Featured
-
-
Article
| Open Access Functional screening in human HSPCs identifies optimized protein-based enhancers of Homology Directed Repair
Functional screening in human HSPCs identifies optimized protein-based enhancers of Homology Directed Repair
Here the authors describe a functional screening platform in human stem cells to identify and optimize protein-based gene editing additives that increase homologous directed recombination and have potential to improve gene therapy workflows.
- Juan A. Perez-Bermejo
- , Oghene Efagene
- & Kristen L. Seim
-
Article
| Open Access Enhancing prime editor activity by directed protein evolution in yeast
Enhancing prime editor activity by directed protein evolution in yeast
Compared to traditional Cas9 nucleases prime editors (PEs) are less active. Here the authors use OrthoRep, a yeast-based platform for directed protein evolution to enhance the editing efficiency of PEs: they identify mutations that have a positive effect on kinetics and use this knowledge to generate an efficient in vivo PE.
- Yanik Weber
- , Desirée Böck
- & Gerald Schwank
-
Article
| Open Access Phage-assisted evolution of highly active cytosine base editors with enhanced selectivity and minimal sequence context preference
Phage-assisted evolution of highly active cytosine base editors with enhanced selectivity and minimal sequence context preference
Existing TadA-derived CBEs exhibit residual A•T-to-G•C editing activity and suffer from lower activity at several sequence contexts and with non-SpCas9 targeting domains. Here, the authors use phage-assisted evolution to evolve CBE6 variants that address these limitations.
- Emily Zhang
- , Monica E. Neugebauer
- & David R. Liu
-
Article
| Open Access Domain-inlaid Nme2Cas9 adenine base editors with improved activity and targeting scope
Domain-inlaid Nme2Cas9 adenine base editors with improved activity and targeting scope
Nme2Cas9 has been well established as a genome editing platform. Here the authors engineer Nme2Cas9 to further increase the activity and targeting scope of compact Nme2Cas9 base editors and validate domain-inlaid Nme2-ABEs for single-AAV delivery in vivo.
- Nathan Bamidele
- , Han Zhang
- & Erik J. Sontheimer
-
Article
| Open Access Compact zinc finger architecture utilizing toxin-derived cytidine deaminases for highly efficient base editing in human cells
Compact zinc finger architecture utilizing toxin-derived cytidine deaminases for highly efficient base editing in human cells
The most recent class of base editors utilize DddAtox, a deaminase domain that can act upon double-stranded DNA. Here the authors target DddAtox fragments and a FokI-based nickase to the human CIITA gene by fusing these domains to arrays of engineered zinc fingers; they also identify a variety of DddAtox orthologues.
- Friedrich Fauser
- , Bhakti N. Kadam
- & Jeffrey C. Miller
-
Article
| Open Access Transient inhibition of 53BP1 increases the frequency of targeted integration in human hematopoietic stem and progenitor cells
Transient inhibition of 53BP1 increases the frequency of targeted integration in human hematopoietic stem and progenitor cells
Here the authors demonstrate that the frequency of HDR in human hematopoietic stem and progenitor cells is increased by the delivery of an inhibitor of 53BP1 as a recombinant peptide. This approach is applicable for a variety of therapeutically relevant loci in HSPCs as well in other primary human cell types.
- Ron Baik
- , M. Kyle Cromer
- & Matthew H. Porteus
-
Article
| Open Access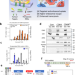 Dual targeted extracellular vesicles regulate oncogenic genes in advanced pancreatic cancer
Dual targeted extracellular vesicles regulate oncogenic genes in advanced pancreatic cancer
KRASG12D mutations frequently co-occur with mutated TP53 tumour suppressor in patients with pancreatic ductal adenocarcinoma (PDAC). Here the authors report the design of dual targeted therapeutic extracellular vesicles containing high copy numbers of TP53 mRNA and siKRASG12D, showing anti-tumor activity in PDAC preclinical models.
