Article
|
Open Access
Featured
-
-
Article
| Open Access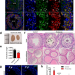 The m6A reader PRRC2A is essential for meiosis I completion during spermatogenesis
The m6A reader PRRC2A is essential for meiosis I completion during spermatogenesis
Modification of RNA with m6A has been shown to be important during spermatogenesis. Here they identify post-transcriptional functions of PRRC2A, showing it promotes transcriptome transition from spermatogonia to spermatocytes and the translation of genes related to cell division.
- Xinshui Tan
- , Caihong Zheng
- & Fengchao Wang
-
Article
| Open Access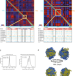 A DNA tumor virus globally reprograms host 3D genome architecture to achieve immortal growth
A DNA tumor virus globally reprograms host 3D genome architecture to achieve immortal growth
The dynamic and temporal changes of host genome architecture during Epstein-Barr virus (EBV) transformation are not well known. Here the authors transform human primary B lymphocyte into lymphoblastoid cell lines (LCLs) with EBV and show that the host 3D genome is rewired to facilitate expression of key oncogenes.
- Chong Wang
- , Xiang Liu
- & Bo Zhao
-
Article
| Open Access Histone H2A monoubiquitination marks are targeted to specific sites by cohesin subunits in Arabidopsis
Histone H2A monoubiquitination marks are targeted to specific sites by cohesin subunits in Arabidopsis
How histone H2A monoubiquitination (H2Aub1) is established at specific genomic locations remains unclear. Here, the authors report that Arabidopsis cohesin subunits SCC3 and SYN4 are involved in H2Aub1 through their direct or indirect interaction with BMI1A/B/C subunits of PRC1, the E3 ligases in PRC1 for H2Aub1.
- Yu Zhang
- , Min Ma
- & Yuda Fang
-
Article
| Open Access Sequence terminus dependent PCR for site-specific mutation and modification detection
Sequence terminus dependent PCR for site-specific mutation and modification detection
Rapid and facile detection of specific nucleic acid modifications could have numerous applications. Here the authors present Specific Terminal Mediated Polymerase Chain Reaction (STEM-PCR) as a generic and accessible approach, and demonstrate proof-of-principle cancer biomarker detection.
- Gaolian Xu
- , Hao Yang
- & Hongchen Gu
-
Article
| Open Access BdLT-Seq as a barcode decay-based method to unravel lineage-linked transcriptome plasticity
BdLT-Seq as a barcode decay-based method to unravel lineage-linked transcriptome plasticity
Cellular plasticity is a core biological process; however, observing diversity in non-genetic inheritance and the resulting phenotypic outputs, is challenging. Here the authors develop a non-genetically based tracing technology which can be used to reveal lineage-linked transcriptome plasticity.
- Yelyzaveta Shlyakhtina
- , Bianca Bloechl
- & Maximiliano M. Portal
-
Article
| Open Access Differential regulation of mRNA stability modulates transcriptional memory and facilitates environmental adaptation
Differential regulation of mRNA stability modulates transcriptional memory and facilitates environmental adaptation
Transcriptional memory is key for cellular adaptation. Here the authors show that differences in mRNA stability and mRNA degradation machinery between naïve and primed cells facilitate faster gene expression response to repeated stimuli.
- Bingnan Li
- , Patrice Zeis
- & Vicent Pelechano
-
Article
| Open Access A CpG island-encoded mechanism protects genes from premature transcription termination
A CpG island-encoded mechanism protects genes from premature transcription termination
Here the authors discover that SET1 complexes function as transcription anti-termination factors that bind to CpG islands and protect low to moderately transcribed genes from the pervasive termination activity of the ZC3H4 complex.
- Amy L. Hughes
- , Aleksander T. Szczurek
- & Robert J. Klose
-
Article
| Open Access Antecedent chromatin organization determines cGAS recruitment to ruptured micronuclei
Antecedent chromatin organization determines cGAS recruitment to ruptured micronuclei
DNA damage-induced micronuclei are linked to downstream viral signalling through the cGAS pattern recognition receptor. Here, the authors identify features of micronuclei chromatin that determine cGAS-MN recruitment and associated pathway activation.
