Featured
-
-
Article |
 UFM1 E3 ligase promotes recycling of 60S ribosomal subunits from the ER
UFM1 E3 ligase promotes recycling of 60S ribosomal subunits from the ER
Structural and biochemical analyses reveal details of how UFM1 conjugation and deconjugation mediate ribosome recycling and quality control.
- Paul A. DaRosa
- , Ivan Penchev
- & Ron R. Kopito
-
Article
| Open Access The UFM1 E3 ligase recognizes and releases 60S ribosomes from ER translocons
The UFM1 E3 ligase recognizes and releases 60S ribosomes from ER translocons
Attachment of the ubiquitin-like modifier UFM1 to 60S ribosomes has a critical function in the release and recycling of stalled or terminated ribosomes from the endoplasmic reticulum membrane.
- Linda Makhlouf
- , Joshua J. Peter
- & Yogesh Kulathu
-
Article
| Open Access Stress response silencing by an E3 ligase mutated in neurodegeneration
Stress response silencing by an E3 ligase mutated in neurodegeneration
The E3 ligase SIFI is identified as a dedicated silencing factor of the integrated stress response, a finding that has implications for the development of therapeutics for neurodegenerative diseases caused by mitochondrial protein import stress.
- Diane L. Haakonsen
- , Michael Heider
- & Michael Rapé
-
Article
| Open Access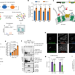 Ubiquitination regulates ER-phagy and remodelling of endoplasmic reticulum
Ubiquitination regulates ER-phagy and remodelling of endoplasmic reticulum
Ubiquitination of the receptor FAM134B regulates ER-phagy and remodelling of the endoplasmic reticulum in response to cellular demands.
- Alexis González
- , Adriana Covarrubias-Pinto
- & Ivan Dikić
-
Article
| Open Access Heteromeric clusters of ubiquitinated ER-shaping proteins drive ER-phagy
Heteromeric clusters of ubiquitinated ER-shaping proteins drive ER-phagy
The membrane-shaping protein ARL6IP1 is involved in the selective degradation of the endoplasmic reticulum, and this process depends on its ubiquitination and interaction with other membrane-shaping proteins such as FAM134B.
- Hector Foronda
- , Yangxue Fu
- & Christian A. Hübner
-
Article
| Open Access Post-translational control of beige fat biogenesis by PRDM16 stabilization
Post-translational control of beige fat biogenesis by PRDM16 stabilization
The ubiquitin E3 ligase CUL2–APPBP2 determines PRDM16 protein stability by catalysing PRDM16 polyubiquitination in beige fat.
- Qiang Wang
- , Huixia Li
- & Shingo Kajimura
-
Article
| Open Access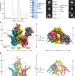 A peroxisomal ubiquitin ligase complex forms a retrotranslocation channel
A peroxisomal ubiquitin ligase complex forms a retrotranslocation channel
The cryo-electron microscopy structure of the membrane-embedded ubiquitin ligase complex reveals its function as a retrotranslocation channel for shuttling mobile receptors out of peroxisomes.
- Peiqiang Feng
- , Xudong Wu
- & Tom A. Rapoport
-
Article |
 Structural insights into Ubr1-mediated N-degron polyubiquitination
Structural insights into Ubr1-mediated N-degron polyubiquitination
Structures of Ubr1 in complex with Ubc2, ubiquitin and two N-degron peptides reveal a Ubc2-binding region and an acceptor ubiquitin-binding loop on Ubr1, providing mechanistic insights into the initiation and elongation steps of ubiquitination catalysed by Ubr1.
- Man Pan
- , Qingyun Zheng
- & Minglei Zhao
-
Article |
 Mechanisms of BRCA1–BARD1 nucleosome recognition and ubiquitylation
Mechanisms of BRCA1–BARD1 nucleosome recognition and ubiquitylation
The authors elucidate the mechanisms for the ubiquitylation specificity and recruitment of the ubiquitin ligase complex BRCA1–BARD1 to damaged DNA within chromatin to facilitate homologous recombination.
