data.
Source Data Extended Data Fig. 5
Statistical source data.
Source Data Extended Data Fig. 6
Statistical source data.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Xu, R., Ning, Y., Ren, F. et al. Structure and mechanism of lysosome transmembrane acetylation by HGSNAT. Nat Struct Mol Biol (2024). https://doi.org/10.1038/s41594-024-01315-5
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41594-024-01315-5