- Chi-Ling Chiang
- , Yifan Ma
- & L. James Lee
-
Article
| Open Access PAM-flexible genome editing with an engineered chimeric Cas9
PAM-flexible genome editing with an engineered chimeric Cas9
CRISPR enzymes require a defined protospacer adjacent motif (PAM) which can be limiting for editing applications. Here the authors recombine the PAM-interacting domain of SpRY with the N-terminus of Sc + + to generate a chimeric enzyme with highly flexible PAM preference: SpRYc.
- Lin Zhao
- , Sabrina R. T. Koseki
- & Pranam Chatterjee
-
Article
| Open Access AAV-mediated base-editing therapy ameliorates the disease phenotypes in a mouse model of retinitis pigmentosa
AAV-mediated base-editing therapy ameliorates the disease phenotypes in a mouse model of retinitis pigmentosa
Base editing technology has great potential in treating pathogenic single-nucleotide variations. Using a dual-AAV base editing system, Wu et al. restored visual functions in a mouse model of retinitis pigmentosa.
- Yidong Wu
- , Xiaoling Wan
- & Xueli Zhang
-
Article
| Open Access Cell cycle arrest and p53 prevent ON-target megabase-scale rearrangements induced by CRISPR-Cas9
Cell cycle arrest and p53 prevent ON-target megabase-scale rearrangements induced by CRISPR-Cas9
ON-target genotoxicity in gene editing is generally underestimated. Here the authors report Fluorescence-Assisted Megabase-scale Rearrangements Detection (FAMReD) systems to detect and characterize rare large loss of heterozygosity: they show that ON-target genotoxicity can be prevented by p53 and cell cycle arrest.
- G. Cullot
- , J. Boutin
- & A. Bedel
-
Article
| Open Access Rapid and definitive treatment of phenylketonuria in variant-humanized mice with corrective editing
Rapid and definitive treatment of phenylketonuria in variant-humanized mice with corrective editing
The PAH P281L variant is one of the most common variants identified in phenylketonuria (PKU) patients. Here, the authors use base editing, enabled by lipid nanoparticle/mRNA technology, to directly correct the P281L variant in the liver in PKU mice and definitively treat the disease within 2 days.
- Dominique L. Brooks
- , Manuel J. Carrasco
- & Xiao Wang
-
Article
| Open Access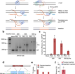 Template-jumping prime editing enables large insertion and exon rewriting in vivo
Template-jumping prime editing enables large insertion and exon rewriting in vivo
Retrotransposons replicate their genetic information through target-primed reverse transcription (TPRT). Here the authors report a template-jumping prime editor (TJ-PE) to act similarly to TPRT and achieve insertions of large DNA fragments at endogenous sites: they rewrite a mutated exon in the mouse liver.
- Chunwei Zheng
- , Bin Liu
- & Wen Xue
-
Article
| Open Access Engineered CRISPR-OsCas12f1 and RhCas12f1 with robust activities and expanded target range for genome editing
Engineered CRISPR-OsCas12f1 and RhCas12f1 with robust activities and expanded target range for genome editing
Cas12f proteins are small and sought after for therapeutic applications. Here the authors report six bacterial Cas12f1 proteins with nuclease activity in mammalian cells, perform sgRNA and protein engineering to generate variants with enhanced editing and broader PAMs, apply an inducible version in vivo.
- Xiangfeng Kong
- , Hainan Zhang
- & Hui Yang
-
Article
| Open Access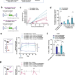 RNAi-mediated rheostat for dynamic control of AAV-delivered transgenes
RNAi-mediated rheostat for dynamic control of AAV-delivered transgenes
Here the authors propose an RNA interference-based switch for dynamic control of AAV transgene expression. In this approach, transgene expression may be silenced by RNAi and subsequently recovered using REVERSIR oligonucleotides.