- Kate M. MacDonald
- , Shirony Nicholson-Puthenveedu
- & Shane M. Harding
-
Article
| Open Access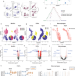 Transcriptional reprogramming of skeletal muscle stem cells by the niche environment
Transcriptional reprogramming of skeletal muscle stem cells by the niche environment
Aging leads to significant alteration in the gene expression of muscle stem cells. In vivo exposure of muscle stem cells from aged mice to a young niche environment restores the expression of a significant portion of age-altered genes in mice.
- Felicia Lazure
- , Rick Farouni
- & Vahab D. Soleimani
-
Article
| Open Access Epigenetic regulation of Neuregulin 1 promotes breast cancer progression associated to hyperglycemia
Epigenetic regulation of Neuregulin 1 promotes breast cancer progression associated to hyperglycemia
Despite hyperglycemia has been associated to breast cancer, the underlying mechanisms are not completely understood. Here, the authors show that epigenetic regulation of Nrg1 gene during hyperglycemia promotes breast cancer development.
- Changhu Lee
- , Min Kim
- & Jiyoung Park
-
Article
| Open Access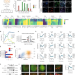 Epigenetic and transcriptional regulations prime cell fate before division during human pluripotent stem cell differentiation
Epigenetic and transcriptional regulations prime cell fate before division during human pluripotent stem cell differentiation
Many stem cells exhibit cell division coupled to differentiation, though the changes occurring between consecutive cell divisions have been difficult to study. Here they use synchronized hPSC culture to show that production of transcription factors and epigenetic changes are linked with cell division timing.
- Pedro Madrigal
- , Siwei Deng
- & Siim Pauklin
-
Article
| Open Access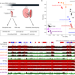 Mechanisms and function of de novo DNA methylation in placental development reveals an essential role for DNMT3B
Mechanisms and function of de novo DNA methylation in placental development reveals an essential role for DNMT3B
DNA methylation is a repressive modification that is essential for development. Here the authors reveal a critical role for DNA methylation in placental development during pregnancy. Failure to properly establish placental DNA methylation patterns compromises not only placental function, but embryo survival.
- Simon Andrews
- , Christel Krueger
- & Courtney W. Hanna
-
Article
| Open Access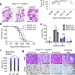 Polycomb deficiency drives a FOXP2-high aggressive state targetable by epigenetic inhibitors
Polycomb deficiency drives a FOXP2-high aggressive state targetable by epigenetic inhibitors
Delineating the specific role of Polycomb Repressive Complex 2 (PRC2) in various cancer systems is desirable as inhibitors for EZH2 inhibitors are approved for some cancers. Here the authors show haplo- and full-insufficiency of EZH2 drive divergent phenotypes in lung cancer. 3D tumoroids recapitulate transcriptional profiles, including FOXP2 derepression, and drug responses of in vivo tumors.
- Fan Chen
- , Aria L. Byrd
- & Christine Fillmore Brainson
-
Article
| Open Access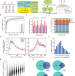 scm6A-seq reveals single-cell landscapes of the dynamic m6A during oocyte maturation and early embryonic development
scm6A-seq reveals single-cell landscapes of the dynamic m6A during oocyte maturation and early embryonic development
Modification of RNA with N6-methyladenosine can regulate RNA metabolism. Here they developed scm6A-seq to profile the methylome and transcriptome in single cells, and reveal the functions of m6A modification during oocyte maturation and early embryo development.
- Huan Yao
- , Chun-Chun Gao
- & Yun-Gui Yang
-
Article
| Open Access Molecular characterization of Richter syndrome identifies de novo diffuse large B-cell lymphomas with poor prognosis
Molecular characterization of Richter syndrome identifies de novo diffuse large B-cell lymphomas with poor prognosis
Richter syndrome (RS) is the transformation of chronic lymphocytic leukaemia (CLL) into aggressive lymphoma, in most cases diffuse large B-cell lymphoma (DLBCL). Here, the authors characterize the DNA methylation and transcriptomic profiles of RS samples, find a clonally-related CLL epigenetic imprint, and develop classifiers for “RS-type” de novo DLBCLs.