- Qi Hu
- , Maria Victoria Botuyan
- & Georges Mer
-
Article |
 Ubiquitylation of lipopolysaccharide by RNF213 during bacterial infection
Ubiquitylation of lipopolysaccharide by RNF213 during bacterial infection
Upon Salmonella invasion of the mammalian cytosol, ubiquitylation of a non-proteinaceous substrate—the lipid A moiety of bacterial lipopolysaccharide—by the E3 ubiquitin ligase RNF213 marks the bacteria as cargo for antibacterial autophagy.
- Elsje G. Otten
- , Emma Werner
- & Felix Randow
-
Article |
 Structural basis for dimerization quality control
Structural basis for dimerization quality control
Structural studies of the dimerization quality control E3 ubiquitin ligase SCF–FBXL17 indicate that its selectivity for aberrant complex formation is based on recognizing both shape and complementarity of interacting domains.
- Elijah L. Mena
- , Predrag Jevtić
- & Michael Rape
-
Article |
 Envelope protein ubiquitination drives entry and pathogenesis of Zika virus
Envelope protein ubiquitination drives entry and pathogenesis of Zika virus
The E3 ubiquitin ligase TRIM7 polyubiquitinates the envelope protein of Zika virus, adding Lys63-linked polyubiquitin chains that interact with the TIM1 receptor of host cells to enhance virus entry and replication.
- Maria I. Giraldo
- , Hongjie Xia
- & Ricardo Rajsbaum
-
Article |
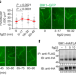 Ligand-induced monoubiquitination of BIK1 regulates plant immunity
Ligand-induced monoubiquitination of BIK1 regulates plant immunity
The detection of microorganism-associated ligands by plant cells activates a signalling cascade in which the kinase BIK1 is monoubiquinated, released from the FLS2–BAK1 complex, and internalized by endocytosis.
- Xiyu Ma
- , Lucas A. N. Claus
- & Libo Shan
-
Article |
 Gene expression and cell identity controlled by anaphase-promoting complex
Gene expression and cell identity controlled by anaphase-promoting complex
WDR5 and TBP recruit anaphase-promoting complex to specific transcription start sites in mitosis, initiating a ubiquitin-dependent mechanism that preserves cell identity by linking gene expression and cell division.
- Eugene Oh
- , Kevin G. Mark
- & Michael Rape
-
Article |
 Stress- and ubiquitylation-dependent phase separation of the proteasome
Stress- and ubiquitylation-dependent phase separation of the proteasome
Hyperosmotic stress leads to a phase separation of the proteasome, triggered by interactions between RAD23B and ubiquitylated proteins, which bring together p97 and proteasome-associated proteins into nuclear proteolytic foci.
- Sayaka Yasuda
- , Hikaru Tsuchiya
- & Yasushi Saeki
-
Article |
 Host-mediated ubiquitination of a mycobacterial protein suppresses immunity
Host-mediated ubiquitination of a mycobacterial protein suppresses immunity
Mycobacterium tuberculosis suppresses the production of inflammatory cytokines by host cells through the host-mediated ubiquitination of a mycobacterial protein, enhancing the interaction of a host signalling inhibitor with another signalling molecule.
- Lin Wang
- , Juehui Wu
- & Baoxue Ge
-
Article |
 Structure of the Fanconi anaemia monoubiquitin ligase complex
Structure of the Fanconi anaemia monoubiquitin ligase complex
The structure of the multiprotein Fanconi anaemia core complex, determined using cryo-electron microscopy and mass spectrometry, shows that the complex adopts an extended asymmetric structure and highlights the structural and functional asymmetry of the RING finger domains.
- Shabih Shakeel
- , Eeson Rajendra
- & Lori A. Passmore
-
Letter |
 Insights into ubiquitin chain architecture using Ub-clipping
Insights into ubiquitin chain architecture using Ub-clipping
Enzymatic cleavage within ubiquitin molecules followed by quantitative mass-spectrometry simplifies complex ubiquitin chains and enables mapping of polyubiquitin architectures.
