Article
|
Open Access
Featured
-
-
Article
| Open Access The proteasome modulates endocytosis specifically in glomerular cells to promote kidney filtration
The proteasome modulates endocytosis specifically in glomerular cells to promote kidney filtration
In the kidney, maintaining permeability of the filtration barrier is critical. Here, Sachs W. et al show that homeostasis of podocytes and glomerular endothelial cells relies on differing proteasome constitutions which orchestrate endocytic activity in addition to protein degradation.
- Wiebke Sachs
- , Lukas Blume
- & Catherine Meyer-Schwesinger
-
Article
| Open Access Molecular basis of SAP05-mediated ubiquitin-independent proteasomal degradation of transcription factors
Molecular basis of SAP05-mediated ubiquitin-independent proteasomal degradation of transcription factors
SAP05, a secreted effector of the obligate parasitic bacteria phytoplasma, bridges host SPL and GATA transcription factors (TFs) to the 26 S proteasome subunit RPN10 for ubiquitination-independent degradation. Here, the authors report the details on how SAP05 interacts with SPL5, GATA18 and RPN10.
- Xiaojie Yan
- , Xinxin Yuan
- & Cheng Dong
-
Article
| Open Access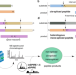 Protein degradation by human 20S proteasomes elucidates the interplay between peptide hydrolysis and splicing
Protein degradation by human 20S proteasomes elucidates the interplay between peptide hydrolysis and splicing
Do proteasomes catalyze peptide splicing? Here, the authors develop and apply a method to identify spliced peptides produced from entire proteins, confirm that proteasomes produce a sizeable variety of cis-spliced peptides with well-defined characteristics, and show that non-spliced and spliced peptides are concentrated in hotspots.
- Wai Tuck Soh
- , Hanna P. Roetschke
- & Michele Mishto
-
Article
| Open Access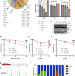 FLIP(C1orf112)-FIGNL1 complex regulates RAD51 chromatin association to promote viability after replication stress
FLIP(C1orf112)-FIGNL1 complex regulates RAD51 chromatin association to promote viability after replication stress
Recombination is essential for life. Here, the authors characterize FLIP as a novel regulator of the key recombination protein RAD51’s functions. FLIP loss caused marked sensitivity to DNA damage, increased DNA breakage and defective replication.
- Jessica D. Tischler
- , Hiroshi Tsuchida
- & Richard O. Adeyemi
-
Article
| Open Access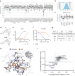 DCAF1-based PROTACs with activity against clinically validated targets overcoming intrinsic- and acquired-degrader resistance
DCAF1-based PROTACs with activity against clinically validated targets overcoming intrinsic- and acquired-degrader resistance
Targeted protein degradation (TPD) is a key modality for drug discovery. Here the authors present the discovery and analysis of reversible DCAF1-PROTACs, which show efficacy in cellular environments resistant to VHL-PROTACs or with acquired resistance to CRBN-PROTACs.
- Martin Schröder
- , Martin Renatus
- & Claudio R. Thoma
-
Article
| Open Access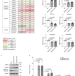 Immunoproteasome-specific subunit PSMB9 induction is required to regulate cellular proteostasis upon mitochondrial dysfunction
Immunoproteasome-specific subunit PSMB9 induction is required to regulate cellular proteostasis upon mitochondrial dysfunction
Mitochondrial dysfunction results in the accumulation of mitochondrial proteins in the cytosol. Here, the authors show that the immunoproteasome subunit PSMB9 promotes protein degradation to maintain cellular protein homeostasis.
- Minji Kim
- , Remigiusz A. Serwa
- & Agnieszka Chacinska
-
Article
| Open Access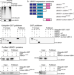 The UBX domain in UBXD1 organizes ubiquitin binding at the C-terminus of the VCP/p97 AAA-ATPase
The UBX domain in UBXD1 organizes ubiquitin binding at the C-terminus of the VCP/p97 AAA-ATPase
The function of VCP/p97 AAA-ATPase cofactor UBXD1 and its UBX domain has been elusive. Here the authors show that the extended UBXD1 UBX domain is located at the p97 pore exit where it binds ubiquitin, suggesting that UBXD1 receives unfolded substrates and hands them off for down-stream processing.
