Article
|
Open Access
Featured
-
-
Article
| Open Access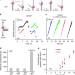 Helicase promotes replication re-initiation from an RNA transcript
Helicase promotes replication re-initiation from an RNA transcript
During DNA replication, replicative helicases play an essential role for DNA unwinding to occur. Here the authors find that bacteriophage T7 helicase is also involved in replication re-initiation by interacting with a non-replicating DNAP and increasing unwinding rate.
- Bo Sun
- , Anupam Singh
- & Michelle D. Wang
-
Article
| Open Access A quantitative mass spectrometry-based approach to monitor the dynamics of endogenous chromatin-associated protein complexes
A quantitative mass spectrometry-based approach to monitor the dynamics of endogenous chromatin-associated protein complexes
Chromatin-associated protein complexes play a critical role in the regulation of gene expression in health and disease. Here, the authors describe a sensitive mass spectrometry-based method to monitor the dynamic interactions of endogenous chromatin-associated protein complexes in clinical samples.
- Evangelia K. Papachristou
- , Kamal Kishore
- & Jason S. Carroll
-
Article
| Open Access Cellular stress alters 3′UTR landscape through alternative polyadenylation and isoform-specific degradation
Cellular stress alters 3′UTR landscape through alternative polyadenylation and isoform-specific degradation
The function and consequences of alternative polyadenylation (APA) in stressed cells are largely unclear. Here, the authors show that stress-induced mRNA degradation depends on 3′UTR length and that APA-mediated 3′UTR shortening is an adaptive stress response mechanism for selective transcript stabilization.
- Dinghai Zheng
- , Ruijia Wang
- & Bin Tian
-
Article
| Open Access Map of synthetic rescue interactions for the Fanconi anemia DNA repair pathway identifies USP48
Map of synthetic rescue interactions for the Fanconi anemia DNA repair pathway identifies USP48
Fanconi anemia is a rare disease caused by defective DNA interstrand crosslink repair. Here the authors observe that USP48 deficiencies reduce chromosomal instability in FA-defective cells, suggesting it might be a potential therapeutic target.
- Georgia Velimezi
- , Lydia Robinson-Garcia
- & Joanna I. Loizou
-
Article
| Open Access Dephosphorylation of the HIV-1 restriction factor SAMHD1 is mediated by PP2A-B55α holoenzymes during mitotic exit
Dephosphorylation of the HIV-1 restriction factor SAMHD1 is mediated by PP2A-B55α holoenzymes during mitotic exit
SAMHD1 is a critical restriction factor for HIV-1 and its antiviral activity is regulated by T592 phosphorylation. Here, Schott et al. show that the phosphatase PP2A-B55α dephosphorylates SAMHD1 during mitotic exit, rendering it antivirally active in G1 phase of primary CD4+ T cells.
- Kerstin Schott
- , Nina V. Fuchs
- & Renate König
-
Article
| Open Access HuR regulates telomerase activity through TERC methylation
HuR regulates telomerase activity through TERC methylation
Mutations in the RNA component TERC can cause telomerase dysfunction but the underlying mechanisms are largely unknown. Here, the authors show that RNA-binding protein HuR regulates telomerase function by enhancing the methylation of TERC, which is impaired by several disease-relevant TERC mutations.
- Hao Tang
- , Hu Wang
- & Wengong Wang
-
Article
| Open Access Parasitic insect-derived miRNAs modulate host development
Parasitic insect-derived miRNAs modulate host development
The moth Plutella xylostella during its larval stage is the host of the endoparasitic wasp Cotesia vestalis. Here the authors show that the parasitoids deliver microRNAs to their hosts through their symbiotic virus and specialized cells leading to induced developmental delay.
- Zhi-zhi Wang
- , Xi-qian Ye
- & Xue-xin Chen
-
Article
| Open Access Precise temporal regulation of alternative splicing during neural development
Precise temporal regulation of alternative splicing during neural development
The precise timing of neurodevelopmental splicing switches and the underlying regulatory mechanisms remain poorly understood. This study identifies two major waves of developmental switches under the control of distinct combinations of RNA-binding proteins in central and peripheral nervous systems.
