Article
|
Open Access
Featured
-
-
Article
| Open Access Alternative splicing of latrophilin-3 controls synapse formation
Alternative splicing of latrophilin-3 controls synapse formation
Latrophilin-3 organizes synapses through a convergent dual-pathway mechanism in which Gαs signalling is activated and phase-separated postsynaptic protein scaffolds are recruited.
- Shuai Wang
- , Chelsea DeLeon
- & Thomas C. Südhof
-
Article
| Open Access Structural basis of catalytic activation in human splicing
Structural basis of catalytic activation in human splicing
Cryogenic electron microscopy images of a spliceosome complex undergoing catalytic activation provide mechanistic insight into how the two ATP-dependent RNA helicases involved in this process, PRP2 and Aquarius, work together.
- Jana Schmitzová
- , Constantin Cretu
- & Vladimir Pena
-
Article |
 CDK11 regulates pre-mRNA splicing by phosphorylation of SF3B1
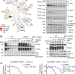
CDK11 regulates pre-mRNA splicing by phosphorylation of SF3B1
CDK11 associates with SF3B1 and phosphorylates threonine residues at the N terminus of SF3B1 during spliceosome activation, and the inhibition of CDK11 blocks the activation and leads to widespread intron retention and the accumulation of non-functional spliceosomes on pre-mRNAs and chromatin.
- Milan Hluchý
- , Pavla Gajdušková
- & Dalibor Blazek
-
Article |
 Transcriptome variation in human tissues revealed by long-read sequencing
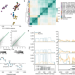
Transcriptome variation in human tissues revealed by long-read sequencing
To understand the contribution of variants to transcript expression regulation, long-read transcriptome data are generated from the GTEx resource, and a new software package to perform allele-specific analysis is developed.
- Dafni A. Glinos
- , Garrett Garborcauskas
- & Beryl B. Cummings
-
Article |
 Silent mutations reveal therapeutic vulnerability in RAS Q61 cancers
Silent mutations reveal therapeutic vulnerability in RAS Q61 cancers
A translationally silent KRASG60G mutation, preventing the formation of a cryptic splice donor site and enabling expression of KRAS(Q61K), reveals a vulnerability in RASQ61 cancers that are therapeutically exploitable in a mutant-selective manner.
- Yoshihisa Kobayashi
- , Chhayheng Chhoeu
- & Pasi A. Jänne
-
Article
| Open Access TDP-43 represses cryptic exon inclusion in the FTD–ALS gene UNC13A
TDP-43 represses cryptic exon inclusion in the FTD–ALS gene UNC13A
TDP-43 controls an exon splicing event in UNC13A that results in the inclusion of a cryptic exon associated with frontotemporal dementia and amyotrophic lateral sclerosis.
- X. Rosa Ma
- , Mercedes Prudencio
- & Aaron D. Gitler
-
Article
| Open Access Structural insights into how Prp5 proofreads the pre-mRNA branch site
Structural insights into how Prp5 proofreads the pre-mRNA branch site
The cryo-electron microscopy structure of a newly identified, early spliceosomal complex reveals the mechanism by which the RNA helicase Prp5 enhances the fidelity of the excision of introns from precursor mRNAs.
- Zhenwei Zhang
- , Norbert Rigo
- & Reinhard Lührmann
-
Article |
 Molecular architecture of the human 17S U2 snRNP
Molecular architecture of the human 17S U2 snRNP
The cryo-EM structure of human U2 small nuclear ribonucleoprotein (snRNP) offers insights into what rearrangements are required for this snRNP to be stably incorporated into the spliceosome, and the role that the DEAD-box ATPase PRP5 may have in these rearrangements.
