Article
|
Open Access
Featured
-
-
Article
| Open Access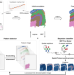 Pianno: a probabilistic framework automating semantic annotation for spatial transcriptomics
Pianno: a probabilistic framework automating semantic annotation for spatial transcriptomics
Recognising spatial spots’ biological identity in spatial transcriptomics remains a challenge. Here, authors introduce Pianno, a tool that helps annotate the biological structures or cell-type constructions across diverse tissues, offering new perspectives on understanding spatial transcriptomics.
- Yuqiu Zhou
- , Wei He
- & Ying Zhu
-
Article
| Open Access Allele-specific transcriptional effects of subclonal copy number alterations enable genotype-phenotype mapping in cancer cells
Allele-specific transcriptional effects of subclonal copy number alterations enable genotype-phenotype mapping in cancer cells
Quantifying the impact of copy-number alterations (CNAs) on gene expression at the subclone level in cancer remains a challenge. Here, the authors develop TreeAlign, a method that integrates sample-matched single-cell DNA and RNA sequencing data to infer the impact of CNAs on subclonal gene expression.
- Hongyu Shi
- , Marc J. Williams
- & Sohrab P. Shah
-
Article
| Open Access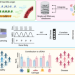 Cell type signatures in cell-free DNA fragmentation profiles reveal disease biology
Cell type signatures in cell-free DNA fragmentation profiles reveal disease biology
Deconvolution of cfDNA fragmentation benefits from cell type-specific reference data. Here, the authors create a disease agnostic cfDNA cell type of origin analysis and show it can successfully predict cell types of origin from plasma samples.
- Kate E. Stanley
- , Tatjana Jatsenko
- & Joris Robert Vermeesch
-
Article
| Open Access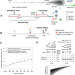 Evolving copy number gains promote tumor expansion and bolster mutational diversification
Evolving copy number gains promote tumor expansion and bolster mutational diversification
Understanding the timing and fitness of somatic copy number alterations (SCNAs) in cancer would shed light on cancer progression and evolution. Here, the authors develop Butte, a computational framework to estimate the timing of clonal SCNAs that encompass multiple gains, and apply it on whole-genome sequencing data from 184 samples.
- Zicheng Wang
- , Yunong Xia
- & Ruping Sun
-
Article
| Open Access Statistical method scDEED for detecting dubious 2D single-cell embeddings and optimizing t-SNE and UMAP hyperparameters
Statistical method scDEED for detecting dubious 2D single-cell embeddings and optimizing t-SNE and UMAP hyperparameters
2D visualisation of single-cell data is highly impacted by the hyperparameter setting of the 2D embedding method, such as t-SNE and UMAP. Here, authors develop a statistical method scDEED to detect dubious cell embeddings and optimise the hyperparameter setting for trustworthy visualisation.
- Lucy Xia
- , Christy Lee
- & Jingyi Jessica Li
-
Comment
| Open Access Fudging the volcano-plot without dredging the data
Fudging the volcano-plot without dredging the data
Selecting omic biomarkers using both their effect size and their differential status significance (i.e., selecting the “volcano-plot outer spray”) has long been equally biologically relevant and statistically troublesome. However, recent proposals are paving the way to resolving this dilemma.
- Thomas Burger
-
Article
| Open Access PheWAS-based clustering of Mendelian Randomisation instruments reveals distinct mechanism-specific causal effects between obesity and educational attainment
PheWAS-based clustering of Mendelian Randomisation instruments reveals distinct mechanism-specific causal effects between obesity and educational attainment
Mendelian Randomisation estimates causal effects between risk factors and complex outcomes using genetic variants as instrumental variables, however it can be affected by certain biases. To alleviate these biases the authors propose an approach based on clustering genetic instruments according to the types of trait they are associated with, and apply this method to revisit the surprisingly large apparent causal effect of body mass index on educational attainment.
- Liza Darrous
- , Gibran Hemani
- & Zoltán Kutalik
-
Article
| Open Access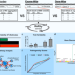 A method to estimate the contribution of rare coding variants to complex trait heritability
A method to estimate the contribution of rare coding variants to complex trait heritability
The contribution of rare variants to complex traits has not been well studied. Here, the authors present RARity, a method to assess rare variant heritability without assuming a particular genetic architecture and enabling both gene-level and exome-wide heritability estimation of continuous traits.
