Article
|
Open Access
Featured
-
-
Article
| Open Access Detecting m6A at single-molecular resolution via direct RNA sequencing and realistic training data
Detecting m6A at single-molecular resolution via direct RNA sequencing and realistic training data
Direct RNA-seq offers the possibility to identify RNA modifications on single molecules. Here, the authors report on the synthesis of biologically realistic training data and the development of mAFiA that accurately detects m6A on single read level.
- Adrian Chan
- , Isabel S. Naarmann-de Vries
- & Christoph Dieterich
-
Article
| Open Access Tracing genetic diversity captures the molecular basis of misfolding disease
Tracing genetic diversity captures the molecular basis of misfolding disease
Pei et al. applied Gaussian process-based machine learning to capture dynamic spatial covariance relationships managed by proteostasis to mediate cooperative folding on a residue basis as a standard model for precision disease management.
- Pei Zhao
- , Chao Wang
- & William E. Balch
-
Article
| Open Access Deep learning predictions of TCR-epitope interactions reveal epitope-specific chains in dual alpha T cells
Deep learning predictions of TCR-epitope interactions reveal epitope-specific chains in dual alpha T cells
Prediction of the specificity of a T cell receptor from amino acid sequence has been performed using different methods and approaches. Here the authors use TCRab sequences with known specificity to develop a deep learning TCR-epitope interaction predictor and use this method to predict specificity of dual alpha chain TCRs and TCRs specific for different antigens.
- Giancarlo Croce
- , Sara Bobisse
- & David Gfeller
-
Article
| Open Access Teacher-student collaborated multiple instance learning for pan-cancer PDL1 expression prediction from histopathology slides
Teacher-student collaborated multiple instance learning for pan-cancer PDL1 expression prediction from histopathology slides
PDL1 expression is a common biomarker for immunotherapy response in cancer, and it is usually quantified using immunohistochemistry. Here, the authors develop a weakly supervised learning approach combining multiple instance learning and a teacher-student framework to predict PDL1 expression from histopathological imaging.
- Darui Jin
- , Shangying Liang
- & Xiangzhi Bai
-
Article
| Open Access scButterfly: a versatile single-cell cross-modality translation method via dual-aligned variational autoencoders
scButterfly: a versatile single-cell cross-modality translation method via dual-aligned variational autoencoders
Technical limitations of simultaneously multi-omics profiling lead to highly noisy multi-modal data and substantial costs. Here, authors proposed a versatile framework and data augmentation schemes, capable of single-cell cross-modality translation and multiple extensive applications.
- Yichuan Cao
- , Xiamiao Zhao
- & Shengquan Chen
-
Article
| Open Access Context-aware deep learning enables high-efficacy localization of high concentration microbubbles for super-resolution ultrasound localization microscopy
Context-aware deep learning enables high-efficacy localization of high concentration microbubbles for super-resolution ultrasound localization microscopy
Ultrasound localisation microscopy enables deep tissue microvascular imaging. Here, authors introduce LOCA-ULM, a deep learning pipeline enhancing localisation accuracy in high microbubble concentrations. LOCA-ULM reveals dense cerebrovascular networks and enhances the sensitivity of functional ULM.
- YiRang Shin
- , Matthew R. Lowerison
- & Pengfei Song
-
Article
| Open Access Genomic language model predicts protein co-regulation and function
Genomic language model predicts protein co-regulation and function
A gene’s function is governed by its sequence, structure and context. Here, the authors develop a genomic language model that learns contextualized functional representations from diverse and large-scale metagenomic datasets.
- Yunha Hwang
- , Andre L. Cornman
- & Peter R. Girguis
-
Article
| Open Access Data-driven identification of predictive risk biomarkers for subgroups of osteoarthritis using interpretable machine learning
Data-driven identification of predictive risk biomarkers for subgroups of osteoarthritis using interpretable machine learning
Osteoarthritis can be caused by multiple biological mechanisms but the drivers of disease risk are not well understood. Here, the authors use data from UK Biobank in machine learning models to identify clinical and biological markers associated with development of osteoarthritis and identify sub-groups with different risk profiles.
