Featured
-
-
Article
| Open Access Ataluren binds to multiple protein synthesis apparatus sites and competitively inhibits release factor-dependent termination
Ataluren binds to multiple protein synthesis apparatus sites and competitively inhibits release factor-dependent termination
Ataluren is the only nonsense suppressor drug currently approved for clinical use. Here, the authors determine where ataluren binds to the ribosome and how it inhibits termination at nonsense codons.
- Shijie Huang
- , Arpan Bhattacharya
- & Barry S. Cooperman
-
Article
| Open Access Structural basis for PoxtA-mediated resistance to phenicol and oxazolidinone antibiotics
Structural basis for PoxtA-mediated resistance to phenicol and oxazolidinone antibiotics
PoxtA confers resistance to ribosome-targeting oxazolidinone (linezolid) and chloramphenicol antibiotics. Here, Crowe-McAuliffe et al. provide structural insights into how binding of PoxtA to the ribosome indirectly promotes drug dissociation.
- Caillan Crowe-McAuliffe
- , Victoriia Murina
- & Vasili Hauryliuk
-
Article
| Open Access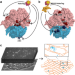 Direct measurements of mRNA translation kinetics in living cells
Direct measurements of mRNA translation kinetics in living cells
Metelev et al. use single-molecule tracking to study kinetics of translation directly in E. coli cells, and how it is affected by translation inhibitors and rRNA mutations. Their results support widespread 70S re-initiation on mRNAs.
- Mikhail Metelev
- , Erik Lundin
- & Magnus Johansson
-
Article
| Open Access eIF6 rebinding dynamically couples ribosome maturation and translation
eIF6 rebinding dynamically couples ribosome maturation and translation
Jaako et al. discover a conserved tier of translational control that dynamically couples ribosome assembly and recycling. This mechanism is corrupted in an inherited bone marrow failure disorder associated with an increased risk of blood cancer.
- Pekka Jaako
- , Alexandre Faille
- & Alan J. Warren
-
Article
| Open Access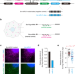 Single-molecule imaging of microRNA-mediated gene silencing in cells
Single-molecule imaging of microRNA-mediated gene silencing in cells
For decades, miRNAs have been studied primarily by ensemble methods, where a bulk collection of molecules is measured outside cells. Here, Kobayashi and Singer report methods to image miRNA function at the single-molecule level inside cells.
- Hotaka Kobayashi
- & Robert H. Singer
-
Article
| Open Access A late-stage assembly checkpoint of the human mitochondrial ribosome large subunit
A late-stage assembly checkpoint of the human mitochondrial ribosome large subunit
Rebelo-Guiomar et al. unveil late stage assembly intermediates of the human mitochondrial ribosome by inactivating the methyltransferase MRM2 in cells. Absence of MRM2 impairs organismal homeostasis, while its catalytic activity is dispensable for mitoribosomal biogenesis.
- Pedro Rebelo-Guiomar
- , Simone Pellegrino
- & Michal Minczuk
-
Article
| Open Access Mutations in Hcfc1 and Ronin result in an inborn error of cobalamin metabolism and ribosomopathy
Mutations in Hcfc1 and Ronin result in an inborn error of cobalamin metabolism and ribosomopathy
Combined methylmalonic acidemia (MMA) and hyperhomocysteinemias are inborn errors of vitamin B12 metabolism, and mutations in the transcriptional regulators HCFC1 and RONIN (THAP11) underlie some forms of these disorders. Here the authors generated mouse models of a human syndrome due to mutations in RONIN (THAP11) and HCFC1, and show that this syndrome is both an inborn error of vitamin B12 metabolism and displays some features of ribosomopathy.
- Tiffany Chern
- , Annita Achilleos
- & Ross A. Poché
-
Article
| Open Access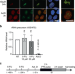 Alteration of ribosome function upon 5-fluorouracil treatment favors cancer cell drug-tolerance
Alteration of ribosome function upon 5-fluorouracil treatment favors cancer cell drug-tolerance
Different mechanisms have been reported to explain resistance to chemotherapy in cancer. Here, the authors show that the chemotherapeutic drug 5-fluorouracil alters the function of ribosomes to promote pro-survival gene translation leading to chemotherapy resistance.