- Nazia Pathan
- , Wei Q. Deng
- & Guillaume Paré
-
Article
| Open Access Improving polygenic risk prediction in admixed populations by explicitly modeling ancestral-differential effects via GAUDI
Improving polygenic risk prediction in admixed populations by explicitly modeling ancestral-differential effects via GAUDI
Most polygenic risk score (PRS) methods focus only on individuals with distinct primary continental ancestry, without accommodating recently-admixed individuals. Here, the authors develop a novel penalized regression-based PRS method specifically designed for admixed individuals.
- Quan Sun
- , Bryce T. Rowland
- & Yun Li
-
Article
| Open Access DeepFocus: fast focus and astigmatism correction for electron microscopy
DeepFocus: fast focus and astigmatism correction for electron microscopy
High-throughput electron microscopy demands minimal human intervention and high image quality. Here, authors introduce DeepFocus, a data-driven method for aberration correction in electron microscopy, robust for low SNR images, fast and easily adaptable to microscopes and samples. Peer Review Information: Nature Communications thanks Yang Zhang and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. A peer review file is available.
- P. J. Schubert
- , R. Saxena
- & J. Kornfeld
-
Article
| Open Access Trajectory inference across multiple conditions with condiments
Trajectory inference across multiple conditions with condiments
scRNA-Seq has enabled the study of dynamic systems such as response to a drug at the individual cell and gene levels. Here the authors introduce a framework to interpret differences at the trajectory, cell populations, and individual gene levels.
- Hector Roux de Bézieux
- , Koen Van den Berge
- & Sandrine Dudoit
-
Article
| Open Access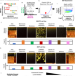 Anti-correlated feature selection prevents false discovery of subpopulations in scRNAseq
Anti-correlated feature selection prevents false discovery of subpopulations in scRNAseq
Typical single-cell RNAseq pipelines will subcluster homogeneous cells. Here, authors present a computational algorithm for accurately identifying cell-type marker genes in single-cell data analysis with a low false discovery rate.
- Scott R. Tyler
- , Daniel Lozano-Ojalvo
- & Eric E. Schadt
-
Article
| Open Access Clinical application of tumour-in-normal contamination assessment from whole genome sequencing
Clinical application of tumour-in-normal contamination assessment from whole genome sequencing
Assessing tumour contamination in normal samples is critical for accurate variant calling in cancer samples. Here, the authors develop TINC, a computational method to determine the level of tumour in normal contamination, and demonstrate its application in the Genomics England 100,000 Genomes Project dataset.
- Jonathan Mitchell
- , Salvatore Milite
- & Giulio Caravagna
-
Article
| Open Access Leveraging single-cell ATAC-seq and RNA-seq to identify disease-critical fetal and adult brain cell types
Leveraging single-cell ATAC-seq and RNA-seq to identify disease-critical fetal and adult brain cell types
This study analyzed data from human cells assayed using single-cell technologies, together with data associating genetic variants to disease, to identify fetal and brain cell types whose biologically critically influences the etiology of disease.
- Samuel S. Kim
- , Buu Truong
- & Alkes L. Price
-
Article
| Open Access DiffDomain enables identification of structurally reorganized topologically associating domains
DiffDomain enables identification of structurally reorganized topologically associating domains
Topologically associating domains (TADs) are critical structural units in 3D genome organization, and their reorganization between health and disease states is associated with essential genome functions. However, computational methods for identifying reorganized TADs are still in the early stages of development. Here, the authors present an algorithm leveraging random matrix theory to identify reorganized TADs.
- Dunming Hua
- , Ming Gu
- & Dechao Tian
-
Article
| Open Access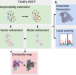 Cryo-EM structure and B-factor refinement with ensemble representation
Cryo-EM structure and B-factor refinement with ensemble representation
Cryo-EM is the go-to method for visualizing large, flexible biomolecules. Here, authors introduce a new Gaussian mixture modelling method for cryo-EM modelling tasks, including refinement, composite map generation and ensemble representation.
