Article
|
Open Access
Featured
-
-
Article
| Open Access Interpretable and tractable models of transcriptional noise for the rational design of single-molecule quantification experiments
Interpretable and tractable models of transcriptional noise for the rational design of single-molecule quantification experiments
Here the authors explore the distributional differences expected from distinct biophysical models of transcription and show how measurements from single-cell genomics experiments can shed light on the underlying biological processes.
- Gennady Gorin
- , John J. Vastola
- & Lior Pachter
-
Article
| Open Access Differential regulation of alternative promoters emerges from unified kinetics of enhancer-promoter interaction
Differential regulation of alternative promoters emerges from unified kinetics of enhancer-promoter interaction
Alternative promoters differ in their expression patterns, whose mechanisms are not well understood. Here the authors show that alternative promoters of a Drosophila embryonic gene hunchback are regulated by different action modes of two enhancers.
- Jingyao Wang
- , Shihe Zhang
- & Heng Xu
-
Article
| Open Access Physics-informed deep learning characterizes morphodynamics of Asian soybean rust disease
Physics-informed deep learning characterizes morphodynamics of Asian soybean rust disease
Deep learning (DL) can be used to automatically extract complex features from dynamic systems. Here, the authors combine high-content imaging, DL and mechanistic models to extract and explain drug-induced morphological changes in the growth of the fungus responsible for Asian soybean rust.
- Henry Cavanagh
- , Andreas Mosbach
- & Robert G. Endres
-
Article
| Open Access Rapid incidence estimation from SARS-CoV-2 genomes reveals decreased case detection in Europe during summer 2020
Rapid incidence estimation from SARS-CoV-2 genomes reveals decreased case detection in Europe during summer 2020
The true number of infections from SARS-Cov-2 is unknown and believed to exceed the reported numbers by several fold. National testing policies, in particular, can strongly affect the proportion of undetected cases. Here, the authors propose a method that reconstructs incidence profiles within minutes, solely from publicly available, time-stamped viral genomes.
- Maureen Rebecca Smith
- , Maria Trofimova
- & Max von Kleist
-
Article
| Open Access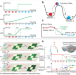 Dissecting transition cells from single-cell transcriptome data through multiscale stochastic dynamics
Dissecting transition cells from single-cell transcriptome data through multiscale stochastic dynamics
How to infer transient cells and cell-fate transitions from snap-shot single cell transcriptome dataset remains a major challenge. Here the authors present a multiscale approach to construct single-cell dynamical manifold, quantify cell stability, and compute transition trajectory and probability between cell states.
- Peijie Zhou
- , Shuxiong Wang
- & Qing Nie
-
Article
| Open Access Exploiting evolutionary steering to induce collateral drug sensitivity in cancer
Exploiting evolutionary steering to induce collateral drug sensitivity in cancer
Evolutionary steering uses therapies to control tumour evolution by exploiting trade-offs. Here, using a barcoding approach applied to large cell populations, the authors explore evolutionary steering in lung cancer cells treated with EGFR inhibitors.
- Ahmet Acar
- , Daniel Nichol
- & Andrea Sottoriva
-
Article
| Open Access Senescent cell turnover slows with age providing an explanation for the Gompertz law
Senescent cell turnover slows with age providing an explanation for the Gompertz law
One of the underlying causes of aging is the accumulation of senescent cells, but their turnover rates and dynamics during ageing are unknown. Here the authors measure and model senescent cell production and removal and explore implications for mortality.
- Omer Karin
- , Amit Agrawal
- & Uri Alon
-
Article
| Open Access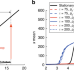 Transient hysteresis and inherent stochasticity in gene regulatory networks
Transient hysteresis and inherent stochasticity in gene regulatory networks
Cell fate commitment is understood in terms of bistable regulatory circuits with hysteresis, but inherent stochasticity in gene expression is incompatible with hysteresis. Here, the authors quantify how, under slow dynamics, the dependency of the non-stationary solutions on the initial state of the cells can lead to transient hysteresis.
