Featured
-
-
Article |
 RIC-seq for global in situ profiling of RNA–RNA spatial interactions
RIC-seq for global in situ profiling of RNA–RNA spatial interactions
RNA in situ conformation sequencing (RIC-seq) enables the generation of three-dimensional interaction maps of RNA in cells, which sheds light on the interactions and regulatory functions of RNA.
- Zhaokui Cai
- , Changchang Cao
- & Yuanchao Xue
-
Letter |
 Mitochondrial double-stranded RNA triggers antiviral signalling in humans
Mitochondrial double-stranded RNA triggers antiviral signalling in humans
Mitochondrial double-stranded RNA can induce an interferon response if released into the cytoplasm, but self-recognition is prevented by SUV3 helicase and PNPase exoribonuclease.
- Ashish Dhir
- , Somdutta Dhir
- & Nicholas J. Proudfoot
-
Letter |
 The helicase Ded1p controls use of near-cognate translation initiation codons in 5′ UTRs
The helicase Ded1p controls use of near-cognate translation initiation codons in 5′ UTRs
The helicase Ded1p associates with the pre-initiation complex and influences translation from near-cognate initiation codons by controlling the levels of mRNA secondary structure in 5′ untranslated regions.
- Ulf-Peter Guenther
- , David E. Weinberg
- & Eckhard Jankowsky
-
Letter |
 Structures of riboswitch RNA reaction states by mix-and-inject XFEL serial crystallography
Structures of riboswitch RNA reaction states by mix-and-inject XFEL serial crystallography
Femtosecond XFEL crystallography is used to identify dynamic changes in the adenine riboswitch aptamer domain, with at least four states identified in real time, two in the apo form before binding and two with the ligand bound.
- J. R. Stagno
- , Y. Liu
- & Y.-X. Wang
-
Article |
 Protein-guided RNA dynamics during early ribosome assembly
Protein-guided RNA dynamics during early ribosome assembly
Three-colour fluorescence resonance energy transfer and molecular dynamics simulations are used to study the events occurring early in assembly of the 30S ribosome; within a non-native intermediate S4 ribosomal protein–16S RNA structure, S4 is capable of altering the RNA helix dynamics to facilitate conformation changes that enable subsequent protein binding.
- Hajin Kim
- , Sanjaya C. Abeysirigunawarden
- & Sarah A. Woodson
-
Letter |
 In vivo genome-wide profiling of RNA secondary structure reveals novel regulatory features
In vivo genome-wide profiling of RNA secondary structure reveals novel regulatory features
RNA adopts vast and complicated secondary structures in the living cell; reported here is a new approach, structure-seq, that characterises RNA structure in vivo on a genome-wide scale at nucleotide resolution and should be applicable to any organism.
- Yiliang Ding
- , Yin Tang
- & Sarah M. Assmann
-
Letter |
 Three-state mechanism couples ligand and temperature sensing in riboswitches
Three-state mechanism couples ligand and temperature sensing in riboswitches
In the human pathogen Vibrio vulnificus, the riboswitch regulating gene expression of the adenosine deaminase is shown to exist in three distinct stable conformational states; this three-state mechanism allows control of gene expression over a broad temperature range, which is essential for Vibrio adaptation.
- Anke Reining
- , Senada Nozinovic
- & Harald Schwalbe
-
Article |
 Single-molecule analysis of Mss116-mediated group II intron folding
Single-molecule analysis of Mss116-mediated group II intron folding
DEAD-box helicases use ATP hydrolysis to unwind duplex RNA and facilitate RNA or RNA–protein remodelling. One such helicase is Mss116, which targets a particular group II intron in RNA. Here, single-molecule fluorescence was used to monitor the effect of Mss16 on a minimal construct containing this intron. The data show that Mss16 stimulates the sampling of different folded states of the RNA. Moreover, the helicase promotes RNA folding through discrete ATP-independent and ATP-dependent steps.
- Krishanthi S. Karunatilaka
- , Amanda Solem
- & David Rueda
-
Letter |
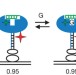 Multiple native states reveal persistent ruggedness of an RNA folding landscape
Multiple native states reveal persistent ruggedness of an RNA folding landscape
The 'thermodynamic hypothesis' proposes that the sequence of a biological macromolecule defines its folded, active structure as a global energy minimum in the folding landscape; however, it is not clear whether there is only one global minimum or several local minima corresponding to active conformations. Here, using single-molecule experiments, an RNA enzyme is shown to fold into multiple distinct native states that interconvert.
- Sergey V. Solomatin
- , Max Greenfeld
- & Daniel Herschlag

 Cryo-EM structures of full-length Tetrahymena ribozyme at 3.1 Å resolution
Cryo-EM structures of full-length Tetrahymena ribozyme at 3.1 Å resolution