- Mike Blueggel
- , Alexander Kroening
- & Christine Beuck
-
Article
| Open Access Allosteric regulation of the 20S proteasome by the Catalytic Core Regulators (CCRs) family
Allosteric regulation of the 20S proteasome by the Catalytic Core Regulators (CCRs) family
A family of regulators named Catalytic Core Regulators (CCRs) oversees the function of the 20S proteasome. Here, the authors show that CCRs function through an allosteric mechanism, coupling the physical binding of the PSMB4 β-subunit with attenuation of the proteasome three proteolytic activities.
- Fanindra Kumar Deshmukh
- , Gili Ben-Nissan
- & Michal Sharon
-
Article
| Open Access Structure of the reduced microsporidian proteasome bound by PI31-like peptides in dormant spores
Structure of the reduced microsporidian proteasome bound by PI31-like peptides in dormant spores
Proteasomes are vital eukaryotic complexes that recycle unneeded proteins. Here, the authors present the structure of a compacted proteasome derived from the dormant stage of parasitic microsporidia and bound by an endogenous inhibitory protein.
- Nathan Jespersen
- , Kai Ehrenbolger
- & Jonas Barandun
-
Article
| Open Access A selective and orally bioavailable VHL-recruiting PROTAC achieves SMARCA2 degradation in vivo
A selective and orally bioavailable VHL-recruiting PROTAC achieves SMARCA2 degradation in vivo
Protein degraders are an emerging drug modality; however, their properties lie beyond typical drug-like space. Here the authors report optimisation via structure-based exit vector and linker design towards the VHL-recruiting PROTAC ACBI2, an orally bioavailable and selective degrader of SMARCA2.
- Christiane Kofink
- , Nicole Trainor
- & William Farnaby
-
Article
| Open Access Regulation of AUXIN RESPONSE FACTOR condensation and nucleo-cytoplasmic partitioning
Regulation of AUXIN RESPONSE FACTOR condensation and nucleo-cytoplasmic partitioning
Auxin-driven transcriptional responses are mediated by ARF transcription factors. Here the authors characterize an F-box protein, AFF1, that regulates the accumulation, condensation, and nucleo-cytoplasmic partitioning of ARF19 and ARF7.
- Hongwei Jing
- , David A. Korasick
- & Lucia C. Strader
-
Article
| Open Access Stress routes clients to the proteasome via a BAG2 ubiquitin-independent degradation condensate
Stress routes clients to the proteasome via a BAG2 ubiquitin-independent degradation condensate
While cellular stress shuts down translation, how protein degradation occurs with stress is incompletely understood. The authors describe a stress-induced phase separated organelle that mediates ubiquitin-independent degradation in the proteasome.
- Daniel C. Carrettiero
- , Maria C. Almeida
- & Kenneth S. Kosik
-
Article
| Open Access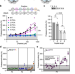 The YΦ motif defines the structure-activity relationships of human 20S proteasome activators
The YΦ motif defines the structure-activity relationships of human 20S proteasome activators
The proteasome complexes, composed of 20S core particles and one or two regulatory particles (proteasome activators), degrade most eukaryotic proteins. Here, the authors identify a sequence motif and resolve its interactions mediating the activation of the human 20S proteasome.
- Kwadwo A. Opoku-Nsiah
- , Andres H. de la Pena
- & Jason E. Gestwicki
-
Article
| Open Access Allosteric control of Ubp6 and the proteasome via a bidirectional switch
Allosteric control of Ubp6 and the proteasome via a bidirectional switch
The interplay of the proteasome and deubiquitinase Ubp6 is crucial for the degradation of ubiquitylated substrates. Here, the authors provide structural insights into the allosteric mechanism by which the activities of both Ubp6 and the proteasome are regulated.
- Ka Ying Sharon Hung
- , Sven Klumpe
- & Daniel Finley
-
Article
| Open Access Structural basis of prokaryotic ubiquitin-like protein engagement and translocation by the mycobacterial Mpa-proteasome complex
Structural basis of prokaryotic ubiquitin-like protein engagement and translocation by the mycobacterial Mpa-proteasome complex
Pup is the bacterial analog of ubiquitin for targeting proteins to the proteasome. Here, the authors use cryoEM to visualize structures of the Mycobacterium tuberculosis proteasome translocating a Pup-tagged substrate.