- Gabriel Therizols
- , Zeina Bash-Imam
- & Jean-Jacques Diaz
-
Article
| Open Access Targeted editing and evolution of engineered ribosomes in vivo by filtered editing
Targeted editing and evolution of engineered ribosomes in vivo by filtered editing
Genome editing methods are limited by the inability to selectively edit repetitive sequences. Here the authors demonstrate precise editing of a repetitive genetic element, a ribosome, while avoiding edits to native sites sharing identical sequence.
- Felix Radford
- , Shane D. Elliott
- & Farren J. Isaacs
-
Article
| Open Access Time-resolved cryo-EM visualizes ribosomal translocation with EF-G and GTP
Time-resolved cryo-EM visualizes ribosomal translocation with EF-G and GTP
EF-G drives ribosomal translocation along mRNA. Time-resolved cryo-EM captured translocation with EF-G•GTP—without inhibitors—revealing how EF-G uses ribosome fluctuations to drive translocation and GTP hydrolysis to leave at the right moment.
- Christine E. Carbone
- , Anna B. Loveland
- & Andrei A. Korostelev
-
Article
| Open Access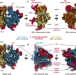 How to build a ribosome from RNA fragments in Chlamydomonas mitochondria
How to build a ribosome from RNA fragments in Chlamydomonas mitochondria
Mitoribosomes are remarkably diverse in their structures and compositions. Here the authors combine biochemistry, genetics, single particle cryo-electron microscopy and in situ cryo-electron tomography to reveal the mitochondrial ribosome of Chlamydomonas reinhardtii as an extreme example of evolution and species-specific adaptation.
- Florent Waltz
- , Thalia Salinas-Giegé
- & Yaser Hashem
-
Article
| Open Access Structural and molecular basis for Cardiovirus 2A protein as a viral gene expression switch
Structural and molecular basis for Cardiovirus 2A protein as a viral gene expression switch
Many RNA viruses employ programmed –1 ribosomal frameshifting (PRF) to expand their coding capacity and optimize production of viral proteins. Here, the authors report structural and biophysical analysis of protein 2A from a cardiovirus, with insights into the mechanism of its PRF-stimulatory function.
- Chris H. Hill
- , Lukas Pekarek
- & Ian Brierley
-
Article
| Open Access Functionally distinct roles for eEF2K in the control of ribosome availability and p-body abundance
Functionally distinct roles for eEF2K in the control of ribosome availability and p-body abundance
Processing bodies are phase separated compartments enriched in translationally repressed mRNAs. Here, Smith et al. show that, in sensory neurons, eukaryotic elongation factor 2 kinase (eEF2K) plays key roles in the regulation of processing body abundance and the formation of translationally inactive ribosomes.
- Patrick R. Smith
- , Sarah Loerch
- & Zachary T. Campbell
-
Article
| Open Access Bi-directional ribosome scanning controls the stringency of start codon selection
Bi-directional ribosome scanning controls the stringency of start codon selection
Start codon selection is commonly thought to occur through the unidirectional scanning of the mRNA by the 40 S ribosome. Here the authors provide evidence that the pre-initiation complex can backslide on the mRNA to initiate translation at upstream AUG codons.
- Yifei Gu
- , Yuanhui Mao
- & Shu-Bing Qian
-
Article
| Open Access Nascent chains can form co-translational folding intermediates that promote post-translational folding outcomes in a disease-causing protein
Nascent chains can form co-translational folding intermediates that promote post-translational folding outcomes in a disease-causing protein
Alpha-1-antitrypsin (AAT) deficiency results from misfolding-prone AAT variants. Here the authors show that AAT forms co-translational folding intermediates on the ribosome that persist upon release and determine its folding fate. They show too that the ribosome can also modulate misfolding-prone AAT intermediates during their synthesis.