- Rikke Linnemann Nielsen
- , Thomas Monfeuga
- & Ramneek Gupta
-
Article
| Open Access Mapping cell-to-tissue graphs across human placenta histology whole slide images using deep learning with HAPPY
Mapping cell-to-tissue graphs across human placenta histology whole slide images using deep learning with HAPPY
Placenta histopathology for maternal and newborn health is highly specialised and time consuming. Here, authors present a deep learning pipeline for quantifying cells and tissues in placenta whole slide images, revealing biological and clinical insights.
- Claudia Vanea
- , Jelisaveta Džigurski
- & Christoffer Nellåker
-
Article
| Open Access SpiDe-Sr: blind super-resolution network for precise cell segmentation and clustering in spatial proteomics imaging
SpiDe-Sr: blind super-resolution network for precise cell segmentation and clustering in spatial proteomics imaging
Imaging mass cytometry (IMC) is a powerful single-cell resolution platform for targeted spatial proteomics, but it can be constrained by imaging noise and resolution. Here, the authors propose SpiDe-Sr, a super-resolution network embedded with a denoising module for IMC spatial resolution enhancement.
- Rui Chen
- , Jiasu Xu
- & Xianting Ding
-
Article
| Open Access 3D molecular generative framework for interaction-guided drug design
3D molecular generative framework for interaction-guided drug design
Designing a molecule that favorably binds to a protein pocket is a keystone of drug discovery. Zhung et al. devise DeepICL, which leverages the generalizable features of non-covalent protein-ligand interactions on a 3D molecular generative model, improving the quality of AI-designed molecules.
- Wonho Zhung
- , Hyeongwoo Kim
- & Woo Youn Kim
-
Article
| Open Access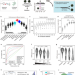 Precise prediction of phase-separation key residues by machine learning
Precise prediction of phase-separation key residues by machine learning
Understanding intracellular phase separation is essential for transcriptional control, cell fate, and disease. Here the authors report PSPHunter which accurately predicts key residues, aiding in disease-associated protein identification and mechanistic insights.
- Jun Sun
- , Jiale Qu
- & Junjun Ding
-
Article
| Open Access Predicting and improving complex beer flavor through machine learning
Predicting and improving complex beer flavor through machine learning
Perception and appreciation of food flavour depends on many factors, posing a challenge for effective prediction. Here, the authors combine extensive chemical and sensory analyses of 250 commercial Belgian beers to train machine learning models that enable flavour and consumer appreciation prediction.
- Michiel Schreurs
- , Supinya Piampongsant
- & Kevin J. Verstrepen
-
Article
| Open Access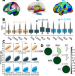 The genetic architecture of multimodal human brain age
The genetic architecture of multimodal human brain age
The biological basis of brain aging is not well understood, but it has implications for human health. Here, the authors explore the genetic basis of human brain aging, finding genetic variants, genes and potential causal relationships with disease.
- Junhao Wen
- , Bingxin Zhao
- & Christos Davatzikos
-
Article
| Open Access Accurate and rapid antibiotic susceptibility testing using a machine learning-assisted nanomotion technology platform
Accurate and rapid antibiotic susceptibility testing using a machine learning-assisted nanomotion technology platform
Sturm et. al developed a 2 to 4 h antibiotic susceptibility test based on bacterial vibrations. This diagnostic test applies to the most frequently found gram-negative bacteria in bloodstream infections and demonstrates its potential in contributing to faster treatment decisions.
- Alexander Sturm
- , Grzegorz Jóźwiak
- & Danuta Cichocka
-
Article
| Open Access Data-driven prediction of colonization outcomes for complex microbial communities
Data-driven prediction of colonization outcomes for complex microbial communities
Predicting the colonization of exogenous species in complex communities is a challenge in ecology. Here, the authors propose a data-driven approach to predict colonization outcomes and perform validation experiments in human gut microbial communities.