- Kirby N. Swatek
- , Joanne L. Usher
- & David Komander
-
Letter |
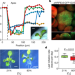 An apical hypoxic niche sets the pace of shoot meristem activity
An apical hypoxic niche sets the pace of shoot meristem activity
Hypoxia in the shoot meristem of Arabidopsis links the regulation of metabolic activity to development by inhibiting proteolysis of a substrate of the N-degron pathway, which controls class-III homeodomain-leucine zipper transcription factors.
- Daan A. Weits
- , Alicja B. Kunkowska
- & Francesco Licausi
-
Letter |
 Distinct proteostasis circuits cooperate in nuclear and cytoplasmic protein quality control
Distinct proteostasis circuits cooperate in nuclear and cytoplasmic protein quality control
Ubiquitin chains linked to cytoplasmic misfolded proteins are different from those linked to nuclear misfolded proteins, each requiring a distinct combination of molecular chaperones and ubiquitination circuitries.
- Rahul S. Samant
- , Christine M. Livingston
- & Judith Frydman
-
Letter |
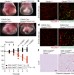 OTULIN limits cell death and inflammation by deubiquitinating LUBAC
OTULIN limits cell death and inflammation by deubiquitinating LUBAC
OTULIN, which removes ubiquitin chains deposited by LUBAC, promotes LUBAC activity by preventing its auto-ubiquitination, thereby supporting normal mouse embryo development and preventing pro-inflammatory cell death in adult mice.
- Klaus Heger
- , Katherine E. Wickliffe
- & Vishva M. Dixit
-
Letter |
 Mechanism of parkin activation by PINK1
Mechanism of parkin activation by PINK1
Structural mass spectrometry of full-length human parkin and a structure of the activated parkin core reveal large-scale domain rearrangements involved in activation of parkin by PINK1.
- Christina Gladkova
- , Sarah L. Maslen
- & David Komander
-
Letter |
 Mechanism of phosphoribosyl-ubiquitination mediated by a single Legionella effector
Mechanism of phosphoribosyl-ubiquitination mediated by a single Legionella effector
Crystal structures of the Legionella effectors SdeA and SdeD uncover the mechanism of a unique phosphoribosyl-ubiquitination reaction.
- Anil Akturk
- , David J. Wasilko
- & Yuxin Mao
-
Letter |
 Insights into catalysis and function of phosphoribosyl-linked serine ubiquitination
Insights into catalysis and function of phosphoribosyl-linked serine ubiquitination
Structural and functional investigations demonstrate how bacterial enzymes ubiquitinate host proteins.
- Sissy Kalayil
- , Sagar Bhogaraju
- & Ivan Dikic
-
Article |
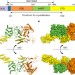 Structural basis of ubiquitin modification by the Legionella effector SdeA
Structural basis of ubiquitin modification by the Legionella effector SdeA
Crystal structures of the Legionella effector SdeA in a ligand-free state and in complex with ubiquitin and NADH provide insight into SdeA-mediated phosphoribosyl-linked ubiquitination.
- Yanan Dong
- , Yajuan Mu
- & Yue Feng
-
Letter |
 LUBAC is essential for embryogenesis by preventing cell death and enabling haematopoiesis
LUBAC is essential for embryogenesis by preventing cell death and enabling haematopoiesis
The HOIL-1 component of the LUBAC ubiquitin ligase complex is required for LUBAC activity, which prevents lethality during embryogenesis by preventing aberrant TNFR1-mediated endothelial cell death and RIPK1-mediated defects in haematopoiesis.
- Nieves Peltzer
- , Maurice Darding
- & Henning Walczak
-
Letter |
 Activity-based E3 ligase profiling uncovers an E3 ligase with esterification activity
Activity-based E3 ligase profiling uncovers an E3 ligase with esterification activity
Non-lysine ubiquitination activity of the E3 ubiquitin ligase MYCBP2 is identified by activity-based profiling; biochemical and structural analysis of MYCBP2 suggests the basis for its mechanism and specificity.