- Sebastien M. Weyn-Vanhentenryck
- , Huijuan Feng
- & Chaolin Zhang
-
Article
| Open Access SETBP1 induces transcription of a network of development genes by acting as an epigenetic hub
SETBP1 induces transcription of a network of development genes by acting as an epigenetic hub
SETBP1 variants occur as somatic mutations in several malignancies and as de novo germline mutations in developmental disorders. Here the authors provide evidence that SETBP1 binds to gDNA in AT-rich promoter regions to promote target gene upregulation, indicating SETBP1 functions directly to regulate transcription.
- Rocco Piazza
- , Vera Magistroni
- & Carlo Gambacorti-Passerini
-
Article
| Open Access In vivo base editing of post-mitotic sensory cells
In vivo base editing of post-mitotic sensory cells
Base editing allows the precise introduction of point mutations into cellular DNA without requiring double-stranded DNA breaks or homology-directed repair, which is inefficient in postmitotic cells. Here the authors demonstrate in vivo base editing of post-mitotic somatic cells in the postnatal mouse inner ear with physiological outcomes.
- Wei-Hsi Yeh
- , Hao Chiang
- & David R. Liu
-
Article
| Open Access Tet1 and Tet2 maintain mesenchymal stem cell homeostasis via demethylation of the P2rX7 promoter
Tet1 and Tet2 maintain mesenchymal stem cell homeostasis via demethylation of the P2rX7 promoter
Tet-mediated DNA oxidation converts 5-methylcytosine (5-mC) to 5-hydroxymethylcytosine (5-hmC), which is essential to regulate different biological processes. Here the authors show that Tet1 and Tet2 regulate mesenchymal stem cell and bone homeostasis through demethylation of P2rX7 promoter.
- Ruili Yang
- , Tingting Yu
- & Songtao Shi
-
Article
| Open Access Structural basis for recognition of 53BP1 tandem Tudor domain by TIRR
Structural basis for recognition of 53BP1 tandem Tudor domain by TIRR
The p53-binding protein 1 (53BP1) regulates the choice of the DNA double-strand break repair pathway. Here the authors present the crystal structure of Tudor-interacting repair regulator (TIRR) bound to the 53BP1 tandem Tudor domain, which reveals how TIRR blocks H4K20me2 binding to 53BP1 Tudor and functionally differs from its paralog Nudt16.
- Yaxin Dai
- , Aili Zhang
- & Zheng Zhou
-
Article
| Open Access The histone demethylase Phf2 acts as a molecular checkpoint to prevent NAFLD progression during obesity
The histone demethylase Phf2 acts as a molecular checkpoint to prevent NAFLD progression during obesity
Steatosis is characterized by initial accumulation of lipids, followed by inflammation and ultimately fibrosis. Here the authors show that the histone demethylase Plant Homeodomain Finger 2 protects liver form steatosis progression by acting as a co-activator of ChREBP, thus, favouring lipid accumulation without inflammation.
- Julien Bricambert
- , Marie-Clotilde Alves-Guerra
- & Renaud Dentin
-
Article
| Open Access The RPAP3-Cterminal domain identifies R2TP-like quaternary chaperones
The RPAP3-Cterminal domain identifies R2TP-like quaternary chaperones
R2TP is an HSP90 co-chaperone composed of an RPAP3-PIH1D1 heterodimer, which binds two essential AAA+ ATPases RUVBL1/RUVBL2. Here authors use a structural approach to study RPAP3 and find an RPAP3-like protein (SPAG1) which also forms a co-chaperone complex with PIH1D2 and RUVBL1/2 enriched in testis.
