Featured
-
-
Article |
 Structures and mechanisms of tRNA methylation by METTL1–WDR4
Structures and mechanisms of tRNA methylation by METTL1–WDR4
Using cryo-electron microscopy, structural and mechanistic insights into how the METTL1–WDR4 complex catalyses methylation of tRNAs are shown.
- Victor M. Ruiz-Arroyo
- , Rishi Raj
- & Yunsun Nam
-
Article
| Open Access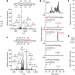 Reversible RNA phosphorylation stabilizes tRNA for cellular thermotolerance
Reversible RNA phosphorylation stabilizes tRNA for cellular thermotolerance
Reversible internal RNA phosphrylation contributes to thermal stability and nuclease resistance of tRNA, and cellular thermotolerance of hyperthermophiles.
- Takayuki Ohira
- , Keiichi Minowa
- & Tsutomu Suzuki
-
Article |
 N6-methyladenosine in poly(A) tails stabilize VSG transcripts
N6-methyladenosine in poly(A) tails stabilize VSG transcripts
N6-methyladenosine is enriched in poly(A) tails of VSG transcripts in Trypanosoma brucei, and when lacking result in mRNA degradation.
- Idálio J. Viegas
- , Juan Pereira de Macedo
- & Luisa M. Figueiredo
-
Article |
 Small-molecule inhibition of METTL3 as a strategy against myeloid leukaemia
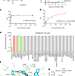
Small-molecule inhibition of METTL3 as a strategy against myeloid leukaemia
Treatment with a specific inhibitor of the N6-methyladenosine methyltransferase METTL3 leads to reduced growth of cancer cells, indicating the potential of approaches targeting RNA-modifying enzymes for anticancer therapy.
- Eliza Yankova
- , Wesley Blackaby
- & Tony Kouzarides
-
Article |
 METTL3 regulates heterochromatin in mouse embryonic stem cells
METTL3 regulates heterochromatin in mouse embryonic stem cells
Binding of METTL3 to chromatin is enriched over IAP family endogenous retroviral elements in mouse embryonic stem cells, helping to ensure the integrity of heterochromatin at these elements.
- Wenqi Xu
- , Jiahui Li
- & Hongjie Shen
-
Article |
 m6A RNA methylation regulates the fate of endogenous retroviruses
m6A RNA methylation regulates the fate of endogenous retroviruses
A CRISPR screen in mouse embryonic stem cells shows that transcripts derived from endogenous retroviruses are destabilized by m6A RNA methylation.
- Tomasz Chelmicki
- , Emeline Roger
- & Deborah Bourc’his
-
Article |
 Dynamic RNA acetylation revealed by quantitative cross-evolutionary mapping
Dynamic RNA acetylation revealed by quantitative cross-evolutionary mapping
A method termed ac4C-seq is introduced for the transcriptome-wide mapping of the RNA modification N4-acetylcytidine, revealing widespread temperature-dependent acetylation that facilitates thermoadaptation in hyperthermophilic archaea.
- Aldema Sas-Chen
- , Justin M. Thomas
- & Schraga Schwartz
-
Letter |
 FTSJ3 is an RNA 2′-O-methyltransferase recruited by HIV to avoid innate immune sensing
FTSJ3 is an RNA 2′-O-methyltransferase recruited by HIV to avoid innate immune sensing
HIV-1 uses the host protein FTSJ3 to methylate its own genome, thereby evading detection by the innate immune system.
- Mathieu Ringeard
- , Virginie Marchand
- & Yamina Bennasser
-
Letter |
 Mitochondrial translation requires folate-dependent tRNA methylation
Mitochondrial translation requires folate-dependent tRNA methylation
Mammalian mitochondria use folate-bound one-carbon units generated by the enzyme SHMT2 to methylate tRNA, and this modification is required for mitochondrial translation and thus oxidative phosphorylation.
- Raphael J. Morscher
- , Gregory S. Ducker
- & Joshua D. Rabinowitz
-
Article |
 Visualization of chemical modifications in the human 80S ribosome structure
Visualization of chemical modifications in the human 80S ribosome structure
A high-resolution structure of the human ribosome determined by cryo-electron microscopy visualizes numerous RNA modifications that are concentrated at functional sites with an extended shell, and suggests the possibility of designing more specific ribosome-targeting drugs.
- S. Kundhavai Natchiar
- , Alexander G. Myasnikov
- & Bruno P. Klaholz
-
Letter |
 The m1A landscape on cytosolic and mitochondrial mRNA at single-base resolution
The m1A landscape on cytosolic and mitochondrial mRNA at single-base resolution
Transcriptome-wide mapping of N1-methyladenosine (m1A) at single-nucleotide resolution reveals m1A to be scarce in cytoplasmic mRNA, to inhibit translation, and to be highly dynamic at a single site in a mitochondrial mRNA.