- Zhenwei Zhang
- , Cindy L. Will
- & Holger Stark
-
Article |
 Recurrent noncoding U1 snRNA mutations drive cryptic splicing in SHH medulloblastoma
Recurrent noncoding U1 snRNA mutations drive cryptic splicing in SHH medulloblastoma
Highly recurrent hotspot r.3A>G mutations are identified in U1 splicesomal small nuclear RNAs in about 50% of Sonic hedgehog medulloblastomas, which result in disrupted RNA splicing and the activation of oncogenes.
- Hiromichi Suzuki
- , Sachin A. Kumar
- & Michael D. Taylor
-
Article |
 The U1 spliceosomal RNA is recurrently mutated in multiple cancers
The U1 spliceosomal RNA is recurrently mutated in multiple cancers
A highly recurrent A>C somatic mutation in U1 small nuclear RNA, which alters the splicing pattern of genes that include known drivers of cancer, is identified in several types of tumour.
- Shimin Shuai
- , Hiromichi Suzuki
- & Lincoln D. Stein
-
Letter |
 Coordinated alterations in RNA splicing and epigenetic regulation drive leukaemogenesis
Coordinated alterations in RNA splicing and epigenetic regulation drive leukaemogenesis
Analyses of transcriptomes from patients with acute myeloid leukaemia identified frequently co-occurring mutations of IDH2 and SRSF2, which functional analyses showed to have distinct and coordinated leukaemogenic effects on the epigenome and RNA splicing.
- Akihide Yoshimi
- , Kuan-Ting Lin
- & Omar Abdel-Wahab
-
Article |
 A unified mechanism for intron and exon definition and back-splicing
A unified mechanism for intron and exon definition and back-splicing
The cryo-electron microscopy structures of an early spliceosome complex in yeast reveal a unified mechanism for defining introns and exons and also for back-splicing to generate circular RNA.
- Xueni Li
- , Shiheng Liu
- & Rui Zhao
-
Letter |
 Arabidopsis FLL2 promotes liquid–liquid phase separation of polyadenylation complexes
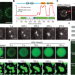
Arabidopsis FLL2 promotes liquid–liquid phase separation of polyadenylation complexes
A genetic screen for factors required by the Arabidopsis RNA-binding protein FCA identifies FLL2 as necessary in the formation of FCA nuclear bodies, and thus a role for FLL2 in liquid–liquid phase separation.
- Xiaofeng Fang
- , Liang Wang
- & Caroline Dean
-
Article |
 Introns are mediators of cell response to starvation
Introns are mediators of cell response to starvation
Transcriptomic and genetic analyses of a deletion set of all known introns in genes of the budding yeast Saccharomyces cerevisiae indicate that introns promote resistance to starvation.
- Julie Parenteau
- , Laurine Maignon
- & Sherif Abou Elela
-
Letter |
 Prespliceosome structure provides insights into spliceosome assembly and regulation
Prespliceosome structure provides insights into spliceosome assembly and regulation
The cryo-electron microscopy structure of the Saccharomyces cerevisiae prespliceosome provides insights into splice-site selection and early spliceosome assembly events.
- Clemens Plaschka
- , Pei-Chun Lin
- & Kiyoshi Nagai
-
Letter |
 piRNA-mediated regulation of transposon alternative splicing in the soma and germ line
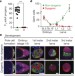
piRNA-mediated regulation of transposon alternative splicing in the soma and germ line
A new mechanism of pre-mRNA splicing regulation is revealed that is mediated by piRNA pathway components and is dependent on heterochromatin histone modifications.
- Felipe Karam Teixeira
- , Martyna Okuniewska
- & Ruth Lehmann
-
Article |
 Structure of a pre-catalytic spliceosome
Structure of a pre-catalytic spliceosome
The cryo-electron microscopy structure of the yeast spliceosome in a pre-catalytic state provides insights into the molecular events leading to formation of the spliceosome active site.
- Clemens Plaschka
- , Pei-Chun Lin
- & Kiyoshi Nagai
-
Letter |
 Structure of a spliceosome remodelled for exon ligation
Structure of a spliceosome remodelled for exon ligation
The cryo-electron microscopy structure of a yeast spliceosome stalled before mature RNA formation provides insight into the mechanism of exon ligation.