- Chloé Maurizy
- , Marc Quinternet
- & Edouard Bertrand
-
Article
| Open Access Tumour-associated missense mutations in the dMi-2 ATPase alters nucleosome remodelling properties in a mutation-specific manner
Tumour-associated missense mutations in the dMi-2 ATPase alters nucleosome remodelling properties in a mutation-specific manner
ATP-dependent chromatin remodelers are often found mutated in human cancers. Here, the authors characterize the nucleosome remodelling properties of cancer-associated mutants of the Drosophila Chd4 homolog dMi-2.
- Kristina Kovač
- , Anja Sauer
- & Alexander Brehm
-
Article
| Open Access Live-cell single-molecule dynamics of PcG proteins imposed by the DIPG H3.3K27M mutation
Live-cell single-molecule dynamics of PcG proteins imposed by the DIPG H3.3K27M mutation
Diffuse intrinsic pontine gliomas exhibit a characteristic mutation of lysine 27 to methionine (K27M) in genes encoding histone H3.3. Here the authors show that the H3.3K27M mutation imposes a specific pattern of H3.3K27 methylation by altering the target search dynamics of PcG proteins.
- Roubina Tatavosian
- , Huy Nguyen Duc
- & Xiaojun Ren
-
Article
| Open Access The CaMKII/NMDA receptor complex controls hippocampal synaptic transmission by kinase-dependent and independent mechanisms
The CaMKII/NMDA receptor complex controls hippocampal synaptic transmission by kinase-dependent and independent mechanisms
Calcium-calmodulin-dependent protein kinase II (CaMKII) is well known for its roles in synaptic plasticity. Using a series of molecular replacement experiments, the authors show that the kinase function of CaMKII is required for long-term plasticity and basal AMPA receptor-mediated transmission.
- Salvatore Incontro
- , Javier Díaz-Alonso
- & Roger A. Nicoll
-
Article
| Open Access Postnatal DNA demethylation and its role in tissue maturation
Postnatal DNA demethylation and its role in tissue maturation
Here the authors show that a large fraction of the tissue-specific methylation pattern is generated postnatally. These changes, which occur in response to hormone signaling, appear to play a major role in the regulation of gene expression and tissue maturation in the liver.
- Yitzhak Reizel
- , Ofra Sabag
- & Howard Cedar
-
Article
| Open Access Distinct roles of XPF-ERCC1 and Rad1-Rad10-Saw1 in replication-coupled and uncoupled inter-strand crosslink repair
Distinct roles of XPF-ERCC1 and Rad1-Rad10-Saw1 in replication-coupled and uncoupled inter-strand crosslink repair
The yeast Rad1–Rad10 complex has multiple roles in DNA damage repair. Here the authors uncover mutants that uncouple the roles in UV excision repair and non-NER functions.
- Ja-Hwan Seol
- , Cory Holland
- & Sang Eun Lee
-
Article
| Open Access Combining laser capture microdissection and proteomics reveals an active translation machinery controlling invadosome formation
Combining laser capture microdissection and proteomics reveals an active translation machinery controlling invadosome formation
Invadosomes degrade extracellular matrix and facilitate cell invasion but their molecular composition is not fully understood. Here, the authors combine laser capture and mass spectrometry to map the proteome of invadosomes, showing that they rely on internal translational activity to maintain their structure.
- Zakaria Ezzoukhry
- , Elodie Henriet
- & Frédéric Saltel
-
Article
| Open Access rbFOX1/MBNL1 competition for CCUG RNA repeats binding contributes to myotonic dystrophy type 1/type 2 differences
rbFOX1/MBNL1 competition for CCUG RNA repeats binding contributes to myotonic dystrophy type 1/type 2 differences
Myotonic dystrophy (DM) type 2 is a neuromuscular pathology caused by large expansions of CCTG repeats. Here the authors find that rbFOX1 RNA binding protein binds to CCUG RNA repeats and competes with MBNL1 for the binding to CCUG repeats, releasing MBNL1 from sequestration in DM2 muscle cells.