- Mikhail Kavalchuk
- , Ahmad Jomaa
- & Eilika Weber-Ban
-
Article
| Open Access The 20S as a stand-alone proteasome in cells can degrade the ubiquitin tag
The 20S as a stand-alone proteasome in cells can degrade the ubiquitin tag
The 20S particle is part of the 26S proteasome, but also exists as a free complex. Here, the authors outline signature activities of the 20S and combine chemical, structural, functional and proteomic assays to show that the 20S can degrade ubiquitin tags along with conjugated substrates.
- Indrajit Sahu
- , Sachitanand M. Mali
- & Michael H. Glickman
-
Article
| Open Access PARylation prevents the proteasomal degradation of topoisomerase I DNA-protein crosslinks and induces their deubiquitylation
PARylation prevents the proteasomal degradation of topoisomerase I DNA-protein crosslinks and induces their deubiquitylation
TOP1 resolves DNA supercoils by forming cleavage complexes (TOP1cc) that are trapped by TOP1 inhibitors. Here the authors provide insights into the mechanistic understanding of TOP1cc PARylation, showing that inhibition of PARG results in stabilization of TOP1cc PARylation that blocks the proteasomal degradation of TOP1cc.
- Yilun Sun
- , Jiji Chen
- & Yves Pommier
-
Article
| Open Access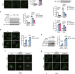 VPS34 K29/K48 branched ubiquitination governed by UBE3C and TRABID regulates autophagy, proteostasis and liver metabolism
VPS34 K29/K48 branched ubiquitination governed by UBE3C and TRABID regulates autophagy, proteostasis and liver metabolism
Autophagy and the ubiquitin–proteasome system (UPS) are cellular quality control processes, but their coordination remains unclear. Here, the authors show that branched ubiquitination of VPS34 functions as a switch between UPS and autophagy and has an important role in lipid metabolism in the liver.
- Yu-Hsuan Chen
- , Tzu-Yu Huang
- & Ruey-Hwa Chen
-
Article
| Open Access Maintenance of type 2 glycolytic myofibers with age by Mib1-Actn3 axis
Maintenance of type 2 glycolytic myofibers with age by Mib1-Actn3 axis
Muscle atrophy is associated with ageing, but the underlying molecular mechanisms are not well understood. Here, they authors show that ablation of the E3 ubiquitin ligase Mib1 is important for myofibre maintenance via a mechanism that involves targeting and degradation of Actn3, and that Mib1 ablation in mice induces muscle atrophy which can be rescued by knockown of Actn3 expression.
- Ji-Yun Seo
- , Jong-Seol Kang
- & Young-Yun Kong
-
Article
| Open Access 20S proteasomes secreted by the malaria parasite promote its growth
20S proteasomes secreted by the malaria parasite promote its growth
Plasmodium falciparum secretes extracellular vesicles (EVs) while growing inside red blood cells (RBCs). Here the authors show that these EVs contain assembled and functional 20S proteasome complexes that remodel the cytoskeleton of naïve human RBCs, priming the RBCs for parasite invasion.
- Elya Dekel
- , Dana Yaffe
- & Neta Regev-Rudzki
-
Article
| Open Access Cryo-EM of mammalian PA28αβ-iCP immunoproteasome reveals a distinct mechanism of proteasome activation by PA28αβ
Cryo-EM of mammalian PA28αβ-iCP immunoproteasome reveals a distinct mechanism of proteasome activation by PA28αβ
The proteasome activator PA28αβ affects MHC class I antigen presentation by associating with immunoproteasome core particles (iCPs). Cryo-EM structures of the mammalian PA28αβ -iCP immunoproteasome and free iCP, combined with cross-linking data, reveal the complex architecture and suggest a distinct immunoproteasome activation mechanism.
