Featured
-
-
Article
| Open Access N6-methyladenosine mRNA marking promotes selective translation of regulons required for human erythropoiesis
N6-methyladenosine mRNA marking promotes selective translation of regulons required for human erythropoiesis
Erythropoiesis can be regulated by transcriptional, epigenetic, and post-transcriptional mechanisms. Here the authors report that N6-methyladenosine mRNA methyltransferase complex stimulates erythropoiesis by promoting translation of specific mRNAs.
- Daniel A. Kuppers
- , Sonali Arora
- & Patrick J. Paddison
-
Article
| Open Access Dissecting splicing decisions and cell-to-cell variability with designed sequence libraries
Dissecting splicing decisions and cell-to-cell variability with designed sequence libraries
Alternative splicing is regulated by multiple mechanisms. Here the authors employed designed splice site libraries and massively parallel reporter assays to dissect the regulatory complexity and cell-to-cell variability of splicing decisions and to build accurate predictive models.
- Martin Mikl
- , Amit Hamburg
- & Eran Segal
-
Article
| Open Access HNRNPK maintains epidermal progenitor function through transcription of proliferation genes and degrading differentiation promoting mRNAs
HNRNPK maintains epidermal progenitor function through transcription of proliferation genes and degrading differentiation promoting mRNAs
Maintenance of high turnover in tissues such as epidermis requires balance between proliferation and differentiation. Here the authors show that HNRNPK promotes RNA Polymerase II binding to proliferation and self-renewal genes as well as degradation of differentiation promoting mRNAs together with DDX6 in epidermis.
- Jingting Li
- , Yifang Chen
- & George L. Sen
-
Article
| Open Access Accurate detection of m6A RNA modifications in native RNA sequences
Accurate detection of m6A RNA modifications in native RNA sequences
We currently lack generic methods to map RNA modifications across the entire transcriptome. Here, the authors demonstrate that m6A RNA modifications can be detected with high accuracy using nanopore direct RNA sequencing.
- Huanle Liu
- , Oguzhan Begik
- & Eva Maria Novoa
-
Article
| Open Access Defects in t6A tRNA modification due to GON7 and YRDC mutations lead to Galloway-Mowat syndrome
Defects in t6A tRNA modification due to GON7 and YRDC mutations lead to Galloway-Mowat syndrome
The biosynthesis of N6-threonylcarbamoylated adenosine 37 in tRNA (t6A) involves the YRDC enzyme and the KEOPS complex. Here, the authors report mutations in YRDC and the KEOPS component GON7 in Galloway-Mowat syndrome and determine the crystal structure of a GON7-containg subcomplex that suggests a role in KEOPS complex stability.
- Christelle Arrondel
- , Sophia Missoury
- & Géraldine Mollet
-
Article
| Open Access RST1 and RIPR connect the cytosolic RNA exosome to the Ski complex in Arabidopsis
RST1 and RIPR connect the cytosolic RNA exosome to the Ski complex in Arabidopsis
Cytosolic RNA degradation by the RNA exosome requires the Ski complex. Here the authors show that the proteins RST1 and RIPR assist the RNA exosome and the Ski complex in RNA degradation, thereby preventing the production of secondary siRNAs from endogenous mRNAs.
- Heike Lange
- , Simon Y. A. Ndecky
- & Dominique Gagliardi
-
Article
| Open Access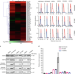 HSP90 inhibitors stimulate DNAJB4 protein expression through a mechanism involving N6-methyladenosine
HSP90 inhibitors stimulate DNAJB4 protein expression through a mechanism involving N6-methyladenosine
Cells respond to heat shock with transcriptional and translational adaptations but how HSP90 inhibition alters the heat shock proteome is largely unclear. Here, the authors analyze proteome changes upon HSP90 inhibition and show that an m6A-mediated mechanism contributes to the heat shock-induced upregulation of DNAJB4.