- Chantal Sellier
- , Estefanía Cerro-Herreros
- & Nicolas Charlet-Berguerand
-
Article
| Open Access L-SCRaMbLE as a tool for light-controlled Cre-mediated recombination in yeast
L-SCRaMbLE as a tool for light-controlled Cre-mediated recombination in yeast
The International Synthetic Yeast Sc2.0 project has built Cre recombinase sites into synthetic chromosomes, enabling rapid genome evolution. Here the authors demonstrate L-SCRaMbLE, a light-controlled recombinase tool with improved control over recombination events.
- Lena Hochrein
- , Leslie A. Mitchell
- & Bernd Mueller-Roeber
-
Article
| Open Access Intron retention and nuclear loss of SFPQ are molecular hallmarks of ALS
Intron retention and nuclear loss of SFPQ are molecular hallmarks of ALS
Intron retention (IR) can increase protein diversity and function, and yet unregulated IR may be detrimental to cellular health. This study shows that aberrant IR occurs in ALS and finds nuclear loss of an RNA-binding protein called SFPQ as a new molecular hallmark in this devastating condition.
- Raphaelle Luisier
- , Giulia E. Tyzack
- & Rickie Patani
-
Article
| Open Access Phenotypic diversification by enhanced genome restructuring after induction of multiple DNA double-strand breaks
Phenotypic diversification by enhanced genome restructuring after induction of multiple DNA double-strand breaks
DNA double-strand break (DSB) leads to genome rearrangements with various genetic and phenotypic effects. Here, the authors develop a tool to induce large-scale genome restructuring by introducing conditional multiple DNA breaks, and produce various traits in yeast and Arabidopsis thaliana.
- Nobuhiko Muramoto
- , Arisa Oda
- & Kunihiro Ohta
-
Article
| Open Access Structure of a cleavage-independent HIV Env recapitulates the glycoprotein architecture of the native cleaved trimer
Structure of a cleavage-independent HIV Env recapitulates the glycoprotein architecture of the native cleaved trimer
Native-like soluble HIV envelope (Env) trimers are potential vaccine immunogens, and elimination of furin-dependence could provide a DNA-based alternative. Here, Sarkar et al. show that a cleavage-independent Env construct recapitulates the architecture and glycosylation of the native cleaved trimer.
- Anita Sarkar
- , Shridhar Bale
- & Ian A. Wilson
-
Article
| Open Access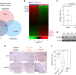 Estrogen-related receptor gamma functions as a tumor suppressor in gastric cancer
Estrogen-related receptor gamma functions as a tumor suppressor in gastric cancer
Very little is known regarding the molecular mechanisms involved in gastric cancer development. Here the authors show estrogen-related receptor gamma (ESRRG) is a tumor suppressor in gastric cancer and suggest the mechanism of this tumor suppression function involves the inhibition of Wnt signaling.
- Myoung-Hee Kang
- , Hyunji Choi
- & Yun-Yong Park
-
Review Article
| Open Access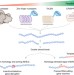 The CRISPR tool kit for genome editing and beyond
The CRISPR tool kit for genome editing and beyond
CRISPR has rapidly become an indispensable tool for biological research. Here Mazhar Adli reviews the current toolbox for editing and manipulating the genome and looks toward future developments in this fast moving field.
- Mazhar Adli
-
Article
| Open Access A CRISPRi screen in E. coli reveals sequence-specific toxicity of dCas9
A CRISPRi screen in E. coli reveals sequence-specific toxicity of dCas9
CRISPR interference (CRISPRi) is a method for targeted silencing of transcription that requires the coexpression of protein dCas9 and a customized guide RNA. Here, Cui et al. show that certain guide RNAs induce toxicity in E. coli, and provide design rules to minimize off-target effects.
- Lun Cui
- , Antoine Vigouroux
- & David Bikard
-
Article
| Open Access DOT1L inhibition attenuates graft-versus-host disease by allogeneic T cells in adoptive immunotherapy models
DOT1L inhibition attenuates graft-versus-host disease by allogeneic T cells in adoptive immunotherapy models
Adoptive T cell therapy using an allogeneic T cell graft is an encouraging therapeutic approach in cancer, but issues such as graft-versus-host disease can hinder applicability. Here, the authors show that DOT1L inhibition or DUSP6 overexpression in T cells attenuates graft-versus-host disease but retains anti-tumour activity in mouse models.