- Kuan-Chuan Pao
- , Nicola T. Wood
- & Satpal Virdee
-
Letter |
 Cyclin D–CDK4 kinase destabilizes PD-L1 via cullin 3–SPOP to control cancer immune surveillance
Cyclin D–CDK4 kinase destabilizes PD-L1 via cullin 3–SPOP to control cancer immune surveillance
Abundance of PD-L1, the ligand of the anti-cancer immunotherapy target PD-1, is negatively regulated by poly-ubiquitination via the cyclin D–CDK4/cullin 3–SPOP axis and PD-1 blockade treatment in mice improved survival when combined with CDK4/6 inhibitors.
- Jinfang Zhang
- , Xia Bu
- & Wenyi Wei
-
Letter |
 A ubiquitin-dependent signalling axis specific for ALKBH-mediated DNA dealkylation repair
A ubiquitin-dependent signalling axis specific for ALKBH-mediated DNA dealkylation repair
A signalling mechanism in human cells for sensing DNA damage induced by alkylation involves ubiquitin-dependent recruitment of the alkylation repair complex ASCC to the vicinity of the damage and co-localization with transcription and splicing factors.
- Joshua R. Brickner
- , Jennifer M. Soll
- & Nima Mosammaparast
-
Article |
 Structure of PINK1 in complex with its substrate ubiquitin
Structure of PINK1 in complex with its substrate ubiquitin
Stabilization of a transient protein kinase–substrate complex using a nanobody provides molecular details about how the Parkinson’s disease-linked protein kinase PINK1 phosphorylates ubiquitin, and suggests new pharmacological strategies.
- Alexander F. Schubert
- , Christina Gladkova
- & David Komander
-
Article |
 Molecular basis of USP7 inhibition by selective small-molecule inhibitors
Molecular basis of USP7 inhibition by selective small-molecule inhibitors
Small molecules are identified that inhibit the ubiquitin-specific protease USP7 with high affinity and specificity as explained by co-crystal structures, and are shown to reduce tumour growth in mice.
- Andrew P. Turnbull
- , Stephanos Ioannidis
- & David Komander
-
Letter |
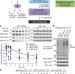 USP7 small-molecule inhibitors interfere with ubiquitin binding
USP7 small-molecule inhibitors interfere with ubiquitin binding
The development of selective ubiquitin-specific protease-7 (USP7) inhibitors GNE-6640 and GNE-6776, which induce tumour cell death and reveal differential kinetics of Lys-48 and Lys-63-linked ubiquitin chain depolymerization by USP7.
- Lorna Kategaya
- , Paola Di Lello
- & Ingrid E. Wertz
-
Letter |
 PTEN counteracts FBXL2 to promote IP3R3- and Ca2+-mediated apoptosis limiting tumour growth
PTEN counteracts FBXL2 to promote IP3R3- and Ca2+-mediated apoptosis limiting tumour growth
PTEN, a known tumour suppressor, inhibits the FXBL2-dependent degradation of IP3R3, an IP3 receptor, thus augmenting IP3R3-mediated calcium release from the endoplasmic reticulum to mitochondria and inducing apoptosis; inhibiting FXBL2 sensitizes PTEN-deficient tumours to photodynamic therapy.
- Shafi Kuchay
- , Carlotta Giorgi
- & Michele Pagano
-
Letter |
 Molecular basis of Lys11-polyubiquitin specificity in the deubiquitinase Cezanne
Molecular basis of Lys11-polyubiquitin specificity in the deubiquitinase Cezanne
The structures of the deubiquitinating enzyme Cezanne alone or in complex with its substrate or product are solved, showing how Cezanne specifically targets Lys11-linked polyubiquitin.
- Tycho E. T. Mevissen
- , Yogesh Kulathu
- & David Komander
-
Article |
 A novel cereblon modulator recruits GSPT1 to the CRL4CRBN ubiquitin ligase
A novel cereblon modulator recruits GSPT1 to the CRL4CRBN ubiquitin ligase
This paper reports the identification of a new cereblon-modulating agent, CC-885, which targets the translation termination factor GSPT1 and demonstrates anti-tumour activity in patient-derived tumour cells; the crystal structure of the cereblon–DDB1–GSPT1–CC-885 complex reveals a common motif for cereblon-substrate recruitment.