- Weili Miao
- , Lin Li
- & Yinsheng Wang
-
Article
| Open Access Modification of messenger RNA by 2′-O-methylation regulates gene expression in vivo
Modification of messenger RNA by 2′-O-methylation regulates gene expression in vivo
The post-transcriptional modification of mRNAs provides an additional layer to gene expression regulation. Here the authors show that 2′-O-methylation mediated by box C/D snoRNAs and fibrillarin can inhibit the translation of target mRNAs.
- Brittany A. Elliott
- , Hsiang-Ting Ho
- & Christopher L. Holley
-
Article
| Open Access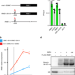 m6A modification of a 3′ UTR site reduces RME1 mRNA levels to promote meiosis
m6A modification of a 3′ UTR site reduces RME1 mRNA levels to promote meiosis
Ime4p is a yeast N6-methyladenosine (m6A) methyltransferase with an unknown role in meiosis. Rme1p is a repressor of meiosis. Here the authors show that Ime4p methylates RME1 3′ UTR to reduce its expression and enable meiosis, thus providing an example of an m6A site with a physiological role.
- G. Guy Bushkin
- , David Pincus
- & Gerald R. Fink
-
Article
| Open Access Time-resolved NMR monitoring of tRNA maturation
Time-resolved NMR monitoring of tRNA maturation
Transfer RNA (tRNA) is regulated by RNA modifications. Here the authors employ time-resolved NMR to monitor modifications of yeast tRNAPhe in cellular extracts, revealing a sequential order and cross-talk between modifications.
- Pierre Barraud
- , Alexandre Gato
- & Carine Tisné
-
Article
| Open Access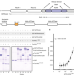 The MTR4 helicase recruits nuclear adaptors of the human RNA exosome using distinct arch-interacting motifs
The MTR4 helicase recruits nuclear adaptors of the human RNA exosome using distinct arch-interacting motifs
The human RNA exosome contains a nuclear co-factor MTR4, which unwinds structural RNAs and recruits adaptors for different RNA processing and decay pathways. Here the authors uncover new variations of the arch-interacting motif (AIM) in NVL and ZCCHC8 and characterise their interaction with MTR4.
- Mahesh Lingaraju
- , Dennis Johnsen
- & Elena Conti
-
Article
| Open Access Reconstitution of recombinant human CCR4-NOT reveals molecular insights into regulated deadenylation
Reconstitution of recombinant human CCR4-NOT reveals molecular insights into regulated deadenylation
The CCR4-NOT complex shortens poly(A) tails of messenger RNAs. By biochemical reconstitution of the entire human CCR4-NOT complex, the authors show the stimulatory roles of non-enzymatic subunits and the importance of the interaction between CAF40 and RNA binding proteins in targeted deadenylation.
- Tobias Raisch
- , Chung-Te Chang
- & Eugene Valkov
-
Article
| Open Access Live cell imaging reveals 3′-UTR dependent mRNA sorting to synapses
Live cell imaging reveals 3′-UTR dependent mRNA sorting to synapses
Asymmetric subcellular mRNA distribution is important for local translation of neuronal mRNAs. Here the authors employed MS2 live-cell imaging and showed that the reporter mRNA containing the 3’ UTR of Rgs4 shows an anterograde transport bias, dependent on neuronal activity and the protein Staufen2, and mediates sustained mRNA recruitment to synapses.
- Karl E. Bauer
- , Inmaculada Segura
- & Michael A. Kiebler
-
Article
| Open Access The mechanism of RNA duplex recognition and unwinding by DEAD-box helicase DDX3X
The mechanism of RNA duplex recognition and unwinding by DEAD-box helicase DDX3X
DEAD-box helicases (DDXs) function in an ATP-dependent, non-processive manner and the conserved helicase core is composed of two RecA-like domains D1 and D2. Here the authors present the crystal structure of the D1D2 core from human DDX3X bound to a 23-base pair dsRNA in the pre-unwound state and discuss the implications for helicase mechanism.