- Yuki Kagoya
- , Munehide Nakatsugawa
- & Naoto Hirano
-
Article
| Open Access PCGF5 is required for neural differentiation of embryonic stem cells
PCGF5 is required for neural differentiation of embryonic stem cells
Polycomb-group proteins are key regulators of transcriptional programs that maintain cell identity. Here the authors provide evidence that PCGF5, a subunit of Polycomb Repressor Complex 1, is important for the differentiation of mouse embryonic stem cells towards a neural cell fate.
- Mingze Yao
- , Xueke Zhou
- & Hongjie Yao
-
Article
| Open Access Aurora A-dependent CENP-A phosphorylation at inner centromeres protects bioriented chromosomes against cohesion fatigue
Aurora A-dependent CENP-A phosphorylation at inner centromeres protects bioriented chromosomes against cohesion fatigue
Sustained spindle tension applied to sister centromeres during mitosis leads to loss of sister chromatid cohesion which is known as cohesion fatigue. Here the authors show that Aurora A-dependent phosphorylation of CENP-A at the inner centromeres protects bioriented chromosomes against cohesion fatigue.
- Grégory Eot-Houllier
- , Laura Magnaghi-Jaulin
- & Christian Jaulin
-
Article
| Open Access Evolutionary instability of CUG-Leu in the genetic code of budding yeasts
Evolutionary instability of CUG-Leu in the genetic code of budding yeasts
The genetic code for amino acids is nearly universal, and among eukaryotic nuclear genomes the only known reassignments are of codon CUG in yeasts. Here, the authors identify a third independent CUG transition in budding yeasts that is still ongoing with alternative tRNAs present in the genome.
- Tadeusz Krassowski
- , Aisling Y. Coughlan
- & Kenneth H. Wolfe
-
Article
| Open Access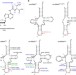 CO2-sensitive tRNA modification associated with human mitochondrial disease
CO2-sensitive tRNA modification associated with human mitochondrial disease
Transfer RNA modifications play critical roles in protein synthesis. Here the authors reveal the t6A37 tRNA modification is dynamically regulated by sensing intracellular CO2 concentration in mitochondria, implying metabolic regulation of protein synthesis.
- Huan Lin
- , Kenjyo Miyauchi
- & Tsutomu Suzuki
-
Article
| Open Access Cyclin K regulates prereplicative complex assembly to promote mammalian cell proliferation
Cyclin K regulates prereplicative complex assembly to promote mammalian cell proliferation
Prereplicative complex (pre-RC) formation during G1 is fundamental for cell replication. Here the authors report a role for cyclin K in regulating pre-RC formation in mammalian cells by affecting cyclin E1 activity.
- Tingjun Lei
- , Peixuan Zhang
- & Qintong Li
-
Article
| Open Access Genome-wide and high-density CRISPR-Cas9 screens identify point mutations in PARP1 causing PARP inhibitor resistance
Genome-wide and high-density CRISPR-Cas9 screens identify point mutations in PARP1 causing PARP inhibitor resistance
The mechanisms of PARP inhibitor (PARPi) resistance are poorly understood. Here the authors employ a CRISPR mutagenesis approach to identify PARP1 mutants causing PARPi resistance and find that PARP1 mutations are tolerated in BRCA1 mutated cells, suggesting alternative resistance mechanisms.
- Stephen J. Pettitt
- , Dragomir B. Krastev
- & Christopher J. Lord
-
Article
| Open Access Structural basis for cofilin binding and actin filament disassembly
Structural basis for cofilin binding and actin filament disassembly
Cofilin is a small actin-binding protein that accelerates actin turnover by disassembling actin filaments. Here the authors present the 3.8 Å cryo-EM structure of a cofilin-decorated actin filament and discuss mechanistic implications.