- Lu Wu
- , Xu-Wen Wang
- & Lei Dai
-
Article
| Open Access Local prediction-learning in high-dimensional spaces enables neural networks to plan
Local prediction-learning in high-dimensional spaces enables neural networks to plan
The task of planning a sequence of actions, and dynamically adjusting the plan in dependence of unforeseen circumstances, remains challenging for artificial intelligence frameworks. The authors introduce a learning approach inspired by cognitive functions, that demonstrates high flexibility and generalization capability in planning tasks, suitable for on-chip learning.
- Christoph Stöckl
- , Yukun Yang
- & Wolfgang Maass
-
Article
| Open Access Enabling large-scale screening of Barrett’s esophagus using weakly supervised deep learning in histopathology
Enabling large-scale screening of Barrett’s esophagus using weakly supervised deep learning in histopathology
Diagnosis of Barrett’s esophagus depends on pathologist assessment of stained slides. Here, the authors utilise a deep learning approach to prioritize potential cases using diagnostic labels in two datasets, with the aim to improve Barrett’s screening capacity.
- Kenza Bouzid
- , Harshita Sharma
- & Javier Alvarez-Valle
-
Article
| Open Access DeepETPicker: Fast and accurate 3D particle picking for cryo-electron tomography using weakly supervised deep learning
DeepETPicker: Fast and accurate 3D particle picking for cryo-electron tomography using weakly supervised deep learning
Picking particles of biological macromolecules is critical for solving their structures in situ using cryo-electron tomograms. Here, authors develop DeepETPicker, a deep learning-based tool for fast, accurate, and automated picking of three-dimensional particles.
- Guole Liu
- , Tongxin Niu
- & Ge Yang
-
Article
| Open Access Biosensor and machine learning-aided engineering of an amaryllidaceae enzyme
Biosensor and machine learning-aided engineering of an amaryllidaceae enzyme
Amaryllidaceae alkaloids, such as the Alzheimer’s medication galantamine, are currently extracted from low-yielding daffodils. Here, authors pair biosensor-assisted screening with machine learning-guided protein design to rapidly engineer an improved Amaryllidaceae enzyme in a microbial host.
- Simon d’Oelsnitz
- , Daniel J. Diaz
- & Andrew D. Ellington
-
Article
| Open Access Systematic analysis of ChatGPT, Google search and Llama 2 for clinical decision support tasks
Systematic analysis of ChatGPT, Google search and Llama 2 for clinical decision support tasks
People will likely use ChatGPT to seek health advice. Here, the authors show promising performance of ChatGPT and open source models, but a lack of high accuracy considering medical question answering. Improvements are expected over time via domain-specific finetuning and integration of regulations.
- Sarah Sandmann
- , Sarah Riepenhausen
- & Julian Varghese
-
Article
| Open Access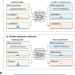 Riboformer: a deep learning framework for predicting context-dependent translation dynamics
Riboformer: a deep learning framework for predicting context-dependent translation dynamics
Riboformer is a deep learning-based framework that predicts changes in translation dynamics with codon-level precision. It corrects experimental artifacts in ribosome profiling data and identifies sequences causing ribosome stalling.
- Bin Shao
- , Jiawei Yan
- & Allen R. Buskirk
-
Article
| Open Access Machine learning-aided design and screening of an emergent protein function in synthetic cells
Machine learning-aided design and screening of an emergent protein function in synthetic cells
Here, the authors introduce a pipeline to screen machine learning generated variants of a protein that forms intracellular spatiotemporal patterns in E. coli, demonstrating the best variants can substitute the wildtype gene.
- Shunshi Kohyama
- , Béla P. Frohn
- & Petra Schwille
-
Article
| Open Access Enhancing the fairness of AI prediction models by Quasi-Pareto improvement among heterogeneous thyroid nodule population
Enhancing the fairness of AI prediction models by Quasi-Pareto improvement among heterogeneous thyroid nodule population
Artificial Intelligence (AI) models for medical diagnosis often face challenges of generalizability and fairness. Here, the authors show that the Quasi-Pareto Improvement approach is widely applicable to improving AI models among less-prevalent subgroups, promoting equitable healthcare outcomes.