- Megha Subramanian
- , James McIninch
- & Vasant Jadhav
-
Article
| Open Access Limitations of gene editing assessments in human preimplantation embryos
Limitations of gene editing assessments in human preimplantation embryos
DNA repair in response to DSBs in the preimplantation embryo is hard to analyze. Here the authors show that over 25% of pre-existing heterozygous loci in control single blastomere samples appeared as homozygous after whole genome amplification, therefore, they validated gene editing seen in human embryos in ESCs.
- Dan Liang
- , Aleksei Mikhalchenko
- & Shoukhrat Mitalipov
-
Review Article
| Open Access Assessing and advancing the safety of CRISPR-Cas tools: from DNA to RNA editing
Assessing and advancing the safety of CRISPR-Cas tools: from DNA to RNA editing
CRISPR-Cas tools have shown exceptional promise in genome engineering over the past decade. Here the authors review the development of CRISPR-Cas9/Cas12/Cas13 nucleases, DNA base editors, prime editors, and RNA base editors, as well as their editing precision, off-target effects, and clinical considerations.
- Jianli Tao
- , Daniel E. Bauer
- & Roberto Chiarle
-
Article
| Open Access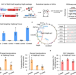 Safeguarding genome integrity during gene-editing therapy in a mouse model of age-related macular degeneration
Safeguarding genome integrity during gene-editing therapy in a mouse model of age-related macular degeneration
Undesired chromosomal translocations, vector integrations, and large deletions remain a problem for therapeutic gene editing in vivo. Here, the authors compare the CRISPR-Cas9TX variant with CRISPR-Cas9 and show elimination of chromosomal translocations and reduction of AVV integration when targeting Vegfa for the treatment of age-related macular degeneration in a mouse model.
- Jianhang Yin
- , Kailun Fang
- & Jiazhi Hu
-
Article
| Open Access Automated high-throughput genome editing platform with an AI learning in situ prediction model
Automated high-throughput genome editing platform with an AI learning in situ prediction model
A large number of cell disease models with pathogenic SNVs are needed. Here the authors report an automated high-throughput platform to perform the genome editing process from gRNA design to the analysis of the editing results; they characterise in situ base editing outcomes.
- Siwei Li
- , Jingjing An
- & Meng Wang
-
Article
| Open Access Low-dose AAV-CRISPR-mediated liver-specific knock-in restored hemostasis in neonatal hemophilia B mice with subtle antibody response
Low-dose AAV-CRISPR-mediated liver-specific knock-in restored hemostasis in neonatal hemophilia B mice with subtle antibody response
Here, the authors develop an AAV-CRISPR mediated somatic knock-in. They apply this system to restore hemostasis in neonatal hemophilia B mice and show liver-specificity of the knock-in and low serum antibody production.
- Xiangjun He
- , Zhenjie Zhang
- & Bo Feng
-
Article
| Open Access Compact zinc finger base editors that edit mitochondrial or nuclear DNA in vitro and in vivo
Compact zinc finger base editors that edit mitochondrial or nuclear DNA in vitro and in vivo
Zinc finger (ZF) arrays are programmable DNA-binding proteins. Here the authors report ZF-DddA-derived cytosine base editors (DdCBEs) and optimise their architectures to improve targeting; they apply these variants in vitro and in vivo to mitochondrial base editing and show higher editing than ZF deaminases.
- Julian C. W. Willis
- , Pedro Silva-Pinheiro
- & David R. Liu
-
Article
| Open Access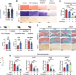 Cholesterol-induced LRP3 downregulation promotes cartilage degeneration in osteoarthritis by targeting Syndecan-4
Cholesterol-induced LRP3 downregulation promotes cartilage degeneration in osteoarthritis by targeting Syndecan-4
This study demonstrates a role of cholesterol metabolism-related gene, Lrp3, in cartilage degeneration and osteoarthritis pathogenesis. LRP3 positively regulates cartilage extracellular matrix metabolism by targeting syndecan-4 via Ras signalling, implicating the cholesterol-LRP3-SDC4 axis in osteoarthritic cartilage degeneration.