- Kotaro Tanaka
- , Shuichi Takeda
- & Akihiro Narita
-
Article
| Open Access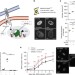 NOTCH-mediated non-cell autonomous regulation of chromatin structure during senescence
NOTCH-mediated non-cell autonomous regulation of chromatin structure during senescence
Notch can drive senescence in a cell contact dependent manner. Here the authors show that NOTCH signalling can modulate chromatin structure autonomously and non-autonomously via the JAG1-NOTCH-HMGA1 interplay during senescence.
- Aled J. Parry
- , Matthew Hoare
- & Masashi Narita
-
Article
| Open Access Molecular mechanism of influenza A NS1-mediated TRIM25 recognition and inhibition
Molecular mechanism of influenza A NS1-mediated TRIM25 recognition and inhibition
NS1 of influenza A virus inhibits TRIM25 activity, which is an E3 ligase important for induction of the interferon response. Here, Koliopoulos et al. present structures of TRIM25 and NS1 and show how NS1 binding interferes with substrate recognition of TRIM25.
- Marios G. Koliopoulos
- , Mathilde Lethier
- & Katrin Rittinger
-
Article
| Open Access Generation of App knock-in mice reveals deletion mutations protective against Alzheimer’s disease-like pathology
Generation of App knock-in mice reveals deletion mutations protective against Alzheimer’s disease-like pathology
To date, only one mutation in the gene for amyloid-beta precursor protein APP has been suggested to be protective against Alzheimer’s disease. Here, authors found using gene editing of a mutant App knock-in mouse line that deletion of the 3’UTR region is protective against amyloid-β accumulation in vivo, and subsequently identify a 52-bp element in the 3’UTR region that is responsible for this effect.
- Kenichi Nagata
- , Mika Takahashi
- & Takaomi C. Saido
-
Article
| Open Access Heterochromatin protein 1a functions for piRNA biogenesis predominantly from pericentric and telomeric regions in Drosophila
Heterochromatin protein 1a functions for piRNA biogenesis predominantly from pericentric and telomeric regions in Drosophila
Heterochromatin protein 1a (HP1a) is thought to function downstream of transposon repression in the Drosophila female germline. Here the authors show that HP1a also functions upstream of piRNA processing by repressing splicing of piRNA precursors, predominantly at telomeric and centromeric regions.
- Ryan Yee Wei Teo
- , Amit Anand
- & Toshie Kai
-
Article
| Open Access Depletion of Nsd2-mediated histone H3K36 methylation impairs adipose tissue development and function
Depletion of Nsd2-mediated histone H3K36 methylation impairs adipose tissue development and function
The epigenetic mechanisms regulating adipose tissue development are poorly understood. Here the authors show that reduction of H3K36 methylation in preadipocytes, both by H3.3K36M expression and depletion of H3K36 methyltransferase Nsd2, impairs adipogenesis by increasing H3K27me3.
- Lenan Zhuang
- , Younghoon Jang
- & Kai Ge
-
Article
| Open Access Clinical and genomic landscape of gastric cancer with a mesenchymal phenotype
Clinical and genomic landscape of gastric cancer with a mesenchymal phenotype
The prognosis and treatment of gastric cancer is complicated by heterogeneity. Here, the authors reveal two molecular subtypes, the mesenchymal subtype associated with poor survival and chemoresistance, and the epithelial phenotype associated with better survival and sensitivity to chemotherapy.
- Sang Cheul Oh
- , Bo Hwa Sohn
- & Ju-Seog Lee
-
Article
| Open Access Nuclear fate of yeast snoRNA is determined by co-transcriptional Rnt1 cleavage
Nuclear fate of yeast snoRNA is determined by co-transcriptional Rnt1 cleavage
Small nucleolar ribonucleoprotein complexes (snoRNP) are fundamental for ribosome biogenesis. Here the authors provide insight into the 5ʹend processing of S. cerevisiae snoRNA and its important role in downstream nuclear events.