- Siqiong Yao
- , Fang Dai
- & Hui Lu
-
Article
| Open Access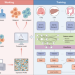 Domain generalization enables general cancer cell annotation in single-cell and spatial transcriptomics
Domain generalization enables general cancer cell annotation in single-cell and spatial transcriptomics
Efficient and accurate annotation of malignant cells is crucial for single-cell and spatial transcriptomics in cancer. Here, the authors develop Cancer-Finder, a deep-learning algorithm that can identify malignant cells in cancer single-cell and spatial transcriptomics data with speed and precision.
- Zhixing Zhong
- , Junchen Hou
- & Jia Song
-
Article
| Open Access Drug target prediction through deep learning functional representation of gene signatures
Drug target prediction through deep learning functional representation of gene signatures
Large-scale OMICs investigations of biological systems can be used to predict functional relationships between compounds, genes and proteins. Here, the authors develop a deep learning-based approach that significantly increases the number of high-quality compound-target predictions relative to existing methods.
- Hao Chen
- , Frederick J. King
- & Yingyao Zhou
-
Article
| Open Access A framework for evaluating clinical artificial intelligence systems without ground-truth annotations
A framework for evaluating clinical artificial intelligence systems without ground-truth annotations
Estimating the performance of clinical AI systems on data in the wild is complicated by distribution shift and the absence of ground-truth annotations. Here, we introduce SUDO, a framework for more reliably evaluating AI systems on data in the wild.
- Dani Kiyasseh
- , Aaron Cohen
- & Nicholas Altieri
-
Article
| Open Access Prediction of plasma ctDNA fraction and prognostic implications of liquid biopsy in advanced prostate cancer
Prediction of plasma ctDNA fraction and prognostic implications of liquid biopsy in advanced prostate cancer
Metastatic castration-resistant prostate cancer is a highly aggressive disease, with a variable response to treatment. Here, the authors validate ctDNA fraction as a poor prognostic factor and develop a model to predict whether patients harbor sufficient ctDNA for informative blood-based genotyping.
- Nicolette M. Fonseca
- , Corinne Maurice-Dror
- & Alexander W. Wyatt
-
Article
| Open Access SEMORE: SEgmentation and MORphological fingErprinting by machine learning automates super-resolution data analysis
SEMORE: SEgmentation and MORphological fingErprinting by machine learning automates super-resolution data analysis
There is a lack of universal tools to analyse protein assemblies and quantify underlying structures in single-molecule localization microscopy. Here, the authors present SEMORE, a semi-automatic machine learning framework for system- and input-dependent analysis of super-resolution data.
- Steen W. B. Bender
- , Marcus W. Dreisler
- & Nikos S. Hatzakis
-
Article
| Open Access Rapid deep learning-assisted predictive diagnostics for point-of-care testing
Rapid deep learning-assisted predictive diagnostics for point-of-care testing
A key aim in the development of diagnostic assays is improving diagnostic speed while maintaining sensitivity. Here the authors report an approach for the rapid and accurate analysis of lateral flow tests, which integrates time-series deep learning and AI verification, achieving a diagnostic time of 1-2 minutes.
- Seungmin Lee
- , Jeong Soo Park
- & Jeong Hoon Lee
-
Article
| Open Access Metabolomic machine learning predictor for diagnosis and prognosis of gastric cancer
Metabolomic machine learning predictor for diagnosis and prognosis of gastric cancer
Gastric cancer detection by endoscopy is intrusive and time-consuming, and early detection is key to improving survival. Here, the authors propose a metabolite-based model to enable early detection.
- Yangzi Chen
- , Bohong Wang
- & Zeping Hu
-
Article
| Open Access scCASE: accurate and interpretable enhancement for single-cell chromatin accessibility sequencing data
scCASE: accurate and interpretable enhancement for single-cell chromatin accessibility sequencing data
Single-cell chromatin accessibility sequencing (scCAS) data suffers from high sparsity and dimensionality. Here, authors propose an accurate and interpretable computational framework for enhancing scCAS data that considers cell-to-cell similarity.
