Featured
-
-
Article
| Open Access Properties of structural variants and short tandem repeats associated with gene expression and complex traits
Properties of structural variants and short tandem repeats associated with gene expression and complex traits
Genetic variation associated with gene expression changes has mostly been studied in the context of single nucleotide variants. Here, Jakubosky et al. report eQTL analysis of structural variants and short tandem repeats and find properties, such as length of variation, that affect the association.
- David Jakubosky
- , Matteo D’Antonio
- & Kelly A. Frazer
-
Article
| Open Access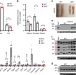 Eosinophil function in adipose tissue is regulated by Krüppel-like factor 3 (KLF3)
Eosinophil function in adipose tissue is regulated by Krüppel-like factor 3 (KLF3)
Immune cells are important regulators of adipose tissue function, including adaptive thermogenesis. Here the authors show that mice with Krüppel-like factor 3 (KLF3) deficiency in bone marrow-derived cells have increased adipose tissue beiging which may at least in part be due to altered eosinophil paracrine signaling.
- Alexander J. Knights
- , Emily J. Vohralik
- & Kate G. R. Quinlan
-
Article
| Open Access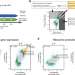 A multi-omics analysis reveals the unfolded protein response regulon and stress-induced resistance to folate-based antimetabolites
A multi-omics analysis reveals the unfolded protein response regulon and stress-induced resistance to folate-based antimetabolites
The unfolded protein response (UPR) is a stress response pathway implicated in numerous diseases and chemotherapy resistance. Here, the authors define the UPR regulon with a multi-omics strategy, uncovering changes to mitochondrial one-carbon metabolism and concomitant resistance to folate-based therapeutics.
- Stefan Reich
- , Chi D. L. Nguyen
- & Jan Medenbach
-
Article
| Open Access Dynamics of the 4D genome during in vivo lineage specification and differentiation
Dynamics of the 4D genome during in vivo lineage specification and differentiation
The relationship between regulatory elements, chromatin interactions and gene expression during development remains poorly understood. Here the authors present Tiled-C, a low-input 3C approach to study genome architecture at high resolution, and apply it to mouse erythroid differentiation in vivo, finding that enhancer-promoter interactions are formed gradually during differentiation, concomitant with progressive upregulation of gene activity.
- A. Marieke Oudelaar
- , Robert A. Beagrie
- & Jim R. Hughes
-
Article
| Open Access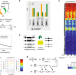 A genome-scale map of DNA methylation turnover identifies site-specific dependencies of DNMT and TET activity
A genome-scale map of DNA methylation turnover identifies site-specific dependencies of DNMT and TET activity
Local activity of the DNA methylation machinery remains poorly understood. Here, the authors present a theoretical and experimental framework to infer methylation and demethylation rates at genome scale in mouse embryonic stem cells, finding that maintenance methylation activity is reduced at transcription factor binding sites, while methylation turnover is elevated in transcribed gene bodies.
- Paul Adrian Ginno
- , Dimos Gaidatzis
- & Dirk Schübeler
-
Article
| Open Access Methylation of a CGATA element inhibits binding and regulation by GATA-1
Methylation of a CGATA element inhibits binding and regulation by GATA-1
While DNA methylation is thought to play a regulatory role, there are few examples where modification of a single CpG dinucleotide directly affects transcription factor binding. Here the authors show that methylation of a single CGATA element within the c-Kit gene inhibits binding and regulation by erythroid transcription factor GATA-1, both in cells and in mice, suggesting that methylation at this site plays an essential role in erythropoiesis.
- Lu Yang
- , Zhiliang Chen
- & Merlin Crossley
-
Article
| Open Access HOX13-dependent chromatin accessibility underlies the transition towards the digit development program
HOX13-dependent chromatin accessibility underlies the transition towards the digit development program
Pioneer factors direct cell fate through switching inaccessible chromatin to an accessible state at specific target enhancers. Here the authors show that HOX13 transcription factors have a pioneer activity which is required for the proper implementation of the distal limb developmental program.