- Pawel Grzechnik
- , Sylwia A. Szczepaniak
- & Nicholas J. Proudfoot
-
Article
| Open Access C/EBPβ regulates delta-secretase expression and mediates pathogenesis in mouse models of Alzheimer’s disease
C/EBPβ regulates delta-secretase expression and mediates pathogenesis in mouse models of Alzheimer’s disease
Delta-secretase cleaves both APP and Tau, and contributes to Alzheimer’s disease-like pathology. Here the authors show that C/EBPβ, a regulator of inflammation, also regulates transcription of delta-secretase in an age-dependent manner and contributes to Alzheimer’s disease-like pathology in mouse models.
- Zhi-Hao Wang
- , Ke Gong
- & Keqiang Ye
-
Article
| Open Access Activity dependent LoNA regulates translation by coordinating rRNA transcription and methylation
Activity dependent LoNA regulates translation by coordinating rRNA transcription and methylation
Non-coding RNAs have been shown to be key components in translational regulation. Here the authors characterize long nucleolus specific lncRNA termed LoNA, identified to play a role in protein biosynthesis by inhibiting rRNA production and ribosome biosynthesis in nucleoli.
- Dingfeng Li
- , Juan Zhang
- & Qiang Liu
-
Article
| Open Access Widespread intronic polyadenylation diversifies immune cell transcriptomes
Widespread intronic polyadenylation diversifies immune cell transcriptomes
Recognition of intronic polyadenylation (IpA) signals can lead to expression of truncated proteins lacking C terminal domains. Analysis of 3ʹ -seq and RNA-seq shows that IpA is widespread in circulating immune cells, while multiple myeloma cells show loss of IpA isoforms that are normally expressed in plasma cells, impacting key genes in the disease.
- Irtisha Singh
- , Shih-Han Lee
- & Christina S. Leslie
-
Article
| Open Access Linear mitochondrial DNA is rapidly degraded by components of the replication machinery
Linear mitochondrial DNA is rapidly degraded by components of the replication machinery
Damaged linearized mtDNA needs to be removed from the cell for mitochondrial genome stability. Here the authors shed light into the identity of the machinery responsible for rapidly degrading linearized DNA, implicating the role of mtDNA replication factors.
- Viktoriya Peeva
- , Daniel Blei
- & Wolfram S. Kunz
-
Article
| Open Access The RNA-binding protein YBX1 regulates epidermal progenitors at a posttranscriptional level
The RNA-binding protein YBX1 regulates epidermal progenitors at a posttranscriptional level
The integrity of the stratified epithelia relies on controlled cell turnover but it is unclear how mRNA binding proteins regulates this. Here, the authors show that the RNA binding protein Y-box binding protein-1 translationally represses cytokines, so preventing senescence and maintaining epidermal homeostasis.
- Eunjeong Kwon
- , Kristina Todorova
- & Anna Mandinova
-
Article
| Open Access Theoretical principles of transcription factor traffic on folded chromatin
Theoretical principles of transcription factor traffic on folded chromatin
How transcription factors find their targets in vivo is still poorly understood. Here the authors use molecular dynamics simulations to investigate how transcription factors diffuse on chromatin, providing a theoretical framework for understanding the key role of genome conformation in this process.
- Ruggero Cortini
- & Guillaume J. Filion
Browse broader subjects
Browse narrower subjects
- Cell division
- Chromatin
- Chromosomes
- CRISPR-Cas systems
- DNA damage and repair
- DNA metabolism
- DNA recombination
- DNA replication
- Epigenetics
- Non-coding RNAs
- Nuclear organization
- Post-translational modifications
- Protein folding
- Proteolysis
- Proteomics
- Riboswitches
- Ribozymes
- RNA metabolism
- RNAi
- Single-molecule biophysics
- Transcription
- Transcriptomics
- Translation
- Transposition

 Co-regulatory activity of hnRNP K and NS1-BP in influenza and human mRNA splicing
Co-regulatory activity of hnRNP K and NS1-BP in influenza and human mRNA splicing