- M. Pájaro
- , I. Otero-Muras
- & A. A. Alonso
-
Article
| Open Access Asymmetric division events promote variability in cell cycle duration in animal cells and Escherichia coli
Asymmetric division events promote variability in cell cycle duration in animal cells and Escherichia coli
We know that variations in cell cycle duration between cells naturally occur but the mechanisms are largely unknown. Here, using lineage tracking, hierarchical clustering and Monte Carlo methods, the authors show that large differences in granddaughter cell cycle duration are driven by asymmetric divisions.
- Ulrich Berge
- , Daria Bochenek
- & Ruth Kroschewski
-
Article
| Open Access Mitochondrial origins of fractional control in regulated cell death
Mitochondrial origins of fractional control in regulated cell death
Phenotypic cell-to-cell variability contributes to fractional killing, but the mechanisms are incompletely understood. Here the authors show that mitochondrial density correlates with cell survival in response to TRAIL, and that variable effective concentrations of Bax/Bak contribute to the effect.
- Luís C. Santos
- , Robert Vogel
- & Pablo Meyer
-
Article
| Open Access Escherichia coli can survive stress by noisy growth modulation
Escherichia coli can survive stress by noisy growth modulation
Noisy gene expression leading to phenotypic variability can help organisms to survive in changing environments. Here, Patange et al. show that noisy expression of a stress response regulator, RpoS, allows E. coli cells to modulate their growth rates to survive future adverse environments.
- Om Patange
- , Christian Schwall
- & James C. W. Locke
-
Article
| Open Access An information-theoretic framework for deciphering pleiotropic and noisy biochemical signaling
An information-theoretic framework for deciphering pleiotropic and noisy biochemical signaling
Signalling responses are marked by substantial cell-to-cell variability. Here, the authors propose an information theoretic framework that accounts for multiple inputs and temporal dynamics to analyse how signals flow through shared network components.
- Tomasz Jetka
- , Karol Nienałtowski
- & Michał Komorowski
-
Article
| Open Access Sources, propagation and consequences of stochasticity in cellular growth
Sources, propagation and consequences of stochasticity in cellular growth
The drivers of growth rate variability in bacteria are yet unknown. Here, the authors present a theory to predict the growth dynamics of individual cells and use a stochastic cell model integrating metabolism, gene expression and replication to identify the processes that underlie growth variation.
- Philipp Thomas
- , Guillaume Terradot
- & Andrea Y. Weiße
-
Article
| Open Access Homeostasis of protein and mRNA concentrations in growing cells
Homeostasis of protein and mRNA concentrations in growing cells
For various organisms, mRNA and protein copy numbers scale with cell volume. Here, the authors show that this result emerges naturally when ribosomes and RNAPs limit expression. Furthermore, the authors show that within their model this result breaks down for a sufficiently high volume/DNA ratio.
- Jie Lin
- & Ariel Amir
-
Article
| Open Access Linear mapping approximation of gene regulatory networks with stochastic dynamics
Linear mapping approximation of gene regulatory networks with stochastic dynamics
The intractability of most stochastic models of gene regulatory networks (GRNs) limits their utility. Here, the authors present a linear-mapping approximation mapping models onto simpler ones, giving approximate but accurate analytic or semi- analytic solutions for a wide range of model GRNs.
- Zhixing Cao
- & Ramon Grima
-
Article
| Open Access Large-scale genetic analysis reveals mammalian mtDNA heteroplasmy dynamics and variance increase through lifetimes and generations
Large-scale genetic analysis reveals mammalian mtDNA heteroplasmy dynamics and variance increase through lifetimes and generations
Mitochondrial populations in cells may consist of heteroplasmic mixtures of mtDNA types, and their evolution through development, aging and generations is central to genetic diseases. Here the authors dissect these population dynamics using a large mouse-based data set to characterise the dynamics of heteroplasmy mean and variance throughout life and across generations.
- Joerg P. Burgstaller
- , Thomas Kolbe
- & Iain G. Johnston
-
Article
| Open Access Cell fate in antiviral response arises in the crosstalk of IRF, NF-κB and JAK/STAT pathways
Cell fate in antiviral response arises in the crosstalk of IRF, NF-κB and JAK/STAT pathways
Innate immunity combines intra- and intercellular signalling to develop responses that limit pathogen spread. Here the authors analyse feedback and feedforward loops connecting IRF3, NF-κB and STAT pathways, and suggest they allow coordinating cell fate decisions in cellular populations in response to the virus-mimicking agent poly(I:C).
