Abstract
Although superresolution (SR) approaches have been routinely used for fixed or living material from other organisms, the use of time-lapse structured illumination microscopy (SIM) imaging in plant cells still remains under-developed. Here we describe a validated method for time-lapse SIM that focuses on cortical microtubules of different plant cell types. By using one of the existing commercially available SIM platforms, we provide a user-friendly and easy-to-follow protocol that may be widely applied to the imaging of plant cells. This protocol includes steps describing calibration of the microscope and channel alignment, generation of an experimental point spread function (PSF), preparation of appropriate observation chambers with available plant material, image acquisition, reconstruction and validation. This protocol can be carried out within two to three working days.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
Renz, M. Fluorescence microscopy-a historical and technical perspective. Cytometry A 83, 767–779 (2013).
Terai, T. & Nagano, T. Small-molecule fluorophores and fluorescent probes for bioimaging. Pflugers Arch. 465, 347–359 (2013).
Zhang, J. The colorful journey of green fluorescent protein. ACS Chem. Biol. 4, 85–88 (2009).
Amos, W.B. & White, J.G. How the confocal laser-scanning microscope entered biological research. Biol. Cell. 95, 335–342 (2003).
Stehbens, S., Pemble, H., Murrow, L. & Wittmann, T. Imaging intracellular protein dynamics by spinning disk confocal microscopy. Methods Enzymol. 504, 293–313 (2012).
Vizcay-Barrena, G., Webb, S.E., Martin-Fernandez, M.L. & Wilson, Z.A. Subcellular and single-molecule imaging of plant fluorescent proteins using total internal reflection fluorescence microscopy (TIRFM). J. Exp. Bot. 62, 5419–5428 (2011).
Gao, L., Shao, L., Chen, B.C. & Betzig, E. 3D live fluorescence imaging of cellular dynamics using Bessel beam plane illumination microscopy. Nat. Protoc. 9, 1083–1101 (2014).
Weber, M., Mickoleit, M. & Huisken, J. Light sheet microscopy. Methods Cell Biol. 123, 193–215 (2014).
Wang, X. et al. Imaging of dynamic secretory vesicles in living pollen tubes of Picea meyeri using evanescent wave microscopy. Plant Physiol. 141, 1591–1603 (2006).
Requejo-Isidro, J. Fluorescence nanoscopy. Methods and applications. J. Chem. Biol. 6, 97–120 (2013).
Engel, U. Structured illumination superresolution imaging of the cytoskeleton. Methods Cell Biol. 123, 315–333 (2014).
Gustafsson, M.G. Surpassing the lateral resolution limit by a factor of two using structured illumination microscopy. J. Microsc. 198, 82–87 (2000).
Heintzmann, R. & Gustafsson, M.G.L. Subdiffraction resolution in continuous samples. Nat. Photon. 3, 362–364 (2009).
Winter, P.W. & Shroff, H. Faster fluorescence microscopy: advances in high speed biological imaging. Curr. Opin. Chem. Biol. 20, 46–53 (2014).
Kusumi, A., Tsunoyama, T.A., Hirosawa, K.M., Kasai, R.S. & Fujiwara, T.K. Tracking single molecules at work in living cells. Nat. Chem. Biol. 10, 524–532 (2014).
Hensel, M., Klingauf, J. & Piehler, J. Imaging the invisible: resolving cellular microcompartments by superresolution microscopy techniques. Biol. Chem. 394, 1097–1113 (2013).
Sengupta, P., Van Engelenburg, S. & Lippincott-Schwartz, J. Visualizing cell structure and function with point-localization superresolution imaging. Dev. Cell. 23, 1092–1102 (2012).
Verdaasdonk, J.S., Stephens, A.D., Haase, J. & Bloom, K. Bending the rules: widefield microscopy and the Abbe limit of resolution. J. Cell Physiol. 229, 132–138 (2014).
Schermelleh, L., Heintzmann, R. & Leonhardt, H. A guide to super-resolution fluorescence microscopy. J. Cell Biol. 190, 165–175 (2010).
Fiolka, R. Seeing more with structured illumination microscopy. Methods Cell Biol. 123, 295–313 (2014).
Buschmann, H., Sambade, A., Pesquet, E., Calder, G. & Lloyd, C.W. Microtubule dynamics in plant cells. Methods Cell Biol. 97, 373–400 (2010).
Komis, G. et al. Dynamics and organization of cortical microtubules as revealed by superresolution structured illumination microscopy. Plant Physiol. 165, 129–148 (2014).
Marc, J. et al. A GFP-MAP4 reporter gene for visualizing cortical microtubule rearrangements in living epidermal cells. Plant Cell 10, 1927–1940 (1998).
Beck, M., Komis, G., Ziemann, A., Menzel, D. & Šamaj, J. Mitogen-activated protein kinase 4 is involved in the regulation of mitotic and cytokinetic microtubule transitions in Arabidopsis thaliana. New Phytol. 189, 1069–1083 (2011).