- He Song
- & Xinhua Ji
-
Article
| Open Access Dual pathways of tRNA hydroxylation ensure efficient translation by expanding decoding capability
Dual pathways of tRNA hydroxylation ensure efficient translation by expanding decoding capability
5-carboxymethoxyuridine (cmo5U) is one of the RNA modifications found in bacterial tRNA anticodons. Here the authors show that the first step of cmo5U biosynthesis from uridine is mediated by either one of two parallel factors, TrhP or TrhO, and that cmo5U modification is required for efficient translation.
- Yusuke Sakai
- , Satoshi Kimura
- & Tsutomu Suzuki
-
Article
| Open Access Programmable mutually exclusive alternative splicing for generating RNA and protein diversity
Programmable mutually exclusive alternative splicing for generating RNA and protein diversity
Alternative splicing expands the genetic coding capacity and proteomic diversity of the cell. Here the authors create a synthetic biology platform for regulating four programmable exons in modular transcription factors.
- Melina Mathur
- , Cameron M. Kim
- & Christina D. Smolke
-
Article
| Open Access The prion-like domain of Drosophila Imp promotes axonal transport of RNP granules in vivo
The prion-like domain of Drosophila Imp promotes axonal transport of RNP granules in vivo
The physiological role of prion-like domains (PLDs) within RNA-binding proteins is not well understood. Here, authors show in Drosophila that the PLD in the protein Imp is required for localization of ribonucleoprotein granules to axons and axonal remodelling.
- Jeshlee Vijayakumar
- , Charlène Perrois
- & Florence Besse
-
Article
| Open Access DHX36 prevents the accumulation of translationally inactive mRNAs with G4-structures in untranslated regions
DHX36 prevents the accumulation of translationally inactive mRNAs with G4-structures in untranslated regions
Translation efficiency can be affected by mRNA stability and secondary RNA structures. Here the authors reveal that loss of DHX36 helicase activity leads to an accumulation of translationally inactive target mRNAs with G-rich structures in untranslated regions.
- Markus Sauer
- , Stefan A. Juranek
- & Katrin Paeschke
-
Article
| Open Access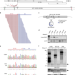 Transforming activity of an oncoprotein-encoding circular RNA from human papillomavirus
Transforming activity of an oncoprotein-encoding circular RNA from human papillomavirus
The authors identify circular RNAs (circRNA) from human papillomavirus and show that circRNA-encoded E7 contributes to cancer cell growth in vitro and in tumor xenografts. Furthermore, circE7 is present in TCGA RNA-Seq data from HPV-positive cancers.
- Jiawei Zhao
- , Eunice E. Lee
- & Richard C. Wang
-
Article
| Open Access Combinatorial recognition of clustered RNA elements by the multidomain RNA-binding protein IMP3
Combinatorial recognition of clustered RNA elements by the multidomain RNA-binding protein IMP3
Multidomain RNA-binding proteins recognize specific target sequences through mechanisms that are not well understood. Here the authors present an integrated approach to define the RNA-binding specificity and RNP topology and apply it to the analysis of the prototypical multidomain RNA-binding protein IMP3.
- Tim Schneider
- , Lee-Hsueh Hung
- & Albrecht Bindereif
-
Article
| Open Access EXOSC10 is required for RPA assembly and controlled DNA end resection at DNA double-strand breaks
EXOSC10 is required for RPA assembly and controlled DNA end resection at DNA double-strand breaks
The exosome is a ribonucleolytic complex that plays part in RNA processing and degradation. Here, the authors reveal an RNA clearance event in homologous recombination and show the function of EXOSC10, a component of the exosome, in the homeostasis of RNA molecules synthesized at DSB sites.
- Judit Domingo-Prim
- , Martin Endara-Coll
- & Neus Visa
-
Article
| Open Access LncRNA EPR controls epithelial proliferation by coordinating Cdkn1a transcription and mRNA decay response to TGF-β
LncRNA EPR controls epithelial proliferation by coordinating Cdkn1a transcription and mRNA decay response to TGF-β
Several lncRNAs are regulated by TGF-β. Here the authors report that an intergenic lncRNA —EPR— is a component of the TGF-β signaling pathway and controls epithelial cell proliferation by altering transcription and mRNA decay of Cdkn1a. EPR overexpression restrains tumor growth of orthotopically transplanted mice.