- Elena Plessa
- , Lien P. Chu
- & Lisa D. Cabrita
-
Article
| Open Access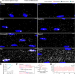 Microtubule-based transport is essential to distribute RNA and nascent protein in skeletal muscle
Microtubule-based transport is essential to distribute RNA and nascent protein in skeletal muscle
It is increasingly recognised that the spatial localisation of RNA is important for proper cellular function. Here, the authors investigate RNA localisation in skeletal muscle and develop methods to show that global active transport of RNA is required to maintain dispersion of gene products in the large muscle syncytium.
- Lance T. Denes
- , Chase P. Kelley
- & Eric T. Wang
-
Article
| Open Access Structural mechanism of GTPase-powered ribosome-tRNA movement
Structural mechanism of GTPase-powered ribosome-tRNA movement
Movement of the ribosome along an mRNA requires the universally-conserved translocase (EF-G in bacteria) that couples GTP hydrolysis to directed movement. Here the authors use time-resolved Cryo-EM to visualize the GTPase-powered step on native translocating ribosomes and capture key translocation intermediates.
- Valentyn Petrychenko
- , Bee-Zen Peng
- & Niels Fischer
-
Article
| Open Access Pathway of Hsp70 interactions at the ribosome
Pathway of Hsp70 interactions at the ribosome
Here, the authors use in vivo site-specific crosslinking to provide molecular-level insight into how the fungal Hsp70 chaperone system — the Ssb:Ssz1:Zuo1 triad — assists the folding process for the nascent peptide chain emerging from the ribosome tunnel.
- Kanghyun Lee
- , Thomas Ziegelhoffer
- & Elizabeth A. Craig
-
Article
| Open Access Directed evolution of rRNA improves translation kinetics and recombinant protein yield
Directed evolution of rRNA improves translation kinetics and recombinant protein yield
Ribosome kinetics are rate-limiting for protein synthesis. Here the authors evolve diverse 16S rRNAs for enhanced protein synthesis rates and genetic code expansion efficiencies in vivo.
- Fan Liu
- , Siniša Bratulić
- & Ahmed H. Badran
-
Article
| Open Access Structural basis for the tryptophan sensitivity of TnaC-mediated ribosome stalling
Structural basis for the tryptophan sensitivity of TnaC-mediated ribosome stalling
Bacteria adjust the expression of some of their metabolic enzymes through metabolite-sensing ribosome nascent chain complexes. Here the authors present a cryo-EM structure of an E. coli ribosome stalled during translation of the TnaC leader peptide and propose a model for L-Trp dependent ribosome stalling where L-Trp competes with release factor 2 for binding to the TnaC-ribosome complex.
- Anne-Xander van der Stel
- , Emily R. Gordon
- & C. Axel Innis
-
Article
| Open Access Humans and other commonly used model organisms are resistant to cycloheximide-mediated biases in ribosome profiling experiments
Humans and other commonly used model organisms are resistant to cycloheximide-mediated biases in ribosome profiling experiments
Ribosome profiling has become the gold standard to analyze mRNA translation dynamics, and the translation inhibitor cycloheximide (CHX) is often used in its application. Here the authors systematically demonstrate that CHX does not bias the outcome of ribosome profiling experiments in most organisms.
- Puneet Sharma
- , Jie Wu
- & Sebastian A. Leidel
-
Article
| Open Access Somatic genetic rescue of a germline ribosome assembly defect
Somatic genetic rescue of a germline ribosome assembly defect
Shwachman-Diamond syndrome (SDS) is a leukemia predisposition disorder that is caused by defective release of eIF6 during ribosome assembly. Here the authors show that acquired somatic EIF6 mutations are frequent in the hematopoietic cells from individuals with SDS and provide a selective advantage over non-modified cells.