- Maciej Czerkies
- , Zbigniew Korwek
- & Tomasz Lipniacki
-
Article
| Open Access The Chemical Fluctuation Theorem governing gene expression
The Chemical Fluctuation Theorem governing gene expression
A unified framework to understand gene expression noise is still lacking. Here the authors derive a universal theorem relating the biological noise with dynamics of birth and death processes and present a model of transcription dynamics, allowing analytical prediction of the dependence of mRNA noise on mRNA lifetime variability.
- Seong Jun Park
- , Sanggeun Song
- & Jaeyoung Sung
-
Article
| Open Access Stochastic gene expression in Arabidopsis thaliana
Stochastic gene expression in Arabidopsis thaliana
Noisy gene expression can cause stochasticity in the expression of plant traits. Here, Araújo et al. use a dual reporter system of protein expression in Arabidopsis to show that expression noise is lowest in stomata relative to other tissues and that leaf cells are coupled with respect to noise.
- Ilka Schultheiß Araújo
- , Jessica Magdalena Pietsch
- & Martin Hülskamp
-
Article
| Open Access Cell fate decisions emerge as phages cooperate or compete inside their host
Cell fate decisions emerge as phages cooperate or compete inside their host
The bacteriophage lambda and its hostEscherichia coli provide a model system to study cell-fate decisions. Here, Trinh et al. develop a four-colour fluorescence system at the single-cell/single-virus/single-viral-DNA level and find phages cooperate during lysogenization and compete during lysis.
- Jimmy T. Trinh
- , Tamás Székely
- & Lanying Zeng
-
Article
| Open Access T-cell stimuli independently sum to regulate an inherited clonal division fate
T-cell stimuli independently sum to regulate an inherited clonal division fate
Why do populations of highly similar T cells have heterogeneous division destinies in response to antigenic stimulus? Here the authors develop a multiplex-dye assay and a mathematical framework to test clonal heterogeneity and show distinction in division destiny is a result of inter-clonal variability as lineage imprinting ensures clones share similar proliferation fates.
- J. M. Marchingo
- , G. Prevedello
- & K. R. Duffy
-
Article
| Open Access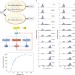 Noise reduction facilitated by dosage compensation in gene networks
Noise reduction facilitated by dosage compensation in gene networks
Cells must function despite the noisiness of their processes by tolerating or reducing such variability. Here, the authors combine experiment and modelling to show that a network motif that mediates network-dosage compensation also reduces noise in network output, suggesting that noise is tuneable.
- Weilin Peng
- , Ruijie Song
- & Murat Acar
-
Article
| Open Access 3D replicon distributions arise from stochastic initiation and domino-like DNA replication progression
3D replicon distributions arise from stochastic initiation and domino-like DNA replication progression
DNA replication in higher eukaryotes is a complex process occurring in a complex genome environment. Here the authors present a model of DNA replication that incorporates random loop chromatin folding and domino-like fork progression reproducing the spatial and temporal characteristics of S-phase.
- D. Löb
- , N. Lengert
- & B. Drossel
-
Article
| Open Access Single-cell analysis and stochastic modelling unveil large cell-to-cell variability in influenza A virus infection
Single-cell analysis and stochastic modelling unveil large cell-to-cell variability in influenza A virus infection
Cell-to-cell variability in viral infection means that cell population measurements may not be an accurate representation of the process. Using both experimental and modelling approaches the authors confirm this notion showing that influenza virus infections are variable processes affected by intrinsic and extrinsic noise.
- Frank S. Heldt
- , Sascha Y. Kupke
- & Timo Frensing
-
Article
| Open Access Stochastic signalling rewires the interaction map of a multiple feedback network during yeast evolution
Stochastic signalling rewires the interaction map of a multiple feedback network during yeast evolution
GALgenes enhance their own transcription via the transcription factor Gal4p, and the number of Galp4 sites in a promoter is expected to strengthen the feedback. In this study, Hsuet al. show that instead the feedback loops are activated by genes that have frequent bursts of expression and fast RNA decay kinetics.
- Chieh Hsu
- , Simone Scherrer
- & Attila Becskei

 Damage dynamics and the role of chance in the timing of E. coli cell death
Damage dynamics and the role of chance in the timing of E. coli cell death