- Joseph G. Beton
- , Thomas Mulvaney
- & Maya Topf
-
Article
| Open Access Integrating genetic regulation and single-cell expression with GWAS prioritizes causal genes and cell types for glaucoma
Integrating genetic regulation and single-cell expression with GWAS prioritizes causal genes and cell types for glaucoma
The molecular and cellular causes of glaucoma are not well understood. Here, the authors integrate GWAS with genetic regulation and single cell expression from multiple eye tissues to identify genes and key cell types that affect glaucoma pathogenesis.
- Andrew R. Hamel
- , Wenjun Yan
- & Ayellet V. Segrè
-
Article
| Open Access ACIDES: on-line monitoring of forward genetic screens for protein engineering
ACIDES: on-line monitoring of forward genetic screens for protein engineering
Screening mutated proteins is a versatile strategy in protein research, producing massive datasets when combined with NGS. Here, authors present ACIDES to estimate mutated protein fitness and aid protein engineering pipelines in a range of applications, including gene therapy.
- Takahiro Nemoto
- , Tommaso Ocari
- & Ulisse Ferrari
-
Article
| Open Access Revealing hidden patterns in deep neural network feature space continuum via manifold learning
Revealing hidden patterns in deep neural network feature space continuum via manifold learning
Existing feature visualisation methods are not well-suited for regression tasks. Here, authors introduce a method to learn the manifold topology related to deep neural network output and target labels and provide insightful visualisations of the high-dimensional features while preserving the local geometry.
- Md Tauhidul Islam
- , Zixia Zhou
- & Lei Xing
-
Article
| Open Access JOINTLY: interpretable joint clustering of single-cell transcriptomes
JOINTLY: interpretable joint clustering of single-cell transcriptomes
Batch integration is a critical yet challenging step in many single-cell RNA-seq analysis workflows. Here, authors present JOINTLY, a hybrid linear and non-linear NMF-based algorithm, providing interpretable and robust cell clustering against over-integration.
- Andreas Fønss Møller
- & Jesper Grud Skat Madsen
-
Article
| Open Access Haplotype-based inference of recent effective population size in modern and ancient DNA samples
Haplotype-based inference of recent effective population size in modern and ancient DNA samples
The authors introduce a new computational method, HapNe, for inferring the recent effective size of human populations. HapNe does not require high-quality genotype data, making it suitable for the study of ancient DNA samples.
- Romain Fournier
- , Zoi Tsangalidou
- & Pier Francesco Palamara
-
Article
| Open Access CurveCurator: a recalibrated F-statistic to assess, classify, and explore significance of dose–response curves
CurveCurator: a recalibrated F-statistic to assess, classify, and explore significance of dose–response curves
Dose-response curves are ubiquitous in pharmacology and biology, yet potency and effect size are often estimated even when there is no response. Here, authors present a statistical framework to assess curve significance and demonstrate how this aids drug mode of action analysis in large public datasets.
- Florian P. Bayer
- , Manuel Gander
- & Matthew The
-
Article
| Open Access Augmenting interpretable models with large language models during training
Augmenting interpretable models with large language models during training
Prediction and interpretation tasks may be challenging in high-stakes applications, such as medical decision-making, or systems with compute-limited hardware. The authors introduce an augmented framework for leveraging the knowledge learned by Large Language Models to build interpretable models which are both accurate and efficient.
- Chandan Singh
- , Armin Askari
- & Jianfeng Gao
-
Article
| Open Access Integrating spatial and single-cell transcriptomics data using deep generative models with SpatialScope
Integrating spatial and single-cell transcriptomics data using deep generative models with SpatialScope
Spatial transcriptomics (ST) is transforming tissue analysis but has limitations. Here, authors introduce SpatialScope, an integrated approach combining scRNA-seq and ST data using deep generative models, enabling comprehensive spatial characterisation at transcriptome-wide single-cell resolution.
- Xiaomeng Wan
- , Jiashun Xiao
- & Can Yang
-
Article
| Open Access Paired single-cell multi-omics data integration with Mowgli
Paired single-cell multi-omics data integration with Mowgli
Mowgli is a novel paired single-cell multi-omics integration method leveraging matrix factorization and Optimal Transport. In-depth benchmarking demonstrates promising cell clustering results and improved biological interpretability.