- Modi Safra
- , Aldema Sas-Chen
- & Schraga Schwartz
-
Letter |
 mRNA 3′ uridylation and poly(A) tail length sculpt the mammalian maternal transcriptome
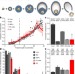
mRNA 3′ uridylation and poly(A) tail length sculpt the mammalian maternal transcriptome
TUT4 and TUT7 mediate 3′ uridylation of mRNA transcripts, preferentially those with short poly(A) tails; in the absence of TUT4 and TUT7, oocytes cannot mature and female mice are infertile.
- Marcos Morgan
- , Christian Much
- & Dónal O’Carroll
-
Letter |
 RNA m6A methylation regulates the ultraviolet-induced DNA damage response
RNA m6A methylation regulates the ultraviolet-induced DNA damage response
Methylation at the 6 position of adenosine (m6A) in RNA is rapidly and transiently induced at DNA damage sites in response to ultraviolet irradiation.
- Yang Xiang
- , Benoit Laurent
- & Yang Shi
-
Letter |
 Editing and methylation at a single site by functionally interdependent activities
Editing and methylation at a single site by functionally interdependent activities
The C-to-U deamination at position 32 of tRNAThr in Trypanosoma brucei requires two enzymatic activities and proceeds via formation of a 3-methylcytosine intermediate, supporting the notion of a coupled modification system.
- Mary Anne T. Rubio
- , Kirk W. Gaston
- & Juan D. Alfonzo
-
Article |
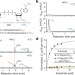 Reversible methylation of m6Am in the 5′ cap controls mRNA stability
Reversible methylation of m6Am in the 5′ cap controls mRNA stability
Fat mass and obesity-associated protein (FTO) preferentially demethylates m6Am, a modified adenosine that, when present at the 5′ end of certain mRNAs, positively influences mRNA stability by preventing DCP2-mediated decapping.
- Jan Mauer
- , Xiaobing Luo
- & Samie R. Jaffrey
-
Article |
 m6A modulates neuronal functions and sex determination in Drosophila
m6A modulates neuronal functions and sex determination in Drosophila
One of the most abundant modifications found in messenger RNAs is N6-methyladenosine (m6A); here, this modification is shown to alter gene expression during sex determination and affect neuronal functions and behaviour in Drosophila via the m6A reader protein YT521-B.
- Tina Lence
- , Junaid Akhtar
- & Jean-Yves Roignant
-
Letter |
 m6A potentiates Sxl alternative pre-mRNA splicing for robust Drosophila sex determination
m6A potentiates Sxl alternative pre-mRNA splicing for robust Drosophila sex determination
Two complementary studies describe how the pervasive N6-methyladenosine modification in mRNA can affect Drosophila sex determination, neuronal function and behaviour.
- Irmgard U. Haussmann
- , Zsuzsanna Bodi
- & Matthias Soller
-
Article |
 m6A RNA methylation promotes XIST-mediated transcriptional repression
m6A RNA methylation promotes XIST-mediated transcriptional repression
The methylation of adenosine residues on the long non-coding RNA XIST is essential for X-chromosome transcriptional repression during female mammalian development.
- Deepak P. Patil
- , Chun-Kan Chen
- & Samie R. Jaffrey
-
Letter |
 The mechanism of RNA 5′ capping with NAD+, NADH and desphospho-CoA
The mechanism of RNA 5′ capping with NAD+, NADH and desphospho-CoA
RNA caps other than the 7-methylguanylate modification are generated by a distinct mechanism in which caps are added during, not after, transcription initiation through the use of non-canonical initiating nucleotides by RNA polymerases, a finding which has functional consequences.
- Jeremy G. Bird
- , Yu Zhang
- & Bryce E. Nickels
-
Article |
 The dynamic N1-methyladenosine methylome in eukaryotic messenger RNA
The dynamic N1-methyladenosine methylome in eukaryotic messenger RNA
Here the m1A modification is discovered in messenger RNA and mapped at the transcriptome-wide level; the modification is conserved, dynamic, accumulates in structured regions around translation initiation sites upstream of the first splice site, and correlates with higher protein expression.
- Dan Dominissini
- , Sigrid Nachtergaele
- & Chuan He
-
Letter |
 N6-methyladenosine marks primary microRNAs for processing
N6-methyladenosine marks primary microRNAs for processing
The addition of the N6-methyladenosine (m6A) mark to primary microRNAs by METTL3 in mammalian cells is found to promote the recognition of these microRNA precursors by DGCR8, a component of the microprocessor complex.
- Claudio R. Alarcón
- , Hyeseung Lee
- & Sohail F. Tavazoie
-
Letter |
 N6-methyladenosine-dependent RNA structural switches regulate RNA–protein interactions
N6-methyladenosine-dependent RNA structural switches regulate RNA–protein interactions
The binding motifs for many RNA-binding proteins are normally buried within structured regions; now, the N6-methyladenosine modification is shown to act as a switch to remodel these regions, expose the motif, and thereby facilitate binding of RNA-binding proteins.