- Jinhuan Chen
- , Yifan Wang
- & Yao Cong
-
Article
| Open Access Conformational maps of human 20S proteasomes reveal PA28- and immuno-dependent inter-ring crosstalks
Conformational maps of human 20S proteasomes reveal PA28- and immuno-dependent inter-ring crosstalks
Immune cells express immunoproteasomes (i20S), which bind to specialized regulators, contain different catalytic subunits and generate immunogenic peptides. HDX-MS—based assessment of the differences between the conformational dynamics of standard and i20s reveals specific, allosteric changes in i20S and upon regulator binding.
- Jean Lesne
- , Marie Locard-Paulet
- & Julien Marcoux
-
Article
| Open Access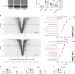 The Hsp70-Hsp90 co-chaperone Hop/Stip1 shifts the proteostatic balance from folding towards degradation
The Hsp70-Hsp90 co-chaperone Hop/Stip1 shifts the proteostatic balance from folding towards degradation
Hop, also known as Stip1 or Sti1, facilitates substrate transfer between the Hsp70 and Hsp90 molecular chaperones. Characterization of proteostasis-related pathways in STIP1 knock-out cell lines reveals that in eukaryotes Stip1 modulates the balance between protein folding and degradation.
- Kaushik Bhattacharya
- , Lorenz Weidenauer
- & Didier Picard
-
Article
| Open Access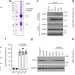 Epsin-mediated degradation of IP3R1 fuels atherosclerosis
Epsin-mediated degradation of IP3R1 fuels atherosclerosis
Endothelial cell (EC) dysfunction and inflammation contribute to plaque destabilization in atherosclerosis, increasing the risk of thrombotic events. Here, the authors show that epsin promotes EC inflammation via a mechanism involving IP3R1 degradation, and that deletion of epsin in the endothelium prevents EC dysfunctoin and atherosclerosis in mice.
- Yunzhou Dong
- , Yang Lee
- & Hong Chen
-
Article
| Open Access A masked initiation region in retinoblastoma protein regulates its proteasomal degradation
A masked initiation region in retinoblastoma protein regulates its proteasomal degradation
Human papilloma virus (HPV) E7 protein destabilizes the retinoblastoma protein (Rb) by inducing its ubiquitination in cervical cancer cells, however proteasomal degradation requires cleavage of Rb after Lys 810 and so far it has been unclear how Rb cleavage contributes to its degradation. Here, the authors combine cell based and in vitro assays and show that calpain cleavage exposes a region in Rb that is recognized by the proteasome, leading to rapid proteolysis of Rb, whereas the proteasome cannot initiate degradation efficiently on full-length Rb.
- Takuya Tomita
- , Jon M. Huibregtse
- & Andreas Matouschek
-
Article
| Open Access The proteasome 19S cap and its ubiquitin receptors provide a versatile recognition platform for substrates
The proteasome 19S cap and its ubiquitin receptors provide a versatile recognition platform for substrates
Ubiquitylated proteins are degraded by the proteasome and the three proteasome subunits Rpn10, Rpn13 and Rpn1 recognize ubiquitin chains. Here the authors employ biochemical and kinetic assays and characterise the ubiquitin chain type specificities of these three ubiquitin receptors.
- Kirby Martinez-Fonts
- , Caroline Davis
- & Andreas Matouschek
-
Article
| Open Access Structural insights into ubiquitin recognition and Ufd1 interaction of Npl4
Structural insights into ubiquitin recognition and Ufd1 interaction of Npl4
The Lys48-linked polyubiquitin-mediated proteasomal degradation in yeast depends on Cdc48 and its cofactors Ufd1 and Npl4. Here, the authors present crystal structures of Npl4 bound to Lys48-linked diubiquitin and the Npl4-binding motif of Ufd1, providing insights into the reaction mechanism of the Cdc48- Ufd1/Npl4 complex.
- Yusuke Sato
- , Hikaru Tsuchiya
- & Shuya Fukai
-
Article
| Open Access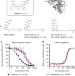 Small molecule degraders of the hepatitis C virus protease reduce susceptibility to resistance mutations
Small molecule degraders of the hepatitis C virus protease reduce susceptibility to resistance mutations
Targeted protein degradation (TPD) is a promising strategy for drug development. In this proof-of-concept study, the authors use telaprevir, which binds hepatitis C virus (HCV) NS3/4A protease, to target the protease for protein degradation, and show inhibition of wildtype as well as drug resistant HCV.