- Geert-Jan Huizing
- , Ina Maria Deutschmann
- & Laura Cantini
-
Article
| Open Access Dimension-agnostic and granularity-based spatially variable gene identification using BSP
Dimension-agnostic and granularity-based spatially variable gene identification using BSP
Identifying spatially variable genes (SVGs) is essential for linking molecular cell functions with tissue phenotypes. Here, authors introduce a non-parametric model that detects SVGs from two or three-dimensional spatial transcriptomics data by comparing gene expression patterns at granularities.
- Juexin Wang
- , Jinpu Li
- & Dong Xu
-
Article
| Open Access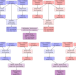 Leveraging information between multiple population groups and traits improves fine-mapping resolution
Leveraging information between multiple population groups and traits improves fine-mapping resolution
Statistical fine-mapping helps to pinpoint likely causal variants underlying genetic association signals, and can be enhanced by using multi-ancestry datasets. Here, the authors introduce MGflashfm, a fine-mapping method for pinpointing likely causal variants amongst multiple traits and population groups.
- Feng Zhou
- , Opeyemi Soremekun
- & Jennifer L. Asimit
-
Article
| Open Access A statistical framework for differential pseudotime analysis with multiple single-cell RNA-seq samples
A statistical framework for differential pseudotime analysis with multiple single-cell RNA-seq samples
Pseudotime analysis is prevalent in single-cell RNA-seq, but it remains challenging to perform it across multiple samples and experimental conditions. Here, the authors develop Lamian, a computational framework for multi-sample pseudotime analysis that adjusts for biological and technical variation to detect gene program changes along cell trajectories and across conditions.
- Wenpin Hou
- , Zhicheng Ji
- & Hongkai Ji
-
Article
| Open Access Unappreciated subcontinental admixture in Europeans and European Americans and implications for genetic epidemiology studies
Unappreciated subcontinental admixture in Europeans and European Americans and implications for genetic epidemiology studies
European ancestry individuals are not typically treated as admixed in genetic studies. Here, the authors detect higher than expected admixture in European populations, which could potentially affect the results of genetic studies if it is not accounted for.
- Mateus H. Gouveia
- , Amy R. Bentley
- & Daniel Shriner
-
Article
| Open Access XMAP: Cross-population fine-mapping by leveraging genetic diversity and accounting for confounding bias
XMAP: Cross-population fine-mapping by leveraging genetic diversity and accounting for confounding bias
Fine-mapping prioritizes risk variants identified by genome-wide association studies to uncover biological mechanisms underlying complex traits. Here, the authors develop a reliable fine-mapping method (XMAP) by leveraging genetic diversity and accounting for confounding bias.
- Mingxuan Cai
- , Zhiwei Wang
- & Can Yang
-
Article
| Open Access Single-cell allele-specific expression analysis reveals dynamic and cell-type-specific regulatory effects
Single-cell allele-specific expression analysis reveals dynamic and cell-type-specific regulatory effects
Here the authors develop DAESC, a statistical method for differential allele-specific expression analysis using single-cell RNA-seq data. Application of DAESC identifies dynamic regulatory effects along endoderm differentiation and differential effects between type 2 diabetes and healthy controls.
- Guanghao Qi
- , Benjamin J. Strober
- & Alexis Battle
-
Article
| Open Access MetaCC allows scalable and integrative analyses of both long-read and short-read metagenomic Hi-C data
MetaCC allows scalable and integrative analyses of both long-read and short-read metagenomic Hi-C data
The authors develop an integrative and scalable framework to eliminate systematic biases and retrieve high-quality metagenome-assembled genomes using either long-read or short-read metagenomic Hi-C data.
- Yuxuan Du
- & Fengzhu Sun
-
Article
| Open Access Global burden of disease due to rifampicin-resistant tuberculosis: a mathematical modeling analysis
Global burden of disease due to rifampicin-resistant tuberculosis: a mathematical modeling analysis
Rifampicin-resistant tuberculosis (RR-TB) requires longer, more toxic therapy than rifampicin-sensitive disease and is associated with a higher occurrence of long-term sequelae. In this mathematical modeling study, the authors estimate that incident RR-TB in 2020 will be responsible for ~6.9 million disability-adjusted life years; 44% due to post-tuberculosis sequelae.