- Mary E. Matyskiela
- , Gang Lu
- & Philip P. Chamberlain
-
Article |
 Cullin–RING ubiquitin E3 ligase regulation by the COP9 signalosome
Cullin–RING ubiquitin E3 ligase regulation by the COP9 signalosome
Much of the intracellular protein degradation in eukaryotes is controlled by cullin–RING ubiquitin ligases (CRLs), a vast class of enzymes which are regulated by the COP9 signalosome (CSN); structural characterization of CSN–N8CRL4A complexes by cryo-electron microscopy reveals an induced-fit mechanism of CSN activation triggered only by catalytically activated CRLs without bound substrate, explaining how CSN acts as a global regulator of CRLs.
- Simone Cavadini
- , Eric S. Fischer
- & Nicolas H. Thomä
-
Letter |
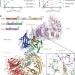 Structural basis of lenalidomide-induced CK1α degradation by the CRL4CRBN ubiquitin ligase
Structural basis of lenalidomide-induced CK1α degradation by the CRL4CRBN ubiquitin ligase
Thalidomide and its derivative lenalidomide bind the CRL4CRBN E3 ubiquitin ligase and target protein substrates for degradation; structural and functional data determined here show that casein kinase 1α and the lymphoid transcription factor Ikaros, the efficacy targets of lenalidomide in two different blood cancers, interact with the CRBN–lenalidomide interface through a β-hairpin destruction motif.
- Georg Petzold
- , Eric S. Fischer
- & Nicolas H. Thomä
-
Letter |
 Structure of a HOIP/E2~ubiquitin complex reveals RBR E3 ligase mechanism and regulation
Structure of a HOIP/E2~ubiquitin complex reveals RBR E3 ligase mechanism and regulation
The first structure of fully active HOIP of the RBR family of RING-type E3 ligases in its transfer complex with an E2~ubiquitin conjugate provides insights into its mechanism of action, including the ideal alignment of the E2 and E3 catalytic centres for ubiquitin transfer and the allosteric regulation of the RBR family.
- Bernhard C. Lechtenberg
- , Akhil Rajput
- & Stefan J. Riedl
-
Letter |
 Histone H1 couples initiation and amplification of ubiquitin signalling after DNA damage
Histone H1 couples initiation and amplification of ubiquitin signalling after DNA damage
At the initiation of DNA double-strand break repair, a number of ubiquitylation events occur; here, the RNF8 ubiquitin E3 ligase and the ubiquitin-conjugating E2 enzyme, UBC13, are shown to primarily modify H1-type linker histones, via a K63 linkage.
- Tina Thorslund
- , Anita Ripplinger
- & Niels Mailand
-
Letter |
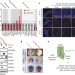 Cell-fate determination by ubiquitin-dependent regulation of translation
Cell-fate determination by ubiquitin-dependent regulation of translation
This study shows that a vertebrate-specific ubiquitin ligase modulates neural crest specification in Xenopus development and human embryonic stem-cell differentiation; a proteomics approach reveals that the CUL3KBTBD8 ligase modulates translation by targeting the modulators of ribosomes production NOLC1 and its paralogue TCOF1, which is mutated in a neural-crest-associated syndrome.
- Achim Werner
- , Shintaro Iwasaki
- & Michael Rape
-
Article |
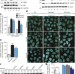 The ubiquitin kinase PINK1 recruits autophagy receptors to induce mitophagy
The ubiquitin kinase PINK1 recruits autophagy receptors to induce mitophagy
The PINK1 ubiquitin kinase is shown to recruit the two autophagy receptors NDP52 and OPTN to mitochondria to activate mitophagy directly, independently of the ubiquitin ligase parkin; once recruited to mitochondria, NDP52 and OPTN recruit autophagy initiation components, and parkin may amplify the phospho-ubiquitin signal generated by PINK1, resulting in robust autophagy induction.
- Michael Lazarou
- , Danielle A. Sliter
- & Richard J. Youle
-
Letter |
 Mechanism of phospho-ubiquitin-induced PARKIN activation
Mechanism of phospho-ubiquitin-induced PARKIN activation
This study provides insights into conformational changes that lead to phospho-ubiquitin-induced PARKIN activation and how PARKIN is recruited to phospho-ubiquitin chains on mitochondria; the crystal structure of PARKIN in complex with phospho-ubiquitin also indicates that the pocket within PARKIN where phospho-ubiquitin binds carries amino acid residues that are mutated in patients with autosomal-recessive juvenile Parkinsonism.