- Mélissanne de Wispelaere
- , Guangyan Du
- & Priscilla L. Yang
-
Article
| Open Access AWD regulates timed activation of BMP signaling in intestinal stem cells to maintain tissue homeostasis
AWD regulates timed activation of BMP signaling in intestinal stem cells to maintain tissue homeostasis
Regeneration after injury in the Drosophila intestine involves early activation of intestinal stem cells (ISCs) and subsequent return to quiescence. Here the authors show that return to quiescence by ISCs involves BMP Type I receptor Tkv protein stabilization along with AWD mediated internalization into endocytic vesicles.
- Xiaoyu Tracy Cai
- , Hongjie Li
- & Heinrich Jasper
-
Article
| Open Access SAGA DUBm-mediated surveillance regulates prompt export of stress-inducible transcripts for proteostasis
SAGA DUBm-mediated surveillance regulates prompt export of stress-inducible transcripts for proteostasis
Stress-inducible transcripts are quickly exported to preserve cell survival when cells are under stress. Here, the authors suggest that Sgf73p of the SAGA deubiquitylating module monitors messenger ribonucleoprotein biogenesis to regulate non-canonical export of stress-inducible transcripts.
- Minhoo Kim
- , Yoonjung Choi
- & Daeyoup Lee
-
Article
| Open Access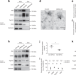 KIBRA controls exosome secretion via inhibiting the proteasomal degradation of Rab27a
KIBRA controls exosome secretion via inhibiting the proteasomal degradation of Rab27a
Exosomes are intercellular signaling vesicles created by fusion of multivesicular bodies (MVBs) and the plasma membrane (PM), but secretory regulation is ill-defined. Song et al. show that KIBRA controls exosome secretion by protecting Rab27a from proteasomal degradation, promoting MVB-PM docking.
- Lin Song
- , Shi Tang
- & Yifeng Du
-
Article
| Open Access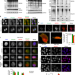 NEDDylation promotes nuclear protein aggregation and protects the Ubiquitin Proteasome System upon proteotoxic stress
NEDDylation promotes nuclear protein aggregation and protects the Ubiquitin Proteasome System upon proteotoxic stress
Protein NEDDylation increases upon proteotoxic stress but the function of this response remains to be elucidated. Here, the authors show that NEDDylation contributes to the cellular defence against proteotoxicity by promoting nuclear protein aggregation and protecting the ubiquitin proteasome system.
- Chantal M. Maghames
- , Sofia Lobato-Gil
- & Dimitris P. Xirodimas
-
Article
| Open Access The structure of the ubiquitin-like modifier FAT10 reveals an alternative targeting mechanism for proteasomal degradation
The structure of the ubiquitin-like modifier FAT10 reveals an alternative targeting mechanism for proteasomal degradation
The ubiquitin-like modifier FAT10 is composed of two ubiquitin-like domains (UBDs). Here the authors present the FAT10 UBD structures and show that the unstructured FAT10 N-terminal heptapeptide together with the poor stability of FAT10 facilitate the rapid proteasomal targeting of FAT10 along with its substrates.
- Annette Aichem
- , Samira Anders
- & Silke Wiesner
-
Article
| Open Access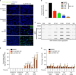 The ubiquitin ligase UBR5 suppresses proteostasis collapse in pluripotent stem cells from Huntington’s disease patients
The ubiquitin ligase UBR5 suppresses proteostasis collapse in pluripotent stem cells from Huntington’s disease patients
Induced pluripotent stem cells (iPSCs) suppress the aggregation of Huntington’s disease (HD) polyQ-expanded huntingtin (HTT). Here the authors show that proteasome activity determines the levels of mutant HTT in HD-iPSCs and find that UBR5 is a modulator of super-vigilant proteostasis of iPSCs.
- Seda Koyuncu
- , Isabel Saez
- & David Vilchez
-
Article
| Open Access Conformational switching in the coiled-coil domains of a proteasomal ATPase regulates substrate processing
Conformational switching in the coiled-coil domains of a proteasomal ATPase regulates substrate processing
Proteasomal ATPases contain functionally important coiled-coil (CC) domains, the mechanistic role of which is not fully understood. Here, the authors provide evidence for three distinct CC conformations, showing that CC conformational changes enable ATPases to switch between active and resting states.