- Ines Desanlis
- , Yacine Kherdjemil
- & Marie Kmita
-
Article
| Open Access Determinants of transcription factor regulatory range
Determinants of transcription factor regulatory range
Characterization of the distance over which TF binding influences gene expression is important for inferring target genes. Here the authors study this relationship using thousands of genomic data sets, finding two classes of TFs with distinct ranges of regulatory influence modulated by chromatin states of topologically associated domains.
- Chen-Hao Chen
- , Rongbin Zheng
- & X. Shirley Liu
-
Article
| Open Access Intron retention is a hallmark and spliceosome represents a therapeutic vulnerability in aggressive prostate cancer
Intron retention is a hallmark and spliceosome represents a therapeutic vulnerability in aggressive prostate cancer
Dysregulation of mRNA alternative splicing is prevalent in cancers. Here, the authors characterized the landscape of aberrant alternative splicing during the development of prostate cancer, progression and therapeutic resistance and show that splicing modulator, E7107, reduces growth in castration-resistant prostate cancer.
- Dingxiao Zhang
- , Qiang Hu
- & Dean G. Tang
-
Article
| Open Access Tunable genetic devices through simultaneous control of transcription and translation
Tunable genetic devices through simultaneous control of transcription and translation
Synthetic genetic circuits are sensitive to their environment and host cell, requiring many rounds of physical reassembly to achieve a desired function. Here the authors use a multi-level regulatory motif to dynamically tune the function of genetic parts as a step towards robust adaptive circuits.
- Vittorio Bartoli
- , Grace A. Meaker
- & Thomas E. Gorochowski
-
Article
| Open Access A high-content RNAi screen reveals multiple roles for long noncoding RNAs in cell division
A high-content RNAi screen reveals multiple roles for long noncoding RNAs in cell division
Long noncoding RNAs (lncRNAs) regulate key steps of cell division. Here, the authors perform a comprehensive RNAi imaging screen targeting more than 2,000 human lncRNAs, and suggest a role of chromatin-associated linc00899 in regulation of cell division by suppressing the transcription of microtubule-binding protein TPPP/p25.
- Lovorka Stojic
- , Aaron T. L. Lun
- & Fanni Gergely
-
Article
| Open Access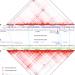 Transposable elements contribute to cell and species-specific chromatin looping and gene regulation in mammalian genomes
Transposable elements contribute to cell and species-specific chromatin looping and gene regulation in mammalian genomes
A fraction of mammalian CTCF binding sites fall within transposable elements (TEs) but their contribution to the evolution of 3D chromatin structure is unknown. Here the authors investigate the effect of TE-driven CTCF binding site expansions on chromatin looping in humans and mice, and provide evidence that TEs contribute to cell-specific and species-specific chromatin looping diversity and variable gene regulation in mammalian genomes.
- Adam G. Diehl
- , Ningxin Ouyang
- & Alan P. Boyle
-
Article
| Open Access An acid-tolerance response system protecting exponentially growing Escherichia coli
An acid-tolerance response system protecting exponentially growing Escherichia coli
The ability to grow at acidic pH is crucial for E. coli colonization of the host’s intestine. Here, the authors identify an acid-tolerance response system that is important for E. coli exponential growth at pH 4.2, survival in the mouse intestine, and production of 3-hydroxypropionate during fermentation.
- Ying Xu
- , Zhe Zhao
- & Guang Zhao
-
Article
| Open Access Dissecting the role of PfAP2-G in malaria gametocytogenesis
Dissecting the role of PfAP2-G in malaria gametocytogenesis
The transcription factor PfAP2-G is a key determinant of sexual commitment in Plasmodium falciparum. Here, Josling et al. define the transcriptional regulatory network of PfAP2-G by identifying its DNA binding sites genome-wide, which vary depending on the route of sexual conversion and rely on interactions with the PfAP2-I transcription factor.