- Martina Rossi
- , Gabriele Bucci
- & Roberto Gherzi
-
Article
| Open Access Crystal structure of the Lin28-interacting module of human terminal uridylyltransferase that regulates let-7 expression
Crystal structure of the Lin28-interacting module of human terminal uridylyltransferase that regulates let-7 expression
Terminal uridylyltransferase 4/7 (TUT4/7) binds to Lin28 and modifies let-7 precursor (pre-let-7) to regulate cell differentiation and proliferation. Here the authors report the crystal structure of the N-terminal Lin28-interacting module of TUT4, showing a role of the N-terminal zinc finger domain in stabilizing the Lin28:pre-let-7:TUT4 ternary complex.
- Seisuke Yamashita
- , Takashi Nagaike
- & Kozo Tomita
-
Article
| Open Access Global identification of functional microRNA-mRNA interactions in Drosophila
Global identification of functional microRNA-mRNA interactions in Drosophila
MicroRNAs are mediators of post-transcriptional gene expression silencing. Here authors provide a transcriptome-wide map of miRNA target sites in Drosophila.
- Hans-Hermann Wessels
- , Svetlana Lebedeva
- & Uwe Ohler
-
Article
| Open Access RNA editing is abundant and correlates with task performance in a social bumblebee
RNA editing is abundant and correlates with task performance in a social bumblebee
Bumblebee workers are genetically highly similar but they show different behaviors such as brood care and foraging. Here the authors report a high level of ADAR-mediated RNA editing in the bumblebee Bombus terrestris and its weak correlation to task performance.
- Hagit T. Porath
- , Esther Hazan
- & Guy Bloch
-
Article
| Open Access Specific inhibition of splicing factor activity by decoy RNA oligonucleotides
Specific inhibition of splicing factor activity by decoy RNA oligonucleotides
Alternative splicing, critical for gene expression, is deregulated in many diseases. Here the authors develop decoy oligonucleotides to specifically downregulate splicing factors activity.
- Polina Denichenko
- , Maxim Mogilevsky
- & Rotem Karni
-
Article
| Open Access An alternative CTCF isoform antagonizes canonical CTCF occupancy and changes chromatin architecture to promote apoptosis
An alternative CTCF isoform antagonizes canonical CTCF occupancy and changes chromatin architecture to promote apoptosis
CTCF plays key roles in gene regulation, chromatin insulation and organizing the higher-order chromatin architecture of mammalian genomes. Here the authors investigate the function an alternatively spliced shorter CTCF isoform, finding that this isoform antagonizes canonical CTCF occupancy and changes chromatin architecture to promote apoptosis.
- Jiao Li
- , Kaimeng Huang
- & Hongjie Yao
-
Article
| Open Access Structure mapping of dengue and Zika viruses reveals functional long-range interactions
Structure mapping of dengue and Zika viruses reveals functional long-range interactions
Here, the authors provide detailed analyses of viral RNA structure in virions and in infected cells for four dengue virus serotypes and four Zika virus strains, and identify conserved structures that are important for virus replication.
- Roland G. Huber
- , Xin Ni Lim
- & Yue Wan
-
Article
| Open Access Molecular interactions between Hel2 and RNA supporting ribosome-associated quality control
Molecular interactions between Hel2 and RNA supporting ribosome-associated quality control
Ribosome-associated quality control (RQC) pathways monitor and respond to stalling of the translating ribosome. Here the authors show that the ribosome associated RQC factor Hel2/ZNF598, an E3 ubiquitin ligase, generally interacts with mRNAs in the vicinity of stop codons.