- Songming Tang
- , Xuejian Cui
- & Shengquan Chen
-
Article
| Open Access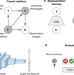 Learning representations for image-based profiling of perturbations
Learning representations for image-based profiling of perturbations
Assessing cell phenotypes in image-based assays requires solid computational methods for transforming images into quantitative data. Here, the authors present a strategy for learning representations of treatment effects from high-throughput imaging, following a causal interpretation.
- Nikita Moshkov
- , Michael Bornholdt
- & Juan C. Caicedo
-
Article
| Open Access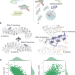 Design of target specific peptide inhibitors using generative deep learning and molecular dynamics simulations
Design of target specific peptide inhibitors using generative deep learning and molecular dynamics simulations
Here the authors report a computational approach which integrates deep learning and structural modelling to design target-specific peptides. They apply this to β-catenin and NF-κB essential modulator, resulting in improved binding, highlighting the efficacy of this strategy.
- Sijie Chen
- , Tong Lin
- & Xiaolin Cheng
-
Article
| Open Access Large language models streamline automated machine learning for clinical studies
Large language models streamline automated machine learning for clinical studies
A knowledge gap persists between machine learning developers and clinicians. Here, the authors show that the Advanced Data Analysis extension of ChatGPT could bridge this gap and simplify complex data analyses, making them more accessible to clinicians.
- Soroosh Tayebi Arasteh
- , Tianyu Han
- & Sven Nebelung
-
Article
| Open Access Efficient encoding of large antigenic spaces by epitope prioritization with Dolphyn
Efficient encoding of large antigenic spaces by epitope prioritization with Dolphyn
Profiling antibody responses to vast antigenic spaces has been challenging using programmable phage display (PhIP-Seq). Here, authors develop a methodology for compressing large proteomic spaces and have discovered human antibodies targeting gut bacteria-infecting phages.
- Anna-Maria Liebhoff
- , Thiagarajan Venkataraman
- & H. Benjamin Larman
-
Article
| Open Access Machine learning-based extrachromosomal DNA identification in large-scale cohorts reveals its clinical implications in cancer
Machine learning-based extrachromosomal DNA identification in large-scale cohorts reveals its clinical implications in cancer
‘Extrachromosomal DNA has been previously linked to tumour progression and heterogeneity, but its potential as a cancer biomarker has not been fully explored. Here, the authors develop a computational framework to refine genomic subtypes and predict response to immunotherapy in gastrointestinal cancer.
- Shixiang Wang
- , Chen-Yi Wu
- & Qi Zhao
-
Article
| Open Access A signal processing and deep learning framework for methylation detection using Oxford Nanopore sequencing
A signal processing and deep learning framework for methylation detection using Oxford Nanopore sequencing
The authors present DeepMod2, a deep-learning based computational method that allows fast and accurate detection of DNA methylation and epihaplotypes from Oxford Nanopore sequencing data.
- Mian Umair Ahsan
- , Anagha Gouru
- & Kai Wang
-
Article
| Open Access A deep-learning-based framework for identifying and localizing multiple abnormalities and assessing cardiomegaly in chest X-ray
A deep-learning-based framework for identifying and localizing multiple abnormalities and assessing cardiomegaly in chest X-ray
Accurate localization of abnormalities is crucial in the interpretation of chest X-rays. Here the authors present a deep learning framework for simultaneous localization of 14 thoracic abnormalities and calculation of cardiothoracic ratio, based on large X-ray dataset with bounding boxes created via a human-in-the-loop approach.
- Weijie Fan
- , Yi Yang
- & Dong Zhang
-
Article
| Open Access Regression-based Deep-Learning predicts molecular biomarkers from pathology slides
Regression-based Deep-Learning predicts molecular biomarkers from pathology slides
Cancer biomarkers are often continuous measurements, which poses challenges for their prediction using classification-based deep learning. Here, the authors develop a regression-based deep learning method to predict continuous biomarkers - such as the homologous repair deficiency score - from cancer histopathology images.
- Omar S. M. El Nahhas
- , Chiara M. L. Loeffler
- & Jakob Nikolas Kather
-
Article
| Open Access Predicting DNA structure using a deep learning method
Predicting DNA structure using a deep learning method
In this work, the authors report a deep learning method, Deep DNAshape, to predict the influence of flanking regions on three-dimensional DNA structure and in structural readout mechanisms of protein-DNA binding.