- Chenxi Cao
- , Yuanyuan Shi
- & Yingfang Ao
-
Article
| Open Access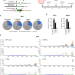 Frequency and mechanisms of LINE-1 retrotransposon insertions at CRISPR/Cas9 sites
Frequency and mechanisms of LINE-1 retrotransposon insertions at CRISPR/Cas9 sites
The identification of events occurring at target sites is critical to determine the safety of CRISPRbased DNA editing tools. Here, the authors show that LINE-1 retrotranspositions can occur frequently at canonical CRISPR/Cas9 editing sites, but are rare with prime editors and base editors.
- Jianli Tao
- , Qi Wang
- & Roberto Chiarle
-
Article
| Open Access Establishment of mouse model of inherited PIGO deficiency and therapeutic potential of AAV-based gene therapy
Establishment of mouse model of inherited PIGO deficiency and therapeutic potential of AAV-based gene therapy
Inherited GPI deficiency (IGD) is caused by PIGO mutations. Here, the authors generate a mouse model of IGD and show that AAV-mediate gene therapy, for knock-in as well as extra-chromosomal expression of Pigo cDNA, ameliorates pathology in the mice.
- Ryoko Kuwayama
- , Keiichiro Suzuki
- & Yoshiko Murakami
-
Article
| Open Access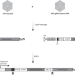 Therapeutic homology-independent targeted integration in retina and liver
Therapeutic homology-independent targeted integration in retina and liver
Limits of AAV-mediated gene therapy include targeting dominant mutations and inducing long-term transgene expression. Here, the authors show that AAV-HITI results in efficient allele-independent integration of a donor DNA in both retina and liver providing therapeutic benefit in mouse models of either a genetic form of blindness or a lysosomal storage disease, respectively.
- Patrizia Tornabene
- , Rita Ferla
- & Alberto Auricchio
-
Article
| Open Access FrCas9 is a CRISPR/Cas9 system with high editing efficiency and fidelity
FrCas9 is a CRISPR/Cas9 system with high editing efficiency and fidelity
Gene editing tools have tremendous potential for biomedical and basic research. Here the authors report a Cas9 from Faecalibaculum rodentium (FrCas9) that achieves efficient and specific gene editing in human cells with a NNTA palindrome PAMs for targeting optimal sites at TATA-boxes to enhance CRISPRa/i screening.
- Zifeng Cui
- , Rui Tian
- & Zheng Hu
-
Article
| Open Access Crosstalk between CRISPR-Cas9 and the human transcriptome
Crosstalk between CRISPR-Cas9 and the human transcriptome
The off-target effects of CRISPR-Cas9 are thought to be mediated by its cognate guide RNA. Here the authors show that Cas9 independently interacts with the human transcriptome, correlating with elevated RNA editing even under guide RNA co-expression.
- Aaron A. Smargon
- , Assael A. Madrigal
- & Gene W. Yeo
-
Article
| Open Access Mutation-specific reporter for optimization and enrichment of prime editing
Mutation-specific reporter for optimization and enrichment of prime editing
While prime editing is a promising technique, some genomic sites remain difficult to edit. Here the authors present fluoPEER, fluorescent prime editing and enrichment reporter, to rank the efficiency of pegRNAs and prime editor variants.
- I. F. Schene
- , I. P. Joore
- & S. A. Fuchs
-
Article
| Open Access Correction of a Factor VIII genomic inversion with designer-recombinases
Correction of a Factor VIII genomic inversion with designer-recombinases
Correction of disease-causing large genomic inversions remains challenging. Here, the authors developed a dual designer-recombinase system (RecF8) that efficiently corrects a 140 kb inversion frequently found in patients with severe Hemophilia A.