- Sebastian M. Fica
- , Chris Oubridge
- & Kiyoshi Nagai
-
Article |
 Cryo-EM structure of a human spliceosome activated for step 2 of splicing
Cryo-EM structure of a human spliceosome activated for step 2 of splicing
The cryo-EM structure of the splicing intermediate known as the C* complex from human.
- Karl Bertram
- , Dmitry E. Agafonov
- & Reinhard Lührmann
-
Letter |
 Genome-wide changes in lncRNA, splicing, and regional gene expression patterns in autism
Genome-wide changes in lncRNA, splicing, and regional gene expression patterns in autism
Gene expression analysis in brain tissue from individuals with and without autism spectrum disorder (ASD) suggests that the transcription factor SOX5 contributes to an ASD-associated reduction in transcriptional differences between brain areas and indicates that common transcriptomic changes occur in different forms of ASD.
- Neelroop N. Parikshak
- , Vivek Swarup
- & Daniel H. Geschwind
-
Letter |
 Splicing factor 1 modulates dietary restriction and TORC1 pathway longevity in C. elegans
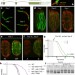
Splicing factor 1 modulates dietary restriction and TORC1 pathway longevity in C. elegans
Precursor mRNA splicing homeostasis is a biomarker and predictor of life expectancy in Caenorhabditis elegans and defects in global pre-mRNA splicing associated with age are reduced by dietary restriction via splicing factor 1.
- Caroline Heintz
- , Thomas K. Doktor
- & William B. Mair
-
Letter |
 Mechanism for DNA transposons to generate introns on genomic scales
Mechanism for DNA transposons to generate introns on genomic scales
The observations that introns are acquired in bursts and that exons are often nucleosome-sized can be explained by the generation of introns from DNA transposons, which insert between nucleosomes.
- Jason T. Huff
- , Daniel Zilberman
- & Scott W. Roy
-
Article |
 Cryo-EM structure of the spliceosome immediately after branching
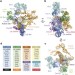
Cryo-EM structure of the spliceosome immediately after branching
Cryo-EM reveals the configuration of substrate pre-mRNA within the active spliceosome and suggests how remodelling occurs prior to exon ligation.
- Wojciech P. Galej
- , Max E. Wilkinson
- & Kiyoshi Nagai
-
Article |
 Cryo-EM structure of the yeast U4/U6.U5 tri-snRNP at 3.7 Å resolution
Cryo-EM structure of the yeast U4/U6.U5 tri-snRNP at 3.7 Å resolution
A 3.7 Å resolution structure for the yeast U4/U6.U5 tri-snRNP, a complex involved in splicing, allows a better appreciation of the architecture of the tri-snRNP, and offers new functional insights into the activation of the spliceosome and the assembly of the catalytic core.
- Thi Hoang Duong Nguyen
- , Wojciech P. Galej
- & Kiyoshi Nagai
-
Letter |
 The spliceosome is a therapeutic vulnerability in MYC-driven cancer
The spliceosome is a therapeutic vulnerability in MYC-driven cancer
Splicing factors such as BUD31 are identified in a synthetic-lethal screen with cells overexpressing the transcription factor MYC; oncogenic MYC leads to an increase in pre-mRNA synthesis, and spliceosome inhibition impairs the growth and tumorigenicity of MYC-dependent breast cancers, suggesting that spliceosome components may be potential therapeutic targets for MYC-driven cancers.
- Tiffany Y.-T. Hsu
- , Lukas M. Simon
- & Thomas F. Westbrook
-
Article |
 The core spliceosome as target and effector of non-canonical ATM signalling
The core spliceosome as target and effector of non-canonical ATM signalling
Transcription-blocking DNA lesions result in chromatin displacement of core spliceosomes containing U2 and U5 snRNPs; consequently, R-loops containing the nascent transcript are formed, which activate ATM in a feed-forward fashion to influence spliceosome dynamics and alternative splicing.