- Aaron Snoberger
- , Evan J. Brettrager
- & David M. Smith
-
Article
| Open Access TRIM11 activates the proteasome and promotes overall protein degradation by regulating USP14
TRIM11 activates the proteasome and promotes overall protein degradation by regulating USP14
The proteasome-bound ubiquitinase USP14 plays an important role in determining proteasome activity and substrate specificity. Here the authors show that TRIM11, a member of the mammalian tripartite motif family, regulates USP14 and is an important activator of the proteasome.
- Liang Chen
- , Guixin Zhu
- & Xiaolu Yang
-
Article
| Open Access Systematic analysis of protein turnover in primary cells
Systematic analysis of protein turnover in primary cells
The proteome-wide characterization of proteostasis depends on robust approaches to determine protein half-lives. Here, the authors improve the accuracy and precision of mass spectrometry-based quantification, enabling reliable protein half-life determination in several non-dividing cell types.
- Toby Mathieson
- , Holger Franken
- & Mikhail M. Savitski
-
Article
| Open Access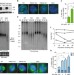 Whole proteome analysis of human tankyrase knockout cells reveals targets of tankyrase-mediated degradation
Whole proteome analysis of human tankyrase knockout cells reveals targets of tankyrase-mediated degradation
Tankyrase 1 and 2 are poly(ADP-ribose) polymerases that mark proteins for degradation, but there is a current lack of knowledge about their distinct functions and substrates. Here, the authors elucidate the cellular roles and substrates of these polymerases using comparative functional and proteomics analyses of tankyrase knockout cell lines.
- Amit Bhardwaj
- , Yanling Yang
- & Susan Smith
-
Article
| Open Access p62/SQSTM1/Sequestosome-1 is an N-recognin of the N-end rule pathway which modulates autophagosome biogenesis
p62/SQSTM1/Sequestosome-1 is an N-recognin of the N-end rule pathway which modulates autophagosome biogenesis
Soluble misfolded proteins that fail to be degraded by the ubiquitin proteasome system (UPS) are redirected to autophagy via specific adaptors, such as p62. Here the authors show that p62 recognises N-degrons in these proteins, acting as a N-recognin from the proteolytic N-end rule pathway, and targets these cargos to autophagosomal degradation.
- Hyunjoo Cha-Molstad
- , Ji Eun Yu
- & Bo Yeon Kim
-
Article
| Open Access Long-range allosteric regulation of the human 26S proteasome by 20S proteasome-targeting cancer drugs
Long-range allosteric regulation of the human 26S proteasome by 20S proteasome-targeting cancer drugs
The proteasome regulates several important cellular processes and has been identified as a target for therapeutic interventions. Here the authors map the conformational and energy landscape of the 26S proteasome upon Oprozomib binding and uncover long-range allosteric effects that control the dynamic behaviour of the proteasome.
- David Haselbach
- , Jil Schrader
- & Holger Stark
-
Article
| Open Access Structure of ubiquitylated-Rpn10 provides insight into its autoregulation mechanism
Structure of ubiquitylated-Rpn10 provides insight into its autoregulation mechanism
Ubiquitin (Ub) receptors are responsible for the recognition of ubiquitylated proteins. Here the authors describe the crystal structure of the ubiquitylated form of the Ub-receptor Rpn10, which suggest that ubiquitylation of Rpn10 promotes its dissociation from the proteasome.
- Tal Keren-Kaplan
- , Lee Zeev Peters
- & Gali Prag
-
Article
| Open Access Deubiquitinase activity is required for the proteasomal degradation of misfolded cytosolic proteins upon heat-stress
Deubiquitinase activity is required for the proteasomal degradation of misfolded cytosolic proteins upon heat-stress
Ubiquitination of misfolded proteins usually results in protein degradation. Here, the authors show that two deubiquitinases—enzymes that remove ubiquitin—are required for the proteasomal degradation of misfolded proteins in response to heat-shock in yeast.