- Marie-Luise Winz
- , Lauri Peil
- & David Tollervey
-
Article
| Open Access Pentatricopeptide repeat poly(A) binding protein KPAF4 stabilizes mitochondrial mRNAs in Trypanosoma brucei
Pentatricopeptide repeat poly(A) binding protein KPAF4 stabilizes mitochondrial mRNAs in Trypanosoma brucei
Polyadenylation stabilizes edited mitochondrial mRNAs in Trypanosoma brucei, but the involved poly(A) binding protein is unknown. Here, Mesitov et al. show that a pentatricopeptide repeat factor KPAF4 binds to A-tail and prevents exonucleolytic degradation as well as translation of incompletely edited mRNAs.
- Mikhail V. Mesitov
- , Tian Yu
- & Inna Aphasizheva
-
Article
| Open Access The splicing factor RBM25 controls MYC activity in acute myeloid leukemia
The splicing factor RBM25 controls MYC activity in acute myeloid leukemia
Splicing factors are often mutated in hematological malignancies. Here, the authors perform an in vivo shRNA screen in a CEBPA mutant AML mouse model and identify that RBM25 controls the splicing of pre-mRNAs encoding BCL-X and BIN1 to exert its tumour suppressor activities in AML.
- Ying Ge
- , Mikkel Bruhn Schuster
- & Bo Torben Porse
-
Article
| Open Access The H/ACA complex disrupts triplex in hTR precursor to permit processing by RRP6 and PARN
The H/ACA complex disrupts triplex in hTR precursor to permit processing by RRP6 and PARN
Telomerase RNA (hTR) is transcribed as a 3′-extended precursor. Here the authors examine the processing of hTR precursors of various lengths and show that processing occurs in distinct steps involving different nucleases PARN and RRP6.
- Chi-Kang Tseng
- , Hui-Fang Wang
- & Peter Baumann
-
Article
| Open Access RNA helicases mediate structural transitions and compositional changes in pre-ribosomal complexes
RNA helicases mediate structural transitions and compositional changes in pre-ribosomal complexes
Pre-ribosomes undergo numerous structural rearrangements during their assembly. Here the authors identify the binding sites of three essential RNA helicases on pre-ribosomal particles, enabling them to provide insights into the structural and compositional changes that occur during biogenesis of the large ribosomal subunit.
- Lukas Brüning
- , Philipp Hackert
- & Markus T. Bohnsack
-
Article
| Open Access Non-invasive monitoring of alternative splicing outcomes to identify candidate therapies for myotonic dystrophy type 1
Non-invasive monitoring of alternative splicing outcomes to identify candidate therapies for myotonic dystrophy type 1
Myotonic dystrophy type 1 (DM1) is associated with aberrant transcript splicing. Here, the authors develop a transgenic mouse model expressing a bi-chromatic reporter system that allows non-invasive monitoring of splicing of a transcript altered in DM1 in vivo, and show that it allows for evaluation of the therapeutic response to treatment with antisense oligonucleotides.
- Ningyan Hu
- , Layal Antoury
- & Thurman M. Wheeler
-
Article
| Open Access Common mechanism of transcription termination at coding and noncoding RNA genes in fission yeast
Common mechanism of transcription termination at coding and noncoding RNA genes in fission yeast
Termination of RNA polymerase II (RNAPII) transcription is an essential step of gene expression. Here the authors provide evidence that in fission yeast termination of ncRNA genes occurs by a cleavage-dependent mechanism involving recruitment of mRNA 3′ end processing factors and requires the conserved Ysh1/CPSF-73 and Dhp1/XRN2 nucleases.
- Marc Larochelle
- , Marc-Antoine Robert
- & François Bachand
-
Article
| Open Access SHQ1 regulation of RNA splicing is required for T-lymphoblastic leukemia cell survival
SHQ1 regulation of RNA splicing is required for T-lymphoblastic leukemia cell survival
T-acute lymphoblastic leukemia is an aggressive cancer. Here the authors provide insights into the functional role of SHQ1, an H/ACA snoRNP assembly factor involved in snRNA pseudouridylation, in T-lymphoblastic leukemia cell survival through regulating the maturation of MYC mRNA.