Shaw, S.L., Kamyar, R. & Ehrhardt, D.W. Sustained microtubule treadmilling in Arabidopsis cortical arrays. Science 300, 1715–1718 (2003).
Phillips, D., Nibau, C., Wnetrzak, J. & Jenkins, G. High resolution analysis of meiotic chromosome structure and behaviour in barley (Hordeum vulgare L.). PLoS ONE 7, e39539 (2012).
Wang, C.J., Carlton, P.M., Golubovskaya, I.N. & Cande, W.Z. Interlock formation and coiling of meiotic chromosome axes during synapsis. Genetics 183, 905–915 (2009).
Dürr, J. et al. The transcript elongation factor SPT4/SPT5 is involved in auxin-related gene expression in Arabidopsis. Nucleic Acids Res. 42, 4332–4347 (2014).
Heckmann, S. et al. Alternative meiotic chromatid segregation in the holocentric plant Luzula elegans. Nat. Commun. 5, 4979 (2014).
Heckmann, S. et al. The holocentric species Luzula elegans shows interplay between centromere and large-scale genome organization. Plant J. 73, 555–565 (2013).
Schubert, V. RNA polymerase II forms transcription networks in rye and Arabidopsis nuclei and its amount increases with endopolyploidy. Cytogenet. Genome Res. 143, 69–77 (2014).
Schubert, V., Lermontova, I. & Schubert, I. The Arabidopsis CAP-D proteins are required for correct chromatin organisation, growth and fertility. Chromosoma 122, 517–533 (2013).
Bell, K., Mitchell, S., Paultre, D., Posch, M. & Oparka, K. Correlative imaging of fluorescent proteins in resin-embedded plant material. Plant Physiol. 161, 1595–1603 (2013).
Fitzgibbon, J., Bell, K., King, E. & Oparka, K. Super-resolution imaging of plasmodesmata using three-dimensional structured illumination microscopy. Plant Physiol. 153, 1453–1463 (2010).
Liesche, J., Ziomkiewicz, I. & Schulz, A. Super-resolution imaging with Pontamine Fast Scarlet 4BS enables direct visualization of cellulose orientation and cell connection architecture in onion epidermis cells. BMC Plant Biol. 13, 226 (2013).
Fiolka, R., Shao, L., Rego, E.H., Davidson, M.W. & Gustafsson, M.G. Time-lapse two-color 3D imaging of live cells with doubled resolution using structured illumination. Proc. Natl. Acad. Sci. USA 109, 5311–5315 (2012).
Conduit, P.T. et al. A molecular mechanism of mitotic centrosome assembly in Drosophila. Elife 3, e03399 (2014).
Kasuboski, J.M., Sigal, Y.J., Joens, M.S., Lillemeier, B.F. & Fitzpatrick, J.A. Super-resolution microscopy: a comparative treatment. Curr. Protoc. Cytom. 62, 2.17.1–2.17.24 (2012).
Fornasiero, E.F. & Opazo, F. Super-resolution imaging for cell biologists: concepts, applications, current challenges and developments. Bioessays 37, 436–451 (2015).
Lee, S., Lim, W.A. & Thorn, K.S. Improved blue, green, and red fluorescent protein tagging vectors for S. cerevisiae. PLoS ONE 8, e67902 (2013).
Shao, L., Kner, P., Rego, E.H. & Gustafsson, M.G. Super-resolution 3D microscopy of live whole cells using structured illumination. Nat. Methods 8, 1044–1046 (2011).
Kner, P., Chhun, B.B., Griffis, E.R., Winoto, L. & Gustafsson, M.G. Super-resolution video microscopy of live cells by structured illumination. Nat. Methods 6, 339–342 (2009).
Chai, Y. et al. Live imaging of cellular dynamics during Caenorhabditis elegans postembryonic development. Nat. Protoc. 7, 2090–2102 (2012).
Shaner, N.C. et al. A bright monomeric green fluorescent protein derived from Branchiostoma lanceolatum. Nat. Methods 10, 407–409 (2013).
Griesbeck, O., Baird, G.S., Campbell, R.E., Zacharias, D.A. & Tsien, R.Y. Reducing the environmental sensitivity of yellow fluorescent protein. Mechanism and applications. J. Biol. Chem. 276, 29188–29194 (2001).
Shaner, N.C. et al. Improving the photostability of bright monomeric orange and red fluorescent proteins. Nat. Methods 5, 545–551 (2008).
Littlejohn, G.R., Gouveia, J.D., Edner, C., Smirnoff, N. & Love, J. Perfluorodecalin enhances in vivo confocal microscopy resolution of Arabidopsis thaliana mesophyll. New Phytol. 186, 1018–1025 (2010).