- Jinsen Li
- , Tsu-Pei Chiu
- & Remo Rohs
-
Article
| Open Access A multicenter clinical AI system study for detection and diagnosis of focal liver lesions
A multicenter clinical AI system study for detection and diagnosis of focal liver lesions
Early detection and accurate diagnosis of focal liver lesions are crucial for effective treatment and prognosis. Here, the authors present a fully automated diagnostic system that leverages multi-phase CT scans and clinical features, for diagnosing liver lesions.
- Hanning Ying
- , Xiaoqing Liu
- & Xiujun Cai
-
Article
| Open Access MAIVeSS: streamlined selection of antigenically matched, high-yield viruses for seasonal influenza vaccine production
MAIVeSS: streamlined selection of antigenically matched, high-yield viruses for seasonal influenza vaccine production
Vaccines combat global influenza threats, relying on timely selection of optimal seed viruses. Here, authors introduce MAIVeSS, a machine learning assisted framework to streamline vaccine seed virus selection using genomic sequence, expediting seasonal flu vaccine production and supply.
- Cheng Gao
- , Feng Wen
- & Xiu-Feng Wan
-
Article
| Open Access Orientation-invariant autoencoders learn robust representations for shape profiling of cells and organelles
Orientation-invariant autoencoders learn robust representations for shape profiling of cells and organelles
In image analysis, the shape properties of cells/organelles should be unaffected by image orientation. Conventional autoencoder (AE) methods can be sensitive to orientation. Here, the authors develop an unsupervised AE method that learns robust, orientation-invariant representations.
- James Burgess
- , Jeffrey J. Nirschl
- & Serena Yeung-Levy
-
Article
| Open Access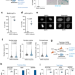 Detection of senescence using machine learning algorithms based on nuclear features
Detection of senescence using machine learning algorithms based on nuclear features
Identifying senescence is complicated by a lack of universal markers. Here, Duran et al. use nuclear morphology features to devise machine-learning classifiers that detect senescence in cell lines and liver sections of patients and mouse models of aging and disease.
- Imanol Duran
- , Joaquim Pombo
- & Jesús Gil
-
Article
| Open Access The impacts of active and self-supervised learning on efficient annotation of single-cell expression data
The impacts of active and self-supervised learning on efficient annotation of single-cell expression data
Cell type annotation for single-cell data is challenging. Here, authors explore active and self-supervised learning and introduce adaptive reweighting as a tailored heuristic, demonstrating competitive performance and showing that incorporating prior knowledge enhances cell type annotation accuracy.
- Michael J. Geuenich
- , Dae-won Gong
- & Kieran R. Campbell
-
Article
| Open Access SiFT: uncovering hidden biological processes by probabilistic filtering of single-cell data
SiFT: uncovering hidden biological processes by probabilistic filtering of single-cell data
Cells simultaneously encode multiple signals, some harder to recover. Here, authors introduce SiFT (Signal FilTering), a kernel-based projection method, revealing underlying biological processes in single-cell data.
- Zoe Piran
- & Mor Nitzan
-
Article
| Open Access Segment anything in medical images
Segment anything in medical images
Segmentation is an important fundamental task in medical image analysis. Here the authors show a deep learning model for efficient and accurate segmentation across a wide range of medical image modalities and anatomies.
- Jun Ma
- , Yuting He
- & Bo Wang
-
Article
| Open Access Using big sequencing data to identify chronic SARS-Coronavirus-2 infections
Using big sequencing data to identify chronic SARS-Coronavirus-2 infections
Chronic SARS-CoV-2 infections have been hypothesised to be sources of new variants. Here, the authors use large-scale genome sequencing data to identify mutations predictive of chronic infections, which may therefore be relevant in future variants.
- Sheri Harari
- , Danielle Miller
- & Adi Stern

 Prospective de novo drug design with deep interactome learning
Prospective de novo drug design with deep interactome learning