- Tobias Wauer
- , Michal Simicek
- & David Komander
-
Article |
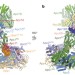 Atomic structure of the APC/C and its mechanism of protein ubiquitination
Atomic structure of the APC/C and its mechanism of protein ubiquitination
A cryo-electron microscopy determination of the atomic structures of anaphase-promoting complex (APC/C)–coactivator complexes with either Emi1 or a UbcH10–ubiquitin conjugate.
- Leifu Chang
- , Ziguo Zhang
- & David Barford
-
Letter |
 Allosteric activation of the RNF146 ubiquitin ligase by a poly(ADP-ribosyl)ation signal
Allosteric activation of the RNF146 ubiquitin ligase by a poly(ADP-ribosyl)ation signal
Structural and biochemical approaches are used to show how RNF146 activity is allosterically regulated by the binding of poly(ADP-ribose) ligand, and how substrate specificity is achieved with protein poly(ADP-ribosyl)ation and ubiquitination occurring in the same protein complex.
- Paul A. DaRosa
- , Zhizhi Wang
- & Wenqing Xu
-
Letter |
 Structural basis for ubiquitin-mediated antiviral signal activation by RIG-I
Structural basis for ubiquitin-mediated antiviral signal activation by RIG-I
RIG-I protein recognizes viral duplex RNA with a 5′-triphosphate group, activating innate immune responses; a crystal structure of its tetrameric CARD signalling domain reveals that non-covalently linked ubiquitin chains stabilize the tetramer in a ‘lock-washer’ structure that serves as a signalling platform for the recruitment and activation of MAVS.
- Alys Peisley
- , Bin Wu
- & Sun Hur
-
Article |
 53BP1 is a reader of the DNA-damage-induced H2A Lys 15 ubiquitin mark
53BP1 is a reader of the DNA-damage-induced H2A Lys 15 ubiquitin mark
This study shows that 53BP1 recruitment to sites of DNA damage involves dual recognition of H4K20me2 and H2AK15 histone ubiquitination; the ubiquitin mark and the surrounding epitope on H2A are read by a region of 53BP1 designated the ubiquitination-dependent recruitment motif.
- Amélie Fradet-Turcotte
- , Marella D. Canny
- & Daniel Durocher
-
Letter |
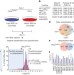 Landscape of the PARKIN-dependent ubiquitylome in response to mitochondrial depolarization
Landscape of the PARKIN-dependent ubiquitylome in response to mitochondrial depolarization
PARKIN, a protein involved in mitochondria clearance by autophagy, is often mutated in early-onset familial Parkinson’s disease; here the cellular repertoire of PARKIN targets is identified by quantitative proteomics.
- Shireen A. Sarraf
- , Malavika Raman
- & J. Wade Harper
-
Letter |
 BTB-ZF factors recruit the E3 ligase cullin 3 to regulate lymphoid effector programs
BTB-ZF factors recruit the E3 ligase cullin 3 to regulate lymphoid effector programs
The E3 ubiquitin ligase cullin 3 is shown to bind BTB-zinc finger transcription factors to direct the ubiquitination of nuclear chromatin-associated factors to control transcription and cell-fate decisions in B- and T-cell populations.
- Rebecca Mathew
- , Michael P. Seiler
- & Albert Bendelac
-
Article |
 Ubiquitin-dependent regulation of COPII coat size and function
Ubiquitin-dependent regulation of COPII coat size and function
The size of COPII vesicles is shown to be controlled by monoubiquitylation, with potential implications for cranio-lenticulo-sutural dysplasia and chylomicron retention disease.
- Lingyan Jin
- , Kanika Bajaj Pahuja
- & Michael Rape
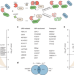
 Proteome-scale discovery of protein degradation and stabilization effectors
Proteome-scale discovery of protein degradation and stabilization effectors