- Shengjiang Tan
- , Laëtitia Kermasson
- & Patrick Revy
-
Article
| Open Access Structures of tmRNA and SmpB as they transit through the ribosome
Structures of tmRNA and SmpB as they transit through the ribosome
Trans-translation, mediated by small protein B (SmpB) and transfer-messenger RNA (tmRNA), enables recycling of the ribosomes stalled on defective mRNAs in bacteria. Here, the authors report structures of the ribosome during trans-translation that reveal a translocation intermediate and elucidate the movements of the tmRNA-SmpB complex in the ribosome.
- Charlotte Guyomar
- , Gaetano D’Urso
- & Reynald Gillet
-
Article
| Open Access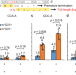 Structural basis for +1 ribosomal frameshifting during EF-G-catalyzed translocation
Structural basis for +1 ribosomal frameshifting during EF-G-catalyzed translocation
Translational frameshifting is a mechanism that expands the coding capabilities of mRNA. Here, structures of 70S ribosome complexes with GTPase elongation factor G (EF-G), a +1-frameshifting-prone mRNA and tRNAs reveal the cooperation between the ribosome and EF-G to induce +1 frameshifting during the translocation step.
- Gabriel Demo
- , Howard B. Gamper
- & Andrei A. Korostelev
-
Article
| Open Access A distinct assembly pathway of the human 39S late pre-mitoribosome
A distinct assembly pathway of the human 39S late pre-mitoribosome
Assembly of the mitoribosome requires assistance from numerous specialized factors. Here, structures of the human 39S late assembly intermediates identify several assembly factors which keep the 16S rRNA in immature conformations, and reveal deacylated tRNA in the ribosomal E-site, suggesting a role in 39S assembly.
- Jingdong Cheng
- , Otto Berninghausen
- & Roland Beckmann
-
Article
| Open Access Structural and mechanistic basis for translation inhibition by macrolide and ketolide antibiotics
Structural and mechanistic basis for translation inhibition by macrolide and ketolide antibiotics
Macrolides and ketolides antibiotics selectively interfere with the translation of a specific subset of proteins. Here the authors show how the macrolide erythromycin and the ketolide telithromycin interplay with the nascent polypeptide chain to arrest translation.
- Bertrand Beckert
- , Elodie C. Leroy
- & Daniel N. Wilson
-
Article
| Open Access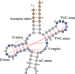 Repurposing tRNAs for nonsense suppression
Repurposing tRNAs for nonsense suppression
Here, the authors report de novo design, optimization and characterization of tRNAs that decode UGA stop codons in E. coli. The structure of the ribosome in a complex with the designed tRNA bound to a UGA stop codon suggests that distinct A-site ligands (tRNAs versus release factors) induce distinct conformation of the stop codon within the mRNA in the decoding center.
- Suki Albers
- , Bertrand Beckert
- & Zoya Ignatova
-
Article
| Open Access Structural basis for late maturation steps of the human mitoribosomal large subunit
Structural basis for late maturation steps of the human mitoribosomal large subunit
Mitochondrial ribosomes (mitoribosomes) are characterized by a distinct architecture and thus biogenesis pathway. Here, cryo-EM structures of mitoribosome large subunit assembly intermediates elucidate final steps of 16 S rRNA folding, methylation and peptidyl transferase centre (PTC) completion, as well as functions of several mitoribosome assembly factors.
- Miriam Cipullo
- , Genís Valentín Gesé
- & Joanna Rorbach
-
Article
| Open Access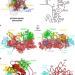 Structural basis of GTPase-mediated mitochondrial ribosome biogenesis and recycling
Structural basis of GTPase-mediated mitochondrial ribosome biogenesis and recycling
Maturation of the ribosomal peptidyl transferase center (PTC) is mediated by universally conserved GTPases. Here, cryo-EM structures of mitochondrial ribosomal large subunit assembly intermediates and of mature ribosomes offer insight into the roles of several assembly factors, including GTPBP6’s role in both ribosome biogenesis and recycling.