- Maria Tresini
- , Daniël O. Warmerdam
- & Jurgen A. Marteijn
-
Article |
 The architecture of the spliceosomal U4/U6.U5 tri-snRNP
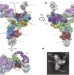
The architecture of the spliceosomal U4/U6.U5 tri-snRNP
This study determines the structure of the spliceosomal tri-snRNP complex (containing three small nuclear RNAs and more than 30 proteins) by single-particle cryo-electron microscopy; the resolution is sufficient to discern the organization of RNA and protein components involved in spliceosome activation, exon alignment and catalysis.
- Thi Hoang Duong Nguyen
- , Wojciech P. Galej
- & Kiyoshi Nagai
-
Letter |
 Recursive splicing in long vertebrate genes
Recursive splicing in long vertebrate genes
Highly conserved recursive splice sites are identified in vertebrates, particularly within long genes encoding proteins that are involved in neuronal development; analysis of the splicing mechanism reveals that such recursive splicing sites can be used to dictate different mRNA isoforms.
- Christopher R. Sibley
- , Warren Emmett
- & Jernej Ule
-
Letter |
 Genome-wide identification of zero nucleotide recursive splicing in Drosophila
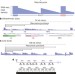
Genome-wide identification of zero nucleotide recursive splicing in Drosophila
In flies, some introns contain internal splice sites that cause ‘recursive splicing’, a multi-step removal of a single intron; this study demonstrates that the scope of this regulatory mechanism is much more extensive in flies than had been appreciated, and provides details about the recursive splicing process.
- Michael O. Duff
- , Sara Olson
- & Brenton R. Graveley
-
Letter |
 MYC regulates the core pre-mRNA splicing machinery as an essential step in lymphomagenesis
MYC regulates the core pre-mRNA splicing machinery as an essential step in lymphomagenesis
The critical effectors of MYC overexpression during lymphomagenesis in transgenic mice are defined.
- Cheryl M. Koh
- , Marco Bezzi
- & Ernesto Guccione
-
Article |
 Crystal structure of a eukaryotic group II intron lariat
Crystal structure of a eukaryotic group II intron lariat
This study determines the structure of a branched lariat RNA, providing insights into rearrangement of the intron between the two steps of RNA splicing.
- Aaron R. Robart
- , Russell T. Chan
- & Navtej Toor
-
Letter |
 Analysis of orthologous groups reveals archease and DDX1 as tRNA splicing factors
Analysis of orthologous groups reveals archease and DDX1 as tRNA splicing factors
Using a phylogenetic approach, the protein archease is identified as being a subunit of the human transfer RNA splicing ligase, and found to be necessary for full ligase activity, in cooperation with DDX1.
- Johannes Popow
- , Jennifer Jurkin
- & Javier Martinez
-
Letter |
 CFIm25 links alternative polyadenylation to glioblastoma tumour suppression
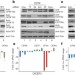
CFIm25 links alternative polyadenylation to glioblastoma tumour suppression
CFIm25 is identified as a factor that prevents messenger RNAs being shortened due to altered 3′ polyadenylation, which typically occurs when cells undergo high proliferation and correlates with increased tumorigenic activity in glioblastoma tumours.
- Chioniso P. Masamha
- , Zheng Xia
- & Eric J. Wagner
-
Article |
 RNA catalyses nuclear pre-mRNA splicing
RNA catalyses nuclear pre-mRNA splicing
The spliceosome is shown to catalyse splicing through the RNA and not the protein components of the spliceosome; pre-messenger RNA splicing requires U6 snRNA acting by a mechanism similar to that used by group II self-splicing introns.