- Gabrielle A. Josling
- , Timothy J. Russell
- & Manuel Llinás
-
Article
| Open Access Candidate silencer elements for the human and mouse genomes
Candidate silencer elements for the human and mouse genomes
Identification of silencer elements by computational or experimental approaches in a genome-wide manner is still challenging. Here authors define uncharacterized cis-regulatory elements (CREs) in human and mouse genomes likely containing silencer elements, and test them in cells using massively parallel reporter assays to identify silencer elements that showed silencer activity.
- Naresh Doni Jayavelu
- , Ajay Jajodia
- & R. David Hawkins
-
Article
| Open Access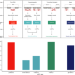 Regulatory sites for splicing in human basal ganglia are enriched for disease-relevant information
Regulatory sites for splicing in human basal ganglia are enriched for disease-relevant information
Regulation of gene expression and splicing are thought to be tissue-specific. Here, the authors obtain genomic and transcriptomic data from putamen and substantia nigra of 117 neurologically healthy human brains and find that splicing eQTLs are enriched for neuron-specific regulatory information.
- Sebastian Guelfi
- , Karishma D’Sa
- & Mina Ryten
-
Article
| Open Access Altered chromatin landscape and enhancer engagement underlie transcriptional dysregulation in MED12 mutant uterine leiomyomas
Altered chromatin landscape and enhancer engagement underlie transcriptional dysregulation in MED12 mutant uterine leiomyomas
Somatic mutations in MED12 have been implicated as the causal genetic lesion in the majority of uterine leiomyomas. Here, the authors profile the chromatin landscape of matched normal and leiomyoma tissues and find that changes in enhancer acetylation, enhancer-promoter interaction strength, differential enhancer usage and transcription factor AP-1 occupancy are significant drivers of transcriptional dysregulation in MED12 mutant leiomyomas.
- Mthabisi B. Moyo
- , J. Brandon Parker
- & Debabrata Chakravarti
-
Article
| Open Access Jouvence a small nucleolar RNA required in the gut extends lifespan in Drosophila
Jouvence a small nucleolar RNA required in the gut extends lifespan in Drosophila
Small non-coding RNAs contribute to the regulation of aging. Here the authors identify a small nucleolar RNA, the snoRNA jouvence, which extends the lifespan of fruit flies through its function in the gut, and is conserved in humans.
- Stéphanie Soulé
- , Lucille Mellottée
- & Jean-René Martin
-
Article
| Open Access Alterations in promoter interaction landscape and transcriptional network underlying metabolic adaptation to diet
Alterations in promoter interaction landscape and transcriptional network underlying metabolic adaptation to diet
Metabolic adaptation to different diets results in changes to gene expression. Here, the authors characterise the chromatin landscape and transcriptional network in mice on a diet of high saturated fat, compared to a diet high in carbohydrate, finding a dramatic reprogramming of the liver transcriptional network.
- Yufeng Qin
- , Sara A. Grimm
- & Paul A. Wade
-
Article
| Open Access A lncRNA-SWI/SNF complex crosstalk controls transcriptional activation at specific promoter regions
A lncRNA-SWI/SNF complex crosstalk controls transcriptional activation at specific promoter regions
SWI/SNF complexes regulate chromatin architecture and gene expression. Here the authors report the RNA interactome of SMARCB1-containing SWI/SNF complexes, showing the function of SMARCB1-interacting long noncoding RNA SWINGN in transcriptional activation of GAS6 and a set of SWI/SNF target genes.
- Elena Grossi
- , Ivan Raimondi
- & Maite Huarte
-
Article
| Open Access In situ dissection of domain boundaries affect genome topology and gene transcription in Drosophila
In situ dissection of domain boundaries affect genome topology and gene transcription in Drosophila
In Drosophila, transcription is thought to be required for TAD formation, while the role of architectural proteins remains controversial. Here, the authors find that deletion of domain boundaries at the fly Notch locus results in TAD fusion and long-range topological defects, loss of architectural protein and RNA Pol II chromatin binding, and transcription defects.