- Hauke S. Hillen
- , Elena Lavdovskaia
- & Ricarda Richter-Dennerlein
-
Article
| Open Access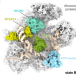 Stepwise maturation of the peptidyl transferase region of human mitoribosomes
Stepwise maturation of the peptidyl transferase region of human mitoribosomes
Mammalian mitoribosomes feature dramatically reduced ribosomal RNAs and follow mitochondria specific assembly pathways. Here the authors describe the process of human mitochondrial ribosome maturation that results in the formation of the ribosomal active site region, including the peptidyl transferase loop and the two tRNA-binding loops.
- Tea Lenarčič
- , Mateusz Jaskolowski
- & Nenad Ban
-
Article
| Open Access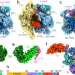 Distinct mechanisms of the human mitoribosome recycling and antibiotic resistance
Distinct mechanisms of the human mitoribosome recycling and antibiotic resistance
High-resolution cryo-EM structures and biochemical analyses of the human mitoribosome, in complex with mitochondria-specific factors mediating mitoribosome recycling, RRFmt and EF-G2mt, offer insight into mechanisms of mitoribosome recycling and resistance to antibiotic fusidic acid.
- Ravi Kiran Koripella
- , Ayush Deep
- & Rajendra K. Agrawal
-
Article
| Open Access Structural basis of ABCF-mediated resistance to pleuromutilin, lincosamide, and streptogramin A antibiotics in Gram-positive pathogens
Structural basis of ABCF-mediated resistance to pleuromutilin, lincosamide, and streptogramin A antibiotics in Gram-positive pathogens
Mitochondrial ribosomes (mitoribosomes) are characterized by a distinct architecture and thus biogenesis pathway. Here, cryo-EM structures of mitoribosome large subunit assembly intermediates elucidate final steps of 16 S rRNA folding, methylation and peptidyl transferase centre (PTC) completion, as well as functions of several mitoribosome assembly factors.
- Caillan Crowe-McAuliffe
- , Victoriia Murina
- & Daniel N. Wilson
-
Article
| Open Access Translation and codon usage regulate Argonaute slicer activity to trigger small RNA biogenesis
Translation and codon usage regulate Argonaute slicer activity to trigger small RNA biogenesis
22G-RNAs are single-stranded antisense small RNAs that are expressed in C. elegans germline. Here the authors show that CSR-1 dependent 22G-RNAs are produced in the cytosol on mRNAs actively engaged in translation and that codon usage of an mRNA regulates the biogenesis of CSR-1 dependent 22G-RNAs.
- Meetali Singh
- , Eric Cornes
- & Germano Cecere
-
Article
| Open Access 40S ribosome profiling reveals distinct roles for Tma20/Tma22 (MCT-1/DENR) and Tma64 (eIF2D) in 40S subunit recycling
40S ribosome profiling reveals distinct roles for Tma20/Tma22 (MCT-1/DENR) and Tma64 (eIF2D) in 40S subunit recycling
Following the termination of translation at stop codons, the eukaryotic 60S subunit of the ribosome is removed by the ATPase ABCE1. Here using 40S ribosome footprinting the authors provide a direct demonstration that the yeast orthologs of eIF2D, MCT-1, and DENR recycle the 40S subunits.
- David J. Young
- , Sezen Meydan
- & Nicholas R. Guydosh
-
Article
| Open Access Context-specific action of macrolide antibiotics on the eukaryotic ribosome
Context-specific action of macrolide antibiotics on the eukaryotic ribosome
Macrolide antibiotics inhibit bacterial translation in a context-specific manner, arresting ribosomes at defined sites within mRNAs and selectively inhibiting synthesis of only a subset of cellular proteins. Here the authors provide a structural basis for the context-specific activity of macrolides on the eukaryotic ribosome.