- Nicolas A. Menzies
- , Brian W. Allwood
- & Ted Cohen
-
Article
| Open Access scBridge embraces cell heterogeneity in single-cell RNA-seq and ATAC-seq data integration
scBridge embraces cell heterogeneity in single-cell RNA-seq and ATAC-seq data integration
Multi-omics data integration can be challenging in the event of cell heterogeneity. Here, the authors present scBridge, a method that exploits heterogeneous omics differences, to progressively integrate cells and narrows omics gap, leading to promising integration and label transfer results.
- Yunfan Li
- , Dan Zhang
- & Xi Peng
-
Article
| Open Access Genome-wide enhancer-gene regulatory maps link causal variants to target genes underlying human cancer risk
Genome-wide enhancer-gene regulatory maps link causal variants to target genes underlying human cancer risk
Here, the authors apply the Activity-by-Contact (ABC) model to infer enhancer-gene regulation and the effect of associated variants across multiple cancer types, integrating genetic and multi-omics data. Then, they explore the mechanisms associated with ABC regulatory variants in colorectal cancer.
- Pingting Ying
- , Can Chen
- & Xiaoping Miao
-
Article
| Open Access The Oncology Biomarker Discovery framework reveals cetuximab and bevacizumab response patterns in metastatic colorectal cancer
The Oncology Biomarker Discovery framework reveals cetuximab and bevacizumab response patterns in metastatic colorectal cancer
Identifying actionable biomarkers remains a challenge. Here, the authors develop a framework Oncology Biomarker Discovery (OncoBird), apply it to a phase III trial and investigate the molecular and biomarker landscape of metastatic colorectal carcinoma patients.
- Alexander J. Ohnmacht
- , Arndt Stahler
- & Michael P. Menden
-
Article
| Open Access Single-cell genomics improves the discovery of risk variants and genes of atrial fibrillation
Single-cell genomics improves the discovery of risk variants and genes of atrial fibrillation
Here the authors combine an experimental and analytical approach that integrates single cell epigenomics with GWAS to prioritize risk variants and genes to provide a comprehensive map of Atrial Fibrillation risk variants and genes.
- Alan Selewa
- , Kaixuan Luo
- & Sebastian Pott
-
Article
| Open Access SnapFISH: a computational pipeline to identify chromatin loops from multiplexed DNA FISH data
SnapFISH: a computational pipeline to identify chromatin loops from multiplexed DNA FISH data
Multiplexed DNA FISH technologies are powerful tools to reveal chromatin spatial organisation. Here, the authors developed SnapFISH, a computational pipeline to identify chromatin loops from multiplexed DNA FISH data.
- Lindsay Lee
- , Hongyu Yu
- & Ming Hu
-
Article
| Open Access Cell-type-specific co-expression inference from single cell RNA-sequencing data
Cell-type-specific co-expression inference from single cell RNA-sequencing data
Inferring co-expressions with scRNA-seq data is challenging, and existing methods suffer from inflated false positives and biases. Here, the authors proposed CS-CORE, which yields unbiased estimates and identifies co-expressions that are more reproducible and biologically relevant for scRNA-seq data.
- Chang Su
- , Zichun Xu
- & Jingfei Zhang
-
Article
| Open Access SONAR enables cell type deconvolution with spatially weighted Poisson-Gamma model for spatial transcriptomics
SONAR enables cell type deconvolution with spatially weighted Poisson-Gamma model for spatial transcriptomics
Spatial transcriptomics reveal cellular profiles with spatial context. Here the authors present SONAR, a computational model that utilizes spatial information to decipher cell types in tissues and validate on various spatial patterns and fine-mapped cell types in complex tissues.
- Zhiyuan Liu
- , Dafei Wu
- & Liang Ma
-
Article
| Open Access Multi-PGS enhances polygenic prediction by combining 937 polygenic scores
Multi-PGS enhances polygenic prediction by combining 937 polygenic scores
Polygenic scores (PGS) have high potential for clinical use but are currently underpowered for many applications. Here, the authors develop an approach that leverages an agnostic library of hundreds of PGS to increase prediction of complex diseases and other traits. This multi-PGS framework is ideal for emerging biobank data.