- Rodrigo G. Arzate-Mejía
- , Angel Josué Cerecedo-Castillo
- & Félix Recillas-Targa
-
Article
| Open Access mTORC1 coordinates an immediate unfolded protein response-related transcriptome in activated B cells preceding antibody secretion
mTORC1 coordinates an immediate unfolded protein response-related transcriptome in activated B cells preceding antibody secretion
Antibody production in plasma cells involves the unfold protein response (UPR), but how this is regulated is not clear. Here the authors show that mTORC1 signalling but not Xbp1-mediated transcription regulation in activated B cells is important for the induction of a UPR-related transcriptome that precedes full plasma cell functions.
- Brian T. Gaudette
- , Derek D. Jones
- & David Allman
-
Article
| Open Access Mechanisms governing the pioneering and redistribution capabilities of the non-classical pioneer PU.1
Mechanisms governing the pioneering and redistribution capabilities of the non-classical pioneer PU.1
PU.1 is a master TF of hematopoietic lineage differentiation. Here the authors analyse properties of PU.1 DNA-binding in vitro and genome-wide in vivo across different cell types with native or ectopic PU.1 expression, and uncover the mechanisms governing the pioneering and redistribution capabilities of the non-classical pioneer PU.1.
- Julia Minderjahn
- , Andreas Schmidt
- & Michael Rehli
-
Article
| Open Access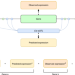 Using regulatory variants to detect gene–gene interactions identifies networks of genes linked to cell immortalisation
Using regulatory variants to detect gene–gene interactions identifies networks of genes linked to cell immortalisation
For most human genes nearby regulatory variants explain only a small proportion of their expression variation between individuals. Here the authors show how the impact of a gene’s set of nearby regulatory variant is often linked to the expression levels of distal genes, providing insights into gene networks.
- D. Wragg
- , Q. Liu
- & J. G. D. Prendergast
-
Article
| Open Access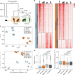 Hominin-specific regulatory elements selectively emerged in oligodendrocytes and are disrupted in autism patients
Hominin-specific regulatory elements selectively emerged in oligodendrocytes and are disrupted in autism patients
The understanding of the changes regulating gene expression relevant for the emergence of the human brain and its susceptibility to disease is limited. Here, the authors identified a set of regulatory elements that evolved in hominins affecting oligodendrocyte function, and link these to autism.
- Bas Castelijns
- , Mirna L. Baak
- & Menno P. Creyghton
-
Article
| Open Access Dual-initiation promoters with intertwined canonical and TCT/TOP transcription start sites diversify transcript processing
Dual-initiation promoters with intertwined canonical and TCT/TOP transcription start sites diversify transcript processing
The functional significance of start site choice in promoter architectures is little understood. Here the authors identify in zebrafish development and mammalian cells a class of dual-initiation promoters, in which non-canonical YC dinucleotides reflecting 5’-TOP/TCT initiation are intertwined with canonical YR-initiation.
- Chirag Nepal
- , Yavor Hadzhiev
- & Ferenc Müller
-
Article
| Open Access Mechanistic insights into transcription factor cooperativity and its impact on protein-phenotype interactions
Mechanistic insights into transcription factor cooperativity and its impact on protein-phenotype interactions
Although transcription factor (TF) cooperativity is widespread, a global mechanistic understanding of the role of TF cooperativity is still lacking. Here the authors introduce a statistical learning framework that provides structural insight into TF cooperativity and its functional consequences based on next generation sequencing data and provide mechanistic insights into TF cooperativity and its impact on protein-phenotype interactions.
- Ignacio L. Ibarra
- , Nele M. Hollmann
- & Judith B. Zaugg
-
Article
| Open Access Chaperone biomarkers of lifespan and penetrance track the dosages of many other proteins
Chaperone biomarkers of lifespan and penetrance track the dosages of many other proteins
In C. elegans the chaperone hsp-16.2 predicts the penetrance of mutations and lifespan after heat shock, but it is not known why cells express different amounts of hsp-16.2. Here the authors show hsp-16.2 tracks differences in global gene expression capacity, rather than differences in signaling or intrinsic noise in adult intestine cells.