Rego, E.H. et al. Nonlinear structured-illumination microscopy with a photoswitchable protein reveals cellular structures at 50-nm resolution. Proc. Natl. Acad. Sci. USA 109, 135–143 (2012).
Schindelin, J. et al. Fiji: an open-source platform for biological-image analysis. Nat. Methods 9, 676–682 (2012).
Cole, R.W., Jinadasa, T. & Brown, C.M. Measuring and interpreting point spread functions to determine confocal microscope resolution and ensure quality control. Nat. Protoc. 6, 1929–1941 (2011).
Fu, J. & Glover, D.M. Structured illumination of the interface between centriole and peri-centriolar material. Open Biol. 2, 120104 (2012).
Righolt, C.H. et al. Image filtering in structured illumination microscopy using the Lukosz bound. Opt. Express 21, 24431–24451 (2013).
Gustafsson, M.G. Nonlinear structured-illumination microscopy: wide-field fluorescence imaging with theoretically unlimited resolution. Proc. Natl. Acad. Sci. USA 102, 13081–13086 (2005).
Acknowledgements
Development of this protocol was financially supported by grant no. P501/11/1764 from the Czech Science Foundation (GACˇR); by grant no. LO1204 from the Czech Ministry of Education, Youth and Sports, National Program of Sustainability I (NPUI) to the Centre of the Region Haná for Biotechnological and Agricultural Research; and by the Czech National program of sustainability, LO1304.
Author information
Authors and Affiliations
Contributions
G.K., M.M., O.Š. and J.Š. performed the experiments. G.K., O.Š., M.M. and J.Š. established material preparation and microscope settings for SIM. M.M., O.Š., J.B. and J.Š. provided material and equipment. G.K. carried out quantitative analyses and validation procedures inherent to the protocol. M.O. prepared Supplementary Figure 1 with help from O.Š. and G.K. G.K. and J.Š. wrote the manuscript with editorial help from M.O., O.Š., M.M. and J.B. J.Š. and G.K. designed the experiments, and J.Š. supervised the whole project.
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Stepwise assembly of the observation chamber for plants.
(a) Placement and orientation of seedling (arrow throughout a-d) in a drop of liquid ½ MS on top of a glass slide. (b) Placement of coverslip and filling of observation chamber with liquid ½ MS. (c) Sealing of the coverslip rim with silicone paste. (d) The observation chamber with the seedling in proper orientation (arrow) fully sealed.
Supplementary Figure 2 Comparative characterization of lateral resolution of individual and bundled cortical microtubules between SIM and WF acquisitions.
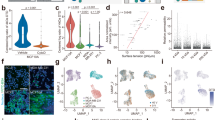
(a) SIM overview of the cortical microtubule array of a hypocotyl cell stably expressing GFP-MBD microtubule marker. (b) The respective WF image corresponding to (a). (c) Transverse 1 μm profile (small diagonal line) across individual microtubule from the left boxed area of (a). (d) Transverse 1 μm profile (small diagonal line) across three closely adjacent microtubules from the right boxed area of (a). (e) Normalized intensity scatterplot corresponding to the individual microtubule profile of (c) by both SIM and WF. The width of the respective curves (green for SIM and red for WF) at 0.5 of normalized intensity corresponds to the FWHM of the individual microtubule. (f) Normalized intensity scatterplots corresponding to the microtubule bundle profile of (d). In both cases SIM (green lines) clearly separates three peaks instead of a single broad one in WF mode (red lines). All scale bars correspond to 5 μm.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1 and 2 (PDF 314 kb)
Time-lapse imaging of cortical microtubule dynamics in a hypocotyl epidermal cell of Arabidopsis thaliana stably transformed with the GFP-MBD microtubule marker.
Video corresponds to Fig. 7a,b. Recording time: 239.4 sec, Frames: 90, Time interval: 2.7 sec, Video frame rate: 16 fps (MOV 24992 kb)
Rights and permissions
About this article
Cite this article
Komis, G., Mistrik, M., Šamajová, O. et al. Superresolution live imaging of plant cells using structured illumination microscopy. Nat Protoc 10, 1248–1263 (2015). https://doi.org/10.1038/nprot.2015.083
Published:
Issue Date:
DOI: https://doi.org/10.1038/nprot.2015.083
This article is cited by
-
Super-resolution microscopy reveals the number and distribution of topoisomerase IIα and CENH3 molecules within barley metaphase chromosomes
Chromosoma (2023)
-
Super-resolution imaging of Douglas fir xylem cell wall nanostructure using SRRF microscopy
Plant Methods (2022)
-
Protein trafficking in plant cells: Tools and markers
Science China Life Sciences (2020)
-
Cytokinin fluoroprobe reveals multiple sites of cytokinin perception at plasma membrane and endoplasmic reticulum
Nature Communications (2020)
-
Multicolour three dimensional structured illumination microscopy of immunolabeled plant microtubules and associated proteins
Plant Methods (2019)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.