- Felix Lansing
- , Liliya Mukhametzyanova
- & Frank Buchholz
-
Article
| Open Access Low immunogenicity of LNP allows repeated administrations of CRISPR-Cas9 mRNA into skeletal muscle in mice
Low immunogenicity of LNP allows repeated administrations of CRISPR-Cas9 mRNA into skeletal muscle in mice
In vivo delivery of CRISPR-Cas9 holds promise for treating muscular dystrophy, however, AAV delivery is known to be immunogenic. Here, the authors show that LNP delivery of CRISPR-Cas9 enables repeated injections into skeletal muscle and leads to restored dystrophin expression in multiple muscle groups.
- Eriya Kenjo
- , Hiroyuki Hozumi
- & Akitsu Hotta
-
Article
| Open Access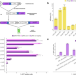 Find and cut-and-transfer (FiCAT) mammalian genome engineering
Find and cut-and-transfer (FiCAT) mammalian genome engineering
Mammalian genome engineering has advanced tremendously over the last decade, however there is still a need for robust gene writing with size scaling capacity. Here the authors present Find Cut-and-Transfer (FiCAT) technology to delivery large targeted payload insertion in cell lines and in vivo in mouse models.
- Maria Pallarès-Masmitjà
- , Dimitrije Ivančić
- & Marc Güell
-
Article
| Open Access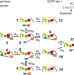 A general theoretical framework to design base editors with reduced bystander effects
A general theoretical framework to design base editors with reduced bystander effects
Base editors can edit target nucleotides, and identical ones that are within the editing window. Here the authors build an analytical model to propose general principles of editor design to reduce bystander effects.
- Qian Wang
- , Jie Yang
- & Anatoly B. Kolomeisky
-
Article
| Open Access Self-inactivating, all-in-one AAV vectors for precision Cas9 genome editing via homology-directed repair in vivo
Self-inactivating, all-in-one AAV vectors for precision Cas9 genome editing via homology-directed repair in vivo
Long-term expression of Cas9 following precision genome editing in vivo may lead to undesirable consequences. Here we show that a single-vector, self-inactivating AAV system containing Cas9 nuclease, guide, and DNA donor can use homology-directed repair to correct disease mutations in vivo.
- Raed Ibraheim
- , Phillip W. L. Tai
- & Erik J. Sontheimer
-
Article
| Open Access CRISPR-Cas9 globin editing can induce megabase-scale copy-neutral losses of heterozygosity in hematopoietic cells
CRISPR-Cas9 globin editing can induce megabase-scale copy-neutral losses of heterozygosity in hematopoietic cells
CRISPR-Cas9 generates double-strand breaks, which pose a threat to genome integrity. Here the authors show loss of heterozygosity in edited hematopoietic progenitor cells.
- J. Boutin
- , J. Rosier
- & A. Bedel
-
Article
| Open Access AsCas12a ultra nuclease facilitates the rapid generation of therapeutic cell medicines
AsCas12a ultra nuclease facilitates the rapid generation of therapeutic cell medicines
The utility of AsCas12a can be limited to poor editing efficiency. Here the authors identify a variant, “AsCas12a Ultra”, that has high on-target specificity demonstrated through editing of clinically relevant T cell genes.
- Liyang Zhang
- , John A. Zuris
- & Christopher A. Vakulskas
-
Article
| Open Access Efficient precise in vivo base editing in adult dystrophic mice
Efficient precise in vivo base editing in adult dystrophic mice
Base editing is one approach used to correct mutations causing cause Duchenne muscular dystrophy (DMD), but limitations are in the requirement for a specific PAM motif and the large size beyond the packaging capacity of adeno-associated virus (AAV). Here, the authors modify the NG-targeting adenine base editor to recognize a broader PAM, devise an intein split strategy to package the otherwise oversized adenine base editor into AAV, and show it efficiently restores dystrophin expression in muscle and heart when systemically injected in a mouse model of DMD
- Li Xu
- , Chen Zhang
- & Renzhi Han
-
Article
| Open Access CRISPECTOR provides accurate estimation of genome editing translocation and off-target activity from comparative NGS data
CRISPECTOR provides accurate estimation of genome editing translocation and off-target activity from comparative NGS data
The control of off-target activity is a challenge for adapting CRISPR to therapeutic use. Here the authors present CRISPECTOR, a software tool to detect, evaluate and quantify editing activity, including translocations, from NGS data.