- Nikolay Burnaevskiy
- , Bryan Sands
- & Alexander Mendenhall
-
Article
| Open Access The epigenomic landscape of transposable elements across normal human development and anatomy
The epigenomic landscape of transposable elements across normal human development and anatomy
Although most are silenced, certain transposable elements (TEs) have been co-opted by the host. Here, the authors quantify the epigenomic status of TEs using data from the Roadmap Epigenomics Project, provide a systematic profile of TE activity across normal human tissues and development, finding that TEs encompass a quarter of the human regulatory epigenome, with 47% of TEs in regulatory states.
- Erica C. Pehrsson
- , Mayank N. K. Choudhary
- & Ting Wang
-
Article
| Open Access Glycogen branching enzyme controls cellular iron homeostasis via Iron Regulatory Protein 1 and mitoNEET
Glycogen branching enzyme controls cellular iron homeostasis via Iron Regulatory Protein 1 and mitoNEET
Higher organisms regulate cellular iron concentrations through Iron Regulatory Proteins (IRPs), which regulate specific messenger RNAs. Here Huynh et al. show that IRP1 requires a Glycogen Branching Enzyme for proper function, and that IRP1 has additional regulatory roles in cell nuclei.
- Nhan Huynh
- , Qiuxiang Ou
- & Kirst King-Jones
-
Article
| Open Access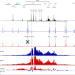 A revised model for promoter competition based on multi-way chromatin interactions at the α-globin locus
A revised model for promoter competition based on multi-way chromatin interactions at the α-globin locus
The coordination of interactions between multiple regulatory elements and genes within a chromatin domain remains poorly understood. Here, the authors use a method to detect multi-way chromatin interactions in a mouse model in which the α-globin domain is extended to include several additional genes, finding that the promoters do not form mutually exclusive interactions with the enhancers, but all interact simultaneously in a single complex.
- A. Marieke Oudelaar
- , Caroline L. Harrold
- & Jim R. Hughes
-
Article
| Open Access Global chromatin conformation differences in the Drosophila dosage compensated chromosome X
Global chromatin conformation differences in the Drosophila dosage compensated chromosome X
In Drosophila, dosage compensation involves a twofold transcriptional upregulation of the single male chromosome X. Here the authors show that global conformational differences are specifically present in the male X chromosome and detectable using Hi-C data, indicating that dosage compensation affects global chromosome structure.
- Koustav Pal
- , Mattia Forcato
- & Francesco Ferrari
-
Article
| Open Access RUNX1 maintains the identity of the fetal ovary through an interplay with FOXL2
RUNX1 maintains the identity of the fetal ovary through an interplay with FOXL2
Proper ovarian differentiation requires RUNX1. Here, the authors show that double knockout of Runx1/Foxl2 results in masculinization of fetal ovaries, and that RUNX1 and FOXL2 jointly occupy common chromatin regions to maintain pre-granulosa cell identity in the fetal ovary.
- Barbara Nicol
- , Sara A. Grimm
- & Humphrey H.-C. Yao
-
Article
| Open Access Regulation of CHD2 expression by the Chaserr long noncoding RNA gene is essential for viability
Regulation of CHD2 expression by the Chaserr long noncoding RNA gene is essential for viability
The conserved long noncoding RNA Chaserr is transcribed upstream of the chromatin remodeler Chd2. Here, using mouse genetics and high throughput assays, the authors show that Chaserr inhibits expression of Chd2 in cis and is required for postnatal mouse development.
- Aviv Rom
- , Liliya Melamed
- & Igor Ulitsky
-
Article
| Open Access An enhancer cluster controls gene activity and topology of the SCN5A-SCN10A locus in vivo
An enhancer cluster controls gene activity and topology of the SCN5A-SCN10A locus in vivo
Mutations associated with SCN5A, which encodes the major cardiac sodium channel, influence impulse conduction and are associated with arrhythmia disorders. Here, the authors identify an evolutionary conserved, super enhancer-like, regulatory cluster downstream of SCN5A and show that it controls the chromatin architecture of the locus and Scn5a expression.
