Article
|
Open Access
Featured
-
-
Article
| Open Access PRO-IP-seq tracks molecular modifications of engaged Pol II complexes at nucleotide resolution
PRO-IP-seq tracks molecular modifications of engaged Pol II complexes at nucleotide resolution
Tracking molecular modifications of engaged Pol II complexes is important for studying transcriptional regulation. Here, the authors combine run-on sequencing (PRO-seq) and immunoprecipitation, revealing dynamics of Pol II CTD phosphorylation at nucleotide-resolution.
- Anniina Vihervaara
- , Philip Versluis
- & John T. Lis
-
Article
| Open Access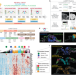 Maintenance of pluripotency-like signature in the entire ectoderm leads to neural crest stem cell potential
Maintenance of pluripotency-like signature in the entire ectoderm leads to neural crest stem cell potential
How the neural crest gains its pluripotency-like stem cell potential is unclear. Here, the authors show that the entire post-gastrula ectoderm maintains expression of pluripotency genes, leading to the high stem cell capacity in the neural crest.
- Ceren Pajanoja
- , Jenny Hsin
- & Laura Kerosuo
-
Article
| Open Access Single cell multiomic analysis reveals diabetes-associated β-cell heterogeneity driven by HNF1A
Single cell multiomic analysis reveals diabetes-associated β-cell heterogeneity driven by HNF1A
The mechanism and disease-relevance of pancreatic b-cell heterogeneity remains elusive. Here the authors show that variable HNF1A-FXYD2 activity drives single b-cell heterogeneity at transcriptomic, epigenomic, and electro-physiological levels, which strongly mark the progression of type 2 diabetes.
- Chen Weng
- , Anniya Gu
- & Yan Li
-
Article
| Open Access High-throughput and high-accuracy single-cell RNA isoform analysis using PacBio circular consensus sequencing
High-throughput and high-accuracy single-cell RNA isoform analysis using PacBio circular consensus sequencing
Long-read single-cell RNA isoform sequencing can elucidate the intricate landscape of alternative RNA splicing in individual cells, but it suffers from a low read throughput. Here, the authors develop circular consensus sequencing methods to allow high-throughput and high-accuracy single-cell RNA isoform sequencing.
- Zhuo-Xing Shi
- , Zhi-Chao Chen
- & Yi-Zhi Liu
-
Article
| Open Access RIViT-seq enables systematic identification of regulons of transcriptional machineries
RIViT-seq enables systematic identification of regulons of transcriptional machineries
Here the authors present their method ‘regulon identification by in vitro transcription-sequencing’ (RIViT-seq), which enables systematic identification of target genes of transcription factors of interest. They applied RIViT-seq to 13 sigma factors from Streptomyces coelicolor A3(2) and successfully identified target genes of 11 of these, expanding the regulatory characterisation in this organism.
- Hiroshi Otani
- & Nigel J. Mouncey
-
Article
| Open Access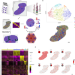 Spatial Transcriptomics to define transcriptional patterns of zonation and structural components in the mouse liver
Spatial Transcriptomics to define transcriptional patterns of zonation and structural components in the mouse liver
Global transcriptional differences across lobular units in the liver remain unknown. Here the authors perform spatial transcriptomics of liver tissue to delineate transcriptional differences in physical space, confirm lobular zonation along transcriptional gradients and suggest the presence of previously uncharacterized structures within liver tissue.
- Franziska Hildebrandt
- , Alma Andersson
- & Johan Ankarklev
-
Article
| Open Access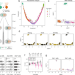 Integrated analysis of Xist upregulation and X-chromosome inactivation with single-cell and single-allele resolution
Integrated analysis of Xist upregulation and X-chromosome inactivation with single-cell and single-allele resolution
X-chromosome inactivation (XCI) ensures dosage compensation between the sexes. Here the authors perform allele-specific single-cell RNA sequencing in differentiating mouse embryonic stem cells to provide a detailed profile of the onset of XCI.
- Guido Pacini
- , Ilona Dunkel
- & Edda G. Schulz
-
Article
| Open Access Spatio-temporal mRNA tracking in the early zebrafish embryo
Spatio-temporal mRNA tracking in the early zebrafish embryo
Early stages of embryogenesis are known to depend on subcellular localization and transport of maternal mRNA, but systematic analyses have been hindered by a lack of methods for tracking of RNA. Here the authors combine spatially-resolved transcriptomics and single-cell RNA labeling to perform a spatio-temporal analysis of the transcriptome during early zebrafish development, revealing insights into this process.