- Hexiu Su
- , Juncheng Hu
- & Hudan Liu
-
Article
| Open Access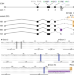 Genetic and mechanistic basis for APOBEC3H alternative splicing, retrovirus restriction, and counteraction by HIV-1 protease
Genetic and mechanistic basis for APOBEC3H alternative splicing, retrovirus restriction, and counteraction by HIV-1 protease
Human APOBEC3H has several haplotypes and splice variants with distinct anti-HIV-1 activities, but the genetics underlying the expression of these variants are unclear. Here, the authors identify an intronic deletion in A3H haplotype II resulting in production of the most active splice variant, which is counteracted by HIV-1 protease.
- Diako Ebrahimi
- , Christopher M. Richards
- & Reuben S. Harris
-
Article
| Open Access Regulatory mechanisms of incomplete huntingtin mRNA splicing
Regulatory mechanisms of incomplete huntingtin mRNA splicing
Incomplete splicing of HTT results in the production of the highly pathogenic exon 1 HTT protein. Here the authors identify the necessary intronic regions and the underlying mechanisms that contribute to this process.
- Andreas Neueder
- , Anaelle A. Dumas
- & Gillian P. Bates
-
Article
| Open Access RNA modification landscape of the human mitochondrial tRNALys regulates protein synthesis
RNA modification landscape of the human mitochondrial tRNALys regulates protein synthesis
Mutations in mitochondrially-encoded tRNA genes can lead to mitochondrial disorders. Here the authors use next generation RNA sequencing to reveal the role of a N1 -methyladenosine modification in tRNALys MERR patients for translation elongation and the stability of selected nascent chains.
- Uwe Richter
- , Molly E. Evans
- & Brendan J. Battersby
-
Article
| Open Access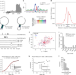 Roquin targets mRNAs in a 3′-UTR-specific manner by different modes of regulation
Roquin targets mRNAs in a 3′-UTR-specific manner by different modes of regulation
Roquin targets are known to contain two types of sequence-structure motifs, the constitutive and the alternative decay elements (CDE and ADE). Here, the authors describe a linear Roquin binding element (LBE) also involved in target recognition, and show that Roquin binding affects the translation of a subset of targeted mRNAs.
- Katharina Essig
- , Nina Kronbeck
- & Vigo Heissmeyer
-
Article
| Open Access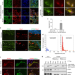 Staufen1 links RNA stress granules and autophagy in a model of neurodegeneration
Staufen1 links RNA stress granules and autophagy in a model of neurodegeneration
Spinocerebellar ataxia type 2 (SCA2) is caused by polyglutamine repeats in the ATXN2 protein. Here the authors demonstrate that Staufen1, known to be an RNA-binding protein, interacts with ATXN2 and contributes to pathology in a mouse model of SCA2.
- Sharan Paul
- , Warunee Dansithong
- & Stefan M. Pulst
-
Article
| Open Access CTD-dependent and -independent mechanisms govern co-transcriptional capping of Pol II transcripts
CTD-dependent and -independent mechanisms govern co-transcriptional capping of Pol II transcripts
The co-transcriptional capping of transcripts synthesized by RNA Pol II is substantially more efficient than capping of free RNA, a process that has been shown to depend on CTD phosphorylation. Here the authors demonstrate that a CTD-independent mechanism functions in parallel with CTD-dependent processes to ensure efficient capping.
- Melvin Noe Gonzalez
- , Shigeo Sato
- & Ronald C. Conaway
-
Article
| Open Access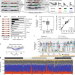 Decoding a cancer-relevant splicing decision in the RON proto-oncogene using high-throughput mutagenesis
Decoding a cancer-relevant splicing decision in the RON proto-oncogene using high-throughput mutagenesis
Alternative splicing is a critical step in eukaryotic gene expression but its molecular rules are not fully understood. Here, the authors develop a high-throughput mutagenesis approach to comprehensively characterise determinants of alternative splicing for the RON proto-oncogene.