- Joyce C. K. Man
- , Rajiv A. Mohan
- & Vincent M. Christoffels
-
Article
| Open Access NSD2 overexpression drives clustered chromatin and transcriptional changes in a subset of insulated domains
NSD2 overexpression drives clustered chromatin and transcriptional changes in a subset of insulated domains
In multiple myeloma, a 4;14 translocation induces overexpression of histone methyltransferase NSD2, resulting in expansion of H3K36me2 and shrinkage of H3K27me3 domains. Here the authors find that CTCF, H3K27ac and gene expression changes cluster within a subset of insulated domains implicating 3D chromosome organization as a key factor in the NSD2-mediated phenotype.
- Priscillia Lhoumaud
- , Sana Badri
- & Jane A. Skok
-
Article
| Open Access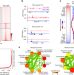 Super-enhancer-guided mapping of regulatory networks controlling mouse trophoblast stem cells
Super-enhancer-guided mapping of regulatory networks controlling mouse trophoblast stem cells
Trophectoderm lineage development is essential for implantation, placentation, and healthy pregnancy. Here the authors map super-enhancers (SEs) in trophoblast stem cells and find both TE-specific master regulators and 150 previous uncharacterised transcription factors that are SE-associated, providing insight into trophectoderm-specific regulatory networks.
- Bum-Kyu Lee
- , Yu jin Jang
- & Jonghwan Kim
-
Article
| Open Access Identification of atrial fibrillation associated genes and functional non-coding variants
Identification of atrial fibrillation associated genes and functional non-coding variants
The majority of disease-associated genetic variants lie in non-coding regions. Here the authors generated and compiled human transcriptomic, epigenomic and chromatin conformation datasets, to identify genes associated with atrial fibrillation and functional non-coding variants.
- Antoinette F. van Ouwerkerk
- , Fernanda M. Bosada
- & Vincent M. Christoffels
-
Article
| Open Access Dissecting splicing decisions and cell-to-cell variability with designed sequence libraries
Dissecting splicing decisions and cell-to-cell variability with designed sequence libraries
Alternative splicing is regulated by multiple mechanisms. Here the authors employed designed splice site libraries and massively parallel reporter assays to dissect the regulatory complexity and cell-to-cell variability of splicing decisions and to build accurate predictive models.
- Martin Mikl
- , Amit Hamburg
- & Eran Segal
-
Article
| Open Access Single cell transcriptome in aneuploidies reveals mechanisms of gene dosage imbalance
Single cell transcriptome in aneuploidies reveals mechanisms of gene dosage imbalance
Gene dosage anomalies such as those caused by aneuploidy underlie diseases including Down syndrome. Here, the authors perform allele-specific single cell transcriptome analysis to investigate the mechanisms of gene dosage imbalance in fibroblasts with trisomies T21, T18, T13 and T8.
- Georgios Stamoulis
- , Marco Garieri
- & Stylianos E. Antonarakis
-
Article
| Open Access BORDER proteins protect expression of neighboring genes by promoting 3′ Pol II pausing in plants
BORDER proteins protect expression of neighboring genes by promoting 3′ Pol II pausing in plants
In plants, 3′ Pol II pausing helps prevent transcriptional interference. Here the authors provide evidence that BORDER proteins are enriched in intergenic regions and inhibit transcriptional interference between closely spaced genes by preventing RNA polymerase from intruding into the promoters of nearby downstream genes.
- Xuhong Yu
- , Pascal G. P. Martin
- & Scott D. Michaels
-
Article
| Open Access Tet inactivation disrupts YY1 binding and long-range chromatin interactions during embryonic heart development
Tet inactivation disrupts YY1 binding and long-range chromatin interactions during embryonic heart development
Tet-mediated DNA demethylation is intimately involved in reguatling embryonic development. Here the authors characterise DNA methylation and hydroxymethylation dynamics during early cardiac development in both human and mice and provide evidence that Tet-mediated DNA demethylation plays a role in regulating chromatin organization during early heart development.