- Ido Amit
- , Ortal Iancu
- & Zohar Yakhini
-
Article
| Open Access Gene therapy via canalostomy approach preserves auditory and vestibular functions in a mouse model of Jervell and Lange-Nielsen syndrome type 2
Gene therapy via canalostomy approach preserves auditory and vestibular functions in a mouse model of Jervell and Lange-Nielsen syndrome type 2
Jervell and Lange-Nielsen syndrome is characterised by congenital deafness and vestibular dysfunction, and is caused by mutations in KCNE1 or KCNQ1. Here, the authors show that gene therapy via canalostomy at early postnatal stage can preserve the morphology of inner ear and auditory and vestibular functions in a mouse model of human JLNS2.
- Xuewen Wu
- , Li Zhang
- & Xi Lin
-
Article
| Open Access Cas9-AAV6 gene correction of beta-globin in autologous HSCs improves sickle cell disease erythropoiesis in mice
Cas9-AAV6 gene correction of beta-globin in autologous HSCs improves sickle cell disease erythropoiesis in mice
CRISPR mediated gene correction of sickle cell disease (SCD) in patient-derived hematopoietic stem cells is a promising avenue for therapy. Here the authors use a humanized SCD mouse model to study gene editing in the context of autologous transplantation.
- Adam C. Wilkinson
- , Daniel P. Dever
- & Matthew H. Porteus
-
Article
| Open Access The TRACE-Seq method tracks recombination alleles and identifies clonal reconstitution dynamics of gene targeted human hematopoietic stem cells
The TRACE-Seq method tracks recombination alleles and identifies clonal reconstitution dynamics of gene targeted human hematopoietic stem cells
Genetic barcoding has been used to track clonal dynamics of cells. Here, the authors develop a Tracking Recombination Alleles in Clonal Engraftment using sequencing (TRACE-Seq), to barcode repaired alleles by introducing silent mutations or outside of coding regions, to show clonal complexity of edited CD34 + cells following engraftment.
- Rajiv Sharma
- , Daniel P. Dever
- & Ravindra Majeti
-
Article
| Open Access MiCas9 increases large size gene knock-in rates and reduces undesirable on-target and off-target indel edits
MiCas9 increases large size gene knock-in rates and reduces undesirable on-target and off-target indel edits
Cas9 fused to DNA damage repair proteins can improve rates of gene knock-in but the chimeric protein is often large. Here the authors fuse Cas9 to a minimal Brex27 motif from BRCA2 consisting of thirty-six amino acids to enhance the efficacy and safety of gene editing.
- Linyuan Ma
- , Jinxue Ruan
- & Jie Xu
-
Article
| Open Access Docking sites inside Cas9 for adenine base editing diversification and RNA off-target elimination
Docking sites inside Cas9 for adenine base editing diversification and RNA off-target elimination
Current SpCas9 adenine base editors that minimise RNA off-target activities have constrained editing windows. Here the authors use domain insertion of TadA into Cas9 to narrow, expand or shift the editing window with RNA off-target minimization simultaneously
- Shuo Li
- , Bo Yuan
- & Tian-Lin Cheng
-
Article
| Open Access Prime editing for functional repair in patient-derived disease models
Prime editing for functional repair in patient-derived disease models
Prime editing uses Cas9 nickase fused to a reverse transcriptase to edit genetic information. Here, the authors prime edit primary adult stem cells in 3D organoid cultures to show functional correction of pathogenic mutations without genome-wide off-target effects.