- Karoline Holler
- , Anika Neuschulz
- & Jan Philipp Junker
-
Article
| Open Access Genome-wide binding potential and regulatory activity of the glucocorticoid receptor’s monomeric and dimeric forms
Genome-wide binding potential and regulatory activity of the glucocorticoid receptor’s monomeric and dimeric forms
Glucocorticoid receptors (GR) are thought to bind DNA as dimers or monomers, to regulate different transcription pathways. Here, the authors perform genome-wide studies on GRs with mutations that impair dimerization and provide evidence that monomeric GRs do not play a significant physiologic role.
- Thomas A. Johnson
- , Ville Paakinaho
- & Diego M. Presman
-
Article
| Open Access High throughput error corrected Nanopore single cell transcriptome sequencing
High throughput error corrected Nanopore single cell transcriptome sequencing
Droplet-based high throughput single cell sequencing techniques can often lose information on transcript splicing and heterogenity. Here the authors introduce ScNaUmi-seq, which uses Oxford Nanopore sequencing and barcoding to generate high accuracy full length sequences.
- Kevin Lebrigand
- , Virginie Magnone
- & Rainer Waldmann
-
Article
| Open Access Epigenome environment interactions accelerate epigenomic aging and unlock metabolically restricted epigenetic reprogramming in adulthood
Epigenome environment interactions accelerate epigenomic aging and unlock metabolically restricted epigenetic reprogramming in adulthood
Early life exposure to environmental stressors, including endocrine disrupting chemicals (EDCs), can impact health later in life. Here, the authors show that neonatal EDC exposure in rats causes epigenetic reprogramming in the liver, which is transcriptionally silent until animals are placed on a Western-style diet.
- Lindsey S. Treviño
- , Jianrong Dong
- & Cheryl Lyn Walker
-
Article
| Open Access Facultative dosage compensation of developmental genes on autosomes in Drosophila and mouse embryonic stem cells
Facultative dosage compensation of developmental genes on autosomes in Drosophila and mouse embryonic stem cells
In Drosophila the Male-Specific Lethal complex (MSLc) mediates upregulation of the single male X chromosome. Here the authors provide evidence that MSL2 also targets autosomal genes required for proper development and that MSL2 binds and similarly regulates mouse orthologues.
- Claudia Isabelle Keller Valsecchi
- , M. Felicia Basilicata
- & Asifa Akhtar
-
Article
| Open Access Sensitive and powerful single-cell RNA sequencing using mcSCRB-seq
Sensitive and powerful single-cell RNA sequencing using mcSCRB-seq
Single-cell RNA-barcoding and sequencing is an efficient, genome-wide method to characterize cellular identities. Here the authors systematically evaluate the protocol and develop molecular crowding SCRB-seq with improved sensitivity and cost-efficiency.
- Johannes W. Bagnoli
- , Christoph Ziegenhain
- & Wolfgang Enard
-
Article
| Open Access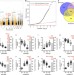 Genome-wide analysis of the genetic regulation of gene expression in human neutrophils
Genome-wide analysis of the genetic regulation of gene expression in human neutrophils
Neutrophils are abundant immune cells important for antimicrobial defence and in autoimmunity. Here, by mapping expression quantitative trait loci (eQTL) in neutrophils of Chinese ethnicity from Singapore, Andiappan et al.provide a resource for understanding immune-related trait associated genetic variants.
- Anand Kumar Andiappan
- , Rossella Melchiotti
- & Olaf Rotzschke
-
Article |
 Transcriptional refractoriness is dependent on core promoter architecture
Transcriptional refractoriness is dependent on core promoter architecture
Genes are often transcribed in random bursts followed by long periods of inactivity. Here the authors show, by a light-inducible transcription system in Neurospora, that refractory promoters carry a physical memory of their previous transcription history.
- François Cesbron
- , Michael Oehler
- & Michael Brunner
-
Article |
 Genome-wide identification of microRNA expression quantitative trait loci
Genome-wide identification of microRNA expression quantitative trait loci
As important post-transcriptional regulators of gene expression, microRNAs play a key role in the generation of complex phenotypes. Here, Huan et al.identify miR-eQTLs in whole blood samples to create a roadmap linking regulation of microRNA expression to complex diseases.
- Tianxiao Huan
- , Jian Rong
- & Jane E. Freedman

 Decoding the gene regulatory network of endosperm differentiation in maize
Decoding the gene regulatory network of endosperm differentiation in maize