- Maxim S. Svetlov
- , Timm O. Koller
- & Alexander S. Mankin
-
Article
| Open Access Translation error clusters induced by aminoglycoside antibiotics
Translation error clusters induced by aminoglycoside antibiotics
Aminoglycoside antibiotics target the ribosome and induce misreading, yet which translation errors induce bacterial cell death is unclear. Here authors use quantitative mass spectrometry and show that bactericidal aminoglycosides induce clusters of errors in full-length proteins in vivo with as many as four amino acid substitutions in a row.
- Ingo Wohlgemuth
- , Raffaella Garofalo
- & Marina V. Rodnina
-
Article
| Open Access trans-Translation inhibitors bind to a novel site on the ribosome and clear Neisseria gonorrhoeae in vivo
trans-Translation inhibitors bind to a novel site on the ribosome and clear Neisseria gonorrhoeae in vivo
Antibiotic-resistant bacterial pathogens pose a substantial threat to human health. Here, aided by structural analyses, the authors describe the molecular mechanism behind the activity of a series of compounds that inhibit trans-translation and are effective in eradicating N. gonorrhoeae infection in mice.
- Zachary D. Aron
- , Atousa Mehrani
- & Kenneth C. Keiler
-
Article
| Open Access A steric gate controls P/E hybrid-state formation of tRNA on the ribosome
A steric gate controls P/E hybrid-state formation of tRNA on the ribosome
The ribosome undergoes multiple large-scale structural rearrangements during protein elongation. Here the authors present an all-atom model of the ribosome to study the energetics of P/E hybrid-state formation, an early conformational rearrangement occurring during translocation.
- Mariana Levi
- , Kelsey Walak
- & Paul C. Whitford
-
Article
| Open Access ArfB can displace mRNA to rescue stalled ribosomes
ArfB can displace mRNA to rescue stalled ribosomes
Alternative rescue factor B (ArfB) is an enzyme that releases peptides from stalled ribosomes to allow ribosome recycling. Here the authors carry-out cryo-EM analyses of 70S ribosomes complexed with ArfB on either a short or longer mRNA to reveal distinct modes of ArfB function.
- Christine E. Carbone
- , Gabriel Demo
- & Andrei A. Korostelev
-
Article
| Open Access Structural insights into assembly of the ribosomal nascent polypeptide exit tunnel
Structural insights into assembly of the ribosomal nascent polypeptide exit tunnel
The nascent polypeptide exit tunnel (NPET) is a functional center of the large ribosomal subunit through which the nascent polypeptide chains travel from the peptidyltransferase center (PTC). Here the authors provide structural insight into NPET maturation and how it is linked to other aspects of ribosome biogenesis.
- Daniel M. Wilson
- , Yu Li
- & John L. Woolford Jr
-
Article
| Open Access Structural basis for the transition from translation initiation to elongation by an 80S-eIF5B complex
Structural basis for the transition from translation initiation to elongation by an 80S-eIF5B complex
The GTPase eIF5B is involved in the correct positioning of the initiator Met-tRNAiMet on the ribosome during the late stages of translation initiation. Here the authors present a cryo-EM structure of the ribosome in complex with eIF5B and Met-tRNAiMet immediately before transition into elongation, providing insight on Met-tRNAiMetdelivery and how the correct reading frame is established.
- Jinfan Wang
- , Jing Wang
- & Israel S. Fernández
-
Article
| Open Access Selective inhibition of human translation termination by a drug-like compound
Selective inhibition of human translation termination by a drug-like compound
The drug-like compound PF846 and its derivatives inhibit the translation of specific mRNAs by the human ribosome. Here the authors show how PF846 arrests translation at the stop codon by slowing hydrolysis of the protein nascent chain at the ribosome P-site tRNA by eukaryotic release factor 1 (eRF1).
- Wenfei Li
- , Stacey Tsai-Lan Chang
- & Jamie H. D. Cate
-
Article
| Open Access A possible universal role for mRNA secondary structure in bacterial translation revealed using a synthetic operon
A possible universal role for mRNA secondary structure in bacterial translation revealed using a synthetic operon
The mechanisms for regulating translation re-initiation in bacteria remain poorly understood. Here, the authors screened a library of synthetic operons and identified a ribosome termination structure that modulates re-initiation efficiency and which is conserved across bacteria.