- Shaohai Fang
- , Jia Li
- & Yun Huang
-
Article
| Open Access Transcription-dependent targeting of Hda1C to hyperactive genes mediates H4-specific deacetylation in yeast
Transcription-dependent targeting of Hda1C to hyperactive genes mediates H4-specific deacetylation in yeast
In yeast, Hda1 histone deacetylase complex (Hda1C) preferentially deacetylates H3 and H2B, but has also been implicated in global H4 deacetylation. Here, the authors provide evidence that Hda1C binds to hyperactive genes, where it specifically deacetylates H4 to partially inhibit elongation, suggesting a role for Hda1C in elongation by specifically deacetylating H4 at highly transcribed regions.
- So Dam Ha
- , Seokjin Ham
- & TaeSoo Kim
-
Article
| Open Access Epigenome editing strategies for the functional annotation of CTCF insulators
Epigenome editing strategies for the functional annotation of CTCF insulators
The role of CTCF-bound insulator elements in enhancer-gene interactions and transcriptional regulation remains poorly understood. Here, the authors investigate multiple epigenome editing strategies for perturbing individual CTCF-bound insulators, and evaluate their effects on genome topology and transcription.
- Daniel R. Tarjan
- , William A. Flavahan
- & Bradley E. Bernstein
-
Article
| Open Access UPF1/SMG7-dependent microRNA-mediated gene regulation
UPF1/SMG7-dependent microRNA-mediated gene regulation
UPF1 mediates the decay of target mRNA in a 3′ untranslated region (UTR)-length-dependent manner. Here the authors reveal that the 3′UTR-length-dependent regulation of UPF1-dependent mRNA decay occurs through EJC-independent but miRNA-dependent regulation.
- Jungyun Park
- , Jwa-Won Seo
- & Jin-Wu Nam
-
Article
| Open Access A high-resolution 3D epigenomic map reveals insights into the creation of the prostate cancer transcriptome
A high-resolution 3D epigenomic map reveals insights into the creation of the prostate cancer transcriptome
In prostate cancer, chromatin structure can impact the transcriptome. Here, the authors develop high resolution chromatin interaction maps in prostate cancer cells using in situ Hi-C, revealing prostate cancer-specific TADs and enhancer-promoter loops surrounding the androgen receptor (AR) locus.
- Suhn Kyong Rhie
- , Andrew A. Perez
- & Peggy J. Farnham
-
Article
| Open Access Engineered CRISPRa enables programmable eukaryote-like gene activation in bacteria
Engineered CRISPRa enables programmable eukaryote-like gene activation in bacteria
CRISPR activation strategies in bacteria are limited due to the reliance on σ70 promoters. Here the authors demonstrate eukaryote-like gene activation with high dynamic ranges using σ54- dependent promoters.
- Yang Liu
- , Xinyi Wan
- & Baojun Wang
-
Article
| Open Access EZHIP constrains Polycomb Repressive Complex 2 activity in germ cells
EZHIP constrains Polycomb Repressive Complex 2 activity in germ cells
Polycomb Repressive Complex 2 (PRC2) plays critical roles in transcriptional silencing during development. Here the authors identify EZHIP as a cofactor of PRC2 expressed predominantly in the gonads, finding that EZHIP limits the enzymatic activity of PRC2 in germ cells in mice.
- Roberta Ragazzini
- , Raquel Pérez-Palacios
- & Raphaël Margueron
-
Article
| Open Access Pioneer and nonpioneer factor cooperation drives lineage specific chromatin opening
Pioneer and nonpioneer factor cooperation drives lineage specific chromatin opening
Pioneer transcription factor Pax7 specifies melanotrope cells, which then allows for the binding of Tpit transcription factor. Here, authors find that while binding of heterochromatin targeting by Pax7 is independent of Tpit, Pax7-dependent chromatin opening requires Tpit.
- Alexandre Mayran
- , Kevin Sochodolsky
- & Jacques Drouin

 Identification of distinct loci for de novo DNA methylation by DNMT3A and DNMT3B during mammalian development
Identification of distinct loci for de novo DNA methylation by DNMT3A and DNMT3B during mammalian development