- Clara Albiñana
- , Zhihong Zhu
- & Bjarni J. Vilhjálmsson
-
Article
| Open Access Atlas-scale single-cell multi-sample multi-condition data integration using scMerge2
Atlas-scale single-cell multi-sample multi-condition data integration using scMerge2
Recent advances in multi-condition single-cell multi-cohort studies enable exploration of diverse cell states. Here, authors present scMerge2, an algorithm that allows integration of a large COVID-19 data collection with over five million cells to uncover distinct signatures of disease progression.
- Yingxin Lin
- , Yue Cao
- & Jean Y. H. Yang
-
Article
| Open Access Multi-batch single-cell comparative atlas construction by deep learning disentanglement
Multi-batch single-cell comparative atlas construction by deep learning disentanglement
Comparing single-cell RNA-seq and ATAC-seq data from multiple batches is challenging due to technical artifacts. Here, the authors propose a method that disentangles technical and biological effects, facilitating batch-confounded chromatin and gene expression state discovery and enhancing the analysis of perturbation effects on cell populations.
- Allen W. Lynch
- , Myles Brown
- & Clifford A. Meyer
-
Article
| Open Access The role of vaccination and public awareness in forecasts of Mpox incidence in the United Kingdom
The role of vaccination and public awareness in forecasts of Mpox incidence in the United Kingdom
An outbreak of Mpox in the UK began in May 2022 and peaked in July. In this modelling study, the authors show that the decline in cases was likely due to behavioural changes among high-risk populations, whilst vaccination could prevent a rebound.
- Samuel P. C. Brand
- , Massimo Cavallaro
- & Matt J. Keeling
-
Article
| Open Access Genome-wide association analysis and Mendelian randomization proteomics identify drug targets for heart failure
Genome-wide association analysis and Mendelian randomization proteomics identify drug targets for heart failure
Here, the authors perform a large-scale meta-analysis of genome-wide association studies and cis-MR proteomics to identify protein biomarkers and drug targets for heart failure.
- Danielle Rasooly
- , Gina M. Peloso
- & Juan P. Casas
-
Article
| Open Access nnSVG for the scalable identification of spatially variable genes using nearest-neighbor Gaussian processes
nnSVG for the scalable identification of spatially variable genes using nearest-neighbor Gaussian processes
The identification of top spatially variable genes is a key step in the analysis of spatially-resolved transcriptomics data. Here, the authors develop a scalable method based on nearest-neighbor Gaussian processes and evaluate performance compared to existing and baseline methods.
- Lukas M. Weber
- , Arkajyoti Saha
- & Stephanie C. Hicks
-
Article
| Open Access Leveraging spatial transcriptomics data to recover cell locations in single-cell RNA-seq with CeLEry
Leveraging spatial transcriptomics data to recover cell locations in single-cell RNA-seq with CeLEry
Cell location information is important for understanding how tissue is spatially organized. Here, the authors develop CeLEry, a machine learning method that aims to recover cell locations for single-cell RNA-seq data by leveraging information learned from spatial transcriptomics.
- Qihuang Zhang
- , Shunzhou Jiang
- & Mingyao Li
-
Article
| Open Access SpatialDM for rapid identification of spatially co-expressed ligand–receptor and revealing cell–cell communication patterns
SpatialDM for rapid identification of spatially co-expressed ligand–receptor and revealing cell–cell communication patterns
Spatial omics are increasingly being recognised to study cell-cell communications. Here, the authors present a bioinformatics toolbox for rapid identification of spatially co-expressed ligand-receptor and revealing cell-cell communication patterns.
- Zhuoxuan Li
- , Tianjie Wang
- & Yuanhua Huang
-
Article
| Open Access Joint analysis of phenotype-effect-generation identifies loci associated with grain quality traits in rice hybrids
Joint analysis of phenotype-effect-generation identifies loci associated with grain quality traits in rice hybrids
Genetic dissection of hybrids is more difficult than inbreds as nonadditive effects are involved. Here, the authors report a pipeline for joint analysis of phenotypes, effects, and generations and demonstrate its usefulness in identification of loci associated with quality traits and improving predict accuracy in genomic selection of hybrid rice.
- Lanzhi Li
- , Xingfei Zheng
- & Zhongli Hu
 Data-driven recombination detection in viral genomes
Data-driven recombination detection in viral genomes