- Imre F. Schene
- , Indi P. Joore
- & Sabine A. Fuchs
-
Article
| Open Access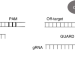 CRISPR GUARD protects off-target sites from Cas9 nuclease activity using short guide RNAs
CRISPR GUARD protects off-target sites from Cas9 nuclease activity using short guide RNAs
Off-target editing remains a concern for therapeutic applications of CRISPR-Cas9. Here the authors present CRISPR GUARD, which uses very short non-cleaving gRNAs to prevent editing at off-target sites.
- Matthew A. Coelho
- , Etienne De Braekeleer
- & Benjamin J. M. Taylor
-
Article
| Open Access Targeted gene correction of human hematopoietic stem cells for the treatment of Wiskott - Aldrich Syndrome
Targeted gene correction of human hematopoietic stem cells for the treatment of Wiskott - Aldrich Syndrome
In recent years, hematopoietic stem cells gene editing has emerged as a promising tool to treat blood disorders. Here the authors develop a CRISPR/Cas9-based genome editing strategy that allows the precise correction of Wiskott-Aldrich Syndrome in vitro and in vivo with high efficiency.
- Rajeev Rai
- , Marianna Romito
- & Alessia Cavazza
-
Article
| Open Access Ex vivo editing of human hematopoietic stem cells for erythroid expression of therapeutic proteins
Ex vivo editing of human hematopoietic stem cells for erythroid expression of therapeutic proteins
A platform for systemic therapeutic transgene expression independent of patient mutations needs a safe and highly transcribed locus. Here the authors ex vivo edit HPSCs using CRISPR-Cas9 to integrate transgenes under the α-globin promoter to achieve erythroid specific expression.
- Giulia Pavani
- , Marine Laurent
- & Mario Amendola
-
Article
| Open Access Engineering monocyte/macrophage−specific glucocerebrosidase expression in human hematopoietic stem cells using genome editing
Engineering monocyte/macrophage−specific glucocerebrosidase expression in human hematopoietic stem cells using genome editing
Gaucher disease is a lysosomal storage disorder caused by insufficient glucocerebrosidase expression. Here, the authors describe a CRISPR/Cas9-based gene-editing approach to re-express this enzyme in human blood stem cells and show that they can engraft in NSG mice and differentiate into functional macrophages.
- Samantha G. Scharenberg
- , Edina Poletto
- & Natalia Gomez-Ospina
-
Article
| Open Access Improving the safety of human pluripotent stem cell therapies using genome-edited orthogonal safeguards
Improving the safety of human pluripotent stem cell therapies using genome-edited orthogonal safeguards
Human pluripotent stem cell derived therapies can have serious safety risks. Here the authors design two drug inducible genetic safeguards to deplete undifferentiated hPSCs and hPSC-derived cell types.
- Renata M. Martin
- , Jonas L. Fowler
- & Kyle M. Loh
-
Article
| Open Access Chemical modifications of adenine base editor mRNA and guide RNA expand its application scope
Chemical modifications of adenine base editor mRNA and guide RNA expand its application scope
Cas9 base editors are promising tools for correcting pathogenic single nucleotide mutations. Here the authors chemically modify mRNA encoding the editor and the gRNA to enhance editing and broaden its application.
- Tingting Jiang
- , Jordana M. Henderson
- & Wen Xue
-
Article
| Open Access Extracellular nanovesicles for packaging of CRISPR-Cas9 protein and sgRNA to induce therapeutic exon skipping
Extracellular nanovesicles for packaging of CRISPR-Cas9 protein and sgRNA to induce therapeutic exon skipping
Expression of Cas9 and gRNA from viral vectors in vivo may cause off-target activity. Here the authors present NanoMEDIC, which uses nanovesicles to transiently deliver editing machinery to hard-to-transfect cells.
- Peter Gee
- , Mandy S. Y. Lung
- & Akitsu Hotta

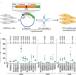 Translation velocity determines the efficacy of engineered suppressor tRNAs on pathogenic nonsense mutations
Translation velocity determines the efficacy of engineered suppressor tRNAs on pathogenic nonsense mutations