- Nian Liu
- , Qing Dai
- & Tao Pan
-
Letter |
 NAD captureSeq indicates NAD as a bacterial cap for a subset of regulatory RNAs
NAD captureSeq indicates NAD as a bacterial cap for a subset of regulatory RNAs
A newly developed method, NAD captureSeq, has been used to show that bacteria cap the 5′-ends of some RNAs to protect against degradation, much as happens with eukaryotic messenger RNAs, although with a different modification: nicotinamide adenine dinucleotide.
- Hana Cahová
- , Marie-Luise Winz
- & Andres Jäschke
-
Letter |
 Pseudouridine profiling reveals regulated mRNA pseudouridylation in yeast and human cells
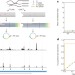
Pseudouridine profiling reveals regulated mRNA pseudouridylation in yeast and human cells
The modification of uridine to pseudouridine is widespread in transfer and ribosomal RNAs but not observed so far in a coding RNA; here a new technique is used to detect this modification on a genome-wide scale, leading to the identification of pseudouridylation in messenger RNAs as well as almost 100 new sites in non-coding RNAs.
- Thomas M. Carlile
- , Maria F. Rojas-Duran
- & Wendy V. Gilbert
-
Letter |
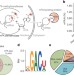 N6-methyladenosine-dependent regulation of messenger RNA stability
N6-methyladenosine-dependent regulation of messenger RNA stability
The mRNAs of higher eukaryotes are extensively modified internally with N6-methyladenosine, but the specific functional role of this modification has been unclear; here this modification on mRNA is shown to be recognized by several proteins, the modification and its recognition serve to regulate the RNA’s lifetime.
- Xiao Wang
- , Zhike Lu
- & Chuan He
-
Article |
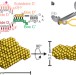 The structure of the box C/D enzyme reveals regulation of RNA methylation
The structure of the box C/D enzyme reveals regulation of RNA methylation
RNAs undergo many types of post-transcriptional modification, including methylation of ribosomal RNAs; here the structure of the archaeal box C/D ribonucleoprotein complex bound to substrate RNA is determined, showing that the two methylation guide sequences exist in different contexts and revealing sequential regulation of methylation at the two sites.
- Audrone Lapinaite
- , Bernd Simon
- & Teresa Carlomagno
-
Letter |
 Unusual base pairing during the decoding of a stop codon by the ribosome
Unusual base pairing during the decoding of a stop codon by the ribosome
Here, the structure of the 30S ribosomal subunit and the 70S ribosome in complex with a messenger RNA with pseudouridine in the place of uridine reveals unexpected base pairing.
- Israel S. Fernández
- , Chyan Leong Ng
- & V. Ramakrishnan
-
Letter |
 Structure-guided discovery of the metabolite carboxy-SAM that modulates tRNA function
Structure-guided discovery of the metabolite carboxy-SAM that modulates tRNA function
Members of the SAM-dependent methyltransferase superfamily are involved in the modification of wobble uridine to 5-oxacetyl uridine in Gram-negative bacteria; CmoA converts SAM to carboxy-SAM (Cx-SAM; a metabolite that was unknown previously), and CmoB uses Cx-SAM to convert 5-hydroxyuridine to 5-oxyacetyl uridine in tRNA.
- Jungwook Kim
- , Hui Xiao
- & Steven C. Almo
-
Article |
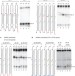 Dicer recognizes the 5′ end of RNA for efficient and accurate processing
Dicer recognizes the 5′ end of RNA for efficient and accurate processing
- Jong-Eun Park
- , Inha Heo
- & V. Narry Kim
-
Letter |
 Converting nonsense codons into sense codons by targeted pseudouridylation
Converting nonsense codons into sense codons by targeted pseudouridylation
- John Karijolich
- & Yi-Tao Yu
-
Letter |
 Identification of a quality-control mechanism for mRNA 5′-end capping
Identification of a quality-control mechanism for mRNA 5′-end capping
Following their synthesis, eukaryotic messenger RNAs have a 7-methylguanosine cap added to their 5′ ends to protect the mRNAs from degradation. Here it is shown that, in vitro and in yeast, caps lacking a methyl group are recognized by the Rai1 protein, which clips off the incomplete cap. The data provide evidence that Rai1 is part of a quality-control mechanism that monitors, and promotes the digestion of, aberrant mRNAs that might arise during stress conditions.
- Xinfu Jiao
- , Song Xiang
- & Megerditch Kiledjian

 mRNA ageing shapes the Cap2 methylome in mammalian mRNA
mRNA ageing shapes the Cap2 methylome in mammalian mRNA