- Yonatan Chemla
- , Michael Peeri
- & Lital Alfonta
-
Article
| Open Access Mechanism of ribosome rescue by alternative ribosome-rescue factor B
Mechanism of ribosome rescue by alternative ribosome-rescue factor B
Rescue of ribosomes stalled on non-stop mRNA is essential for cell viability, and several rescue systems to resolve stalling exist in bacteria. Here, the authors use rapid kinetics and cryo-EM to reveal the pathway and selectivity mechanism of ArfB-mediated ribosome rescue.
- Kai-Hsin Chan
- , Valentyn Petrychenko
- & Marina V. Rodnina
-
Article
| Open Access Structures of the human mitochondrial ribosome bound to EF-G1 reveal distinct features of mitochondrial translation elongation
Structures of the human mitochondrial ribosome bound to EF-G1 reveal distinct features of mitochondrial translation elongation
Translation within mitochondria is carried out by specialized mitoribosomes and translational factors. Here the authors describe cryo-EM structures of the human mitochondrial translation elongation factor G1 in complex with human mitoribosomes, revealing distinct mechanism that include conformational changes at the polypeptide exit tunnel.
- Ravi Kiran Koripella
- , Manjuli R. Sharma
- & Rajendra K. Agrawal
-
Article
| Open Access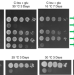 Coupling of 5S RNP rotation with maturation of functional centers during large ribosomal subunit assembly
Coupling of 5S RNP rotation with maturation of functional centers during large ribosomal subunit assembly
As assembling 60S subunits transit from the nucleolus to the nucleoplasm, they undergo significant changes in protein composition and structure. Here, the authors provide a structural view of interconnected events during the middle steps of assembly that include the maturation of the central protuberance, the peptidyltransferase center and the nascent polypeptide exit tunnel.
- Jelena Micic
- , Yu Li
- & John L. Woolford Jr.
-
Article
| Open Access Structural snapshots of human pre-60S ribosomal particles before and after nuclear export
Structural snapshots of human pre-60S ribosomal particles before and after nuclear export
Ribosome biogenesis in eukaryotes is a complex process that involves more than 200 protein factors. Here the authors present a structural analysis of a collection of human pre-60S structures sampled through a nuclear export adaptor NMD3, representing structural snapshots of pre-60S particles immediately before and after passing through nuclear pore complex.
- Xiaomeng Liang
- , Mei-Qing Zuo
- & Ning Gao
-
Article
| Open Access Distinct pre-initiation steps in human mitochondrial translation
Distinct pre-initiation steps in human mitochondrial translation
Translation within mitochondria relies upon specialized mitoribosomes and initiation factors. Here the authors use Cryo-EM and single molecule approaches to describe the early steps leading to translation initiation in human mitochondria, and the roles of mitochondria-specific ribosomal proteins and initiation factors mtIF2 and mtIF3.
- Anas Khawaja
- , Yuzuru Itoh
- & Joanna Rorbach
-
Article
| Open Access Ribosome engineering reveals the importance of 5S rRNA autonomy for ribosome assembly
Ribosome engineering reveals the importance of 5S rRNA autonomy for ribosome assembly
Ribosomes of all organisms have retained 5S rRNA as an autonomous rRNA species. Here the authors engineer a bacterial strain with ribosomes that do not have free 5S rRNA, and carry structural analyses that suggest the evolutionary preservation of 5S rRNA as an independent molecule is based on its role in the dynamic process of ribosome biogenesis.
- Shijie Huang
- , Nikolay A. Aleksashin
- & Alexander S. Mankin

 Substrate recognition and cryo-EM structure of the ribosome-bound TAC toxin of Mycobacterium tuberculosis
Substrate recognition and cryo-EM structure of the ribosome-bound TAC toxin of Mycobacterium tuberculosis