- Sebastian M. Fica
- , Nicole Tuttle
- & Joseph A. Piccirilli
-
Letter |
 Identification of small RNA pathway genes using patterns of phylogenetic conservation and divergence
Identification of small RNA pathway genes using patterns of phylogenetic conservation and divergence
To identify comprehensively factors involved in RNAi and microRNA-mediated gene expression regulation, this study performed a phylogenetic analysis of 86 eukaryotic species; the candidates this approach highlighted were subjected to Bayesian analysis with transcriptional and proteomic interaction data, identifying protein orthologues of already known RNAi silencing factors, as well as other hits involved in splicing, suggesting a connection between the two processes.
- Yuval Tabach
- , Allison C. Billi
- & Gary Ruvkun
-
Article |
 Topology of the human and mouse m6A RNA methylomes revealed by m6A-seq
Topology of the human and mouse m6A RNA methylomes revealed by m6A-seq
N6-methyladenosine (m6A) is the most prevalent internal modification in messenger RNA; here the human and mouse m6A modification landscape is presented in a transcriptome-wide manner, providing insights into this epigenetic modification.
- Dan Dominissini
- , Sharon Moshitch-Moshkovitz
- & Gideon Rechavi
-
Letter |
 DBIRD complex integrates alternative mRNA splicing with RNA polymerase II transcript elongation
DBIRD complex integrates alternative mRNA splicing with RNA polymerase II transcript elongation
Characterization of the human interactome of chromatin-associated messenger ribonucleoprotein particles identifies DBC1 and a new protein (ZIRD) as subunits of a protein complex (DBIRD) that binds directly to RNAPII, regulates alternative splicing of exons embedded in (A + T)-rich DNA, and whose depletion results in region-specific decreases in transcript elongation.
- Pierre Close
- , Philip East
- & Jesper Q. Svejstrup
-
News & Views |
 A hidden ancestral legacy trumped
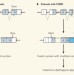
A hidden ancestral legacy trumped
A previously unsuspected genetic mechanism underlies a type of muscular dystrophy common in Japan. A therapeutic approach based on this finding and tested in mice has come up with encouraging results. See Letter p.127
- Masayuki Nakamori
- & Charles Thornton
-
Letter |
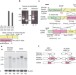 Pathogenic exon-trapping by SVA retrotransposon and rescue in Fukuyama muscular dystrophy
Pathogenic exon-trapping by SVA retrotransposon and rescue in Fukuyama muscular dystrophy
- Mariko Taniguchi-Ikeda
- , Kazuhiro Kobayashi
- & Tatsushi Toda
-
Article |
 Frequent pathway mutations of splicing machinery in myelodysplasia
Frequent pathway mutations of splicing machinery in myelodysplasia
- Kenichi Yoshida
- , Masashi Sanada
- & Seishi Ogawa
-
Letter |
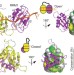 Multi-domain conformational selection underlies pre-mRNA splicing regulation by U2AF
Multi-domain conformational selection underlies pre-mRNA splicing regulation by U2AF
- Cameron D. Mackereth
- , Tobias Madl
- & Michael Sattler
-
Research Highlights |
 Epigenetics: What makes a queen bee?
Epigenetics: What makes a queen bee?
-
Article |
 U1 snRNP protects pre-mRNAs from premature cleavage and polyadenylation
U1 snRNP protects pre-mRNAs from premature cleavage and polyadenylation
Splicing is carried out by a collection of protein–RNA complexes known as snRNPs. The spliceosome contains equal quantities of the U1, U2, U4, U5 and U6 snRNPs, but the U1 snRNP is made in levels excess to the amounts needed to form spliceosomes, leading to the idea that excess U1s might have splicing independent functions. Here it is shown that the U1 snRNA interacts with some pre mRNAs whose introns have cryptic polyadenylation sites. This interaction prevents premature termination and polyadenylation of the pre mRNA.
- Daisuke Kaida
- , Michael G. Berg
- & Gideon Dreyfuss
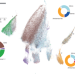
 Autonomous transposons tune their sequences to ensure somatic suppression
Autonomous transposons tune their sequences to ensure somatic suppression