- Simon Braun
- , Mihaela Enculescu
- & Kathi Zarnack
-
Article
| Open Access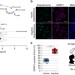 Dedicated surveillance mechanism controls G-quadruplex forming non-coding RNAs in human mitochondria
Dedicated surveillance mechanism controls G-quadruplex forming non-coding RNAs in human mitochondria
G-rich RNAs encoded in mitochondrial DNA are prone to form four-stranded structures called G-quadruplexes (G4s). Here the authors show using in vitro and in vivo approaches that GRSF1 promotes melting of G4 RNA structures in mtRNAs, thus leading to their decay by the hSuv3–PNPase complex.
- Zbigniew Pietras
- , Magdalena A. Wojcik
- & Roman J. Szczesny
-
Article
| Open Access Mutually exclusive acetylation and ubiquitylation of the splicing factor SRSF5 control tumor growth
Mutually exclusive acetylation and ubiquitylation of the splicing factor SRSF5 control tumor growth
Changes in glucose metabolism can lead to tumor development, but the involvement of splicing factors is unclear. Here, the authors screened for SR proteins and identified SRSF5 stability is enhanced in response to glucose elevation to promote alternative splicing of CCAR1 which facilitates tumor growth.
- Yuhan Chen
- , Qingyang Huang
- & Lingqiang Zhang
-
Article
| Open Access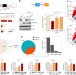 Nuclear PTEN safeguards pre-mRNA splicing to link Golgi apparatus for its tumor suppressive role
Nuclear PTEN safeguards pre-mRNA splicing to link Golgi apparatus for its tumor suppressive role
Cytoplasmic PTEN is a tumor suppressor that antagonises PI3K signalling. Here, the authors show that nuclear PTEN can interact with the spliceosomal proteins and drive pre-mRNA splicing in a phosphatase-independent manner, in particular, PTEN depletion promotes Golgi extension and secretion through GOLGA2 exon skipping.
- Shao-Ming Shen
- , Yan Ji
- & Guo-Qiang Chen
-
Article
| Open Access Co-regulatory activity of hnRNP K and NS1-BP in influenza and human mRNA splicing
Co-regulatory activity of hnRNP K and NS1-BP in influenza and human mRNA splicing
Alternative splicing of influenza A virus (IAV) M transcript is regulated by hnRNP K and NS1-BP, but mechanistic details are unknown. Here, Thompson et al. show how hnRNP K and NS1-BP bind M mRNA and that these proteins regulate splicing of host transcripts in both the absence and presence of IAV infection.
- Matthew G. Thompson
- , Raquel Muñoz-Moreno
- & Kristen W. Lynch
-
Article
| Open Access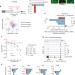 Cellular stress alters 3′UTR landscape through alternative polyadenylation and isoform-specific degradation
Cellular stress alters 3′UTR landscape through alternative polyadenylation and isoform-specific degradation
The function and consequences of alternative polyadenylation (APA) in stressed cells are largely unclear. Here, the authors show that stress-induced mRNA degradation depends on 3′UTR length and that APA-mediated 3′UTR shortening is an adaptive stress response mechanism for selective transcript stabilization.
- Dinghai Zheng
- , Ruijia Wang
- & Bin Tian
-
Article
| Open Access HuR regulates telomerase activity through TERC methylation
HuR regulates telomerase activity through TERC methylation
Mutations in the RNA component TERC can cause telomerase dysfunction but the underlying mechanisms are largely unknown. Here, the authors show that RNA-binding protein HuR regulates telomerase function by enhancing the methylation of TERC, which is impaired by several disease-relevant TERC mutations.
- Hao Tang
- , Hu Wang
- & Wengong Wang

 N6-methyladenosine modification of circNSUN2 facilitates cytoplasmic export and stabilizes HMGA2 to promote colorectal liver metastasis
N6-methyladenosine modification of circNSUN2 facilitates cytoplasmic export and stabilizes HMGA2 to promote colorectal liver metastasis