- Nancy N. Fang
- , Mang Zhu
- & Thibault Mayor
-
Article
| Open Access ELL targets c-Myc for proteasomal degradation and suppresses tumour growth
ELL targets c-Myc for proteasomal degradation and suppresses tumour growth
The expression of the oncogene Myc is carefully controlled and dysregulation often leads to cancer. Here, the authors describe an E3 ligase for Myc—ELL—and show that it likely controls the ubiquitination and degradation of Myc.
- Yu Chen
- , Chi Zhou
- & Wuhan Xiao
-
Article
| Open Access A unified mechanism for proteolysis and autocatalytic activation in the 20S proteasome
A unified mechanism for proteolysis and autocatalytic activation in the 20S proteasome
The proteasome, an essential molecular machine, is a threonine protease, but the evolution and the components of its proteolytic centre are unclear. Here, the authors use structural biology and biochemistry to investigate the role of proteasome active site residues on maturation and activity.
- Eva M. Huber
- , Wolfgang Heinemeyer
- & Michael Groll
-
Article
| Open Access Open-gate mutants of the mammalian proteasome show enhanced ubiquitin-conjugate degradation
Open-gate mutants of the mammalian proteasome show enhanced ubiquitin-conjugate degradation
The proteasome plays a key role in proteostasis by mediating the degradation of ubiquitinated substrates. Here the authors show that an open-gate mutant of the proteasome is hyperactive towards a subset of substrates and can effectively delay the accumulation of toxic protein aggregates.
- Won Hoon Choi
- , Stefanie A. H. de Poot
- & Min Jae Lee
-
Article
| Open Access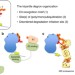 Tripartite degrons confer diversity and specificity on regulated protein degradation in the ubiquitin-proteasome system
Tripartite degrons confer diversity and specificity on regulated protein degradation in the ubiquitin-proteasome system
Degrons are determinants within proteins that direct programmed degradation by the ubiquitinproteasome system. Here, the authors propose a three-part degron architecture which contains an E3-ligase recognition motif, a ubiquitination site(s), and a disordered site to initiate degradation.
- Mainak Guharoy
- , Pallab Bhowmick
- & Peter Tompa
-
Article
| Open Access ATP binding to neighbouring subunits and intersubunit allosteric coupling underlie proteasomal ATPase function
ATP binding to neighbouring subunits and intersubunit allosteric coupling underlie proteasomal ATPase function
The 26S proteasome contains a hexamer of ATPase subunits, which binds, unfolds and translocates substrates in an ATP-dependent manner. Kim et al. use FRET to show that ATP binding preferentially occurs at neighbouring subunits of the hexamer, and identify two allosteric systems that coordinate translocation.
- Young-Chan Kim
- , Aaron Snoberger
- & David M. Smith
-
Article
| Open Access Involvement of a eukaryotic-like ubiquitin-related modifier in the proteasome pathway of the archaeon Sulfolobus acidocaldarius
Involvement of a eukaryotic-like ubiquitin-related modifier in the proteasome pathway of the archaeon Sulfolobus acidocaldarius
Small archaeal ubiquitin-like modifiers (SAMPs) have been hypothesized to be part of an ancestral version of the ubiquitin-proteasome system. Here, Anjum et al. identify a SAMP homologous to the eukaryotic ubiquitin-related modifier-1 and show that it is processed by the 20S core proteasome in S. acidocaldarius.
- Rana S. Anjum
- , Sian M. Bray
- & Nicholas P. Robinson
-
Article
| Open Access Cryo-EM reveals the conformation of a substrate analogue in the human 20S proteasome core
Cryo-EM reveals the conformation of a substrate analogue in the human 20S proteasome core
The proteasome is a highly regulated complex fundamental for cell homeostasis and a target for cancer therapy. Here the authors use cryo-EM and single-particle analysis to obtain a detailed map of the interactions between each active sites of the core 20S proteasome and the irreversible inhibitor AdaAhx3L3VS.
- Paula C.A. da Fonseca
- & Edward P. Morris

 Viscosity-dependent control of protein synthesis and degradation
Viscosity-dependent control of protein synthesis and degradation