- Julien Broséus
- , Sébastien Hergalant
- & Stephan Stilgenbauer
-
Article
| Open Access Targeted systematic evolution of an RNA platform neutralizing DNMT1 function and controlling DNA methylation
Targeted systematic evolution of an RNA platform neutralizing DNMT1 function and controlling DNA methylation
Here the authors generate an RNA-based platform to neutralize the major epigenetic player DNMT1. Using this targeted approach, aberrant DNA methylation in cancer can be corrected.
- Carla L. Esposito
- , Ida Autiero
- & Annalisa Di Ruscio
-
Article
| Open Access Engineering inducible biomolecular assemblies for genome imaging and manipulation in living cells
Engineering inducible biomolecular assemblies for genome imaging and manipulation in living cells
Imaging non-repetitive loci in living cells remains challenging. Here, the authors engineered an inducible system whereby biomolecular assemblies can be guided to specific genomic loci by a nuclease-defective Cas9, allowing the simultaneous imaging and manipulation of the loci.
- Qin Peng
- , Ziliang Huang
- & Yingxiao Wang
-
Article
| Open Access Epigenome-wide meta-analysis identifies DNA methylation biomarkers associated with diabetic kidney disease
Epigenome-wide meta-analysis identifies DNA methylation biomarkers associated with diabetic kidney disease
Approximately 40 percent of people with type 1 diabetes develop kidney disease, but the risk factors are not well understood. Here, the authors identify DNA methylation signatures associated with diabetic kidney disease, of which 21 biomarkers predict the development of kidney failure.
- Laura J. Smyth
- , Emma H. Dahlström
- & Amy Jayne McKnight
-
Article
| Open Access Cell-type specific profiling of histone post-translational modifications in the adult mouse striatum
Cell-type specific profiling of histone post-translational modifications in the adult mouse striatum
Bulk or pooled epigenomic profiling in the heterogenous brain obscures cell-type-specificity and individual subject variability in gene regulation. Here the authors optimized a hybrid protocol, ICuRuS, to profile epigenetic features in neuronal subtypes from a single mouse.
- Marco D. Carpenter
- , Delaney K. Fischer
- & Elizabeth A. Heller
-
Article
| Open Access Histone H3.3 deposition in seed is essential for the post-embryonic developmental competence in Arabidopsis
Histone H3.3 deposition in seed is essential for the post-embryonic developmental competence in Arabidopsis
Producing vigorous seeds that can germinate and establish seedlings is vital for plant propagation. Here, the authors report the critical function of histone variant H3.3 in establishing chromatin accessibility and seed vigour in Arabidopsis.
- Ting Zhao
- , Jingyun Lu
- & Danhua Jiang
-
Article
| Open Access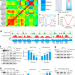 SESAME-catalyzed H3T11 phosphorylation inhibits Dot1-catalyzed H3K79me3 to regulate autophagy and telomere silencing
SESAME-catalyzed H3T11 phosphorylation inhibits Dot1-catalyzed H3K79me3 to regulate autophagy and telomere silencing
How H3T11 phosphorylation exerts biological functions remains poorly understood. Here, authors show that H3pT11 directly inhibits Dot1-catalyzed H3K79 tri-methylation (H3K79me3) and uncover how this histone crosstalk regulates autophagy and telomere silencing.
- Fei He
- , Qi Yu
- & Shanshan Li
-
Article
| Open AccessFood abundance in men before puberty predicts a range of cancers in grandsons
Nutritional conditions experienced early in life may influence the disease risk of future children and grandchildren. Here the authors report that food abundance among boys before puberty associates with the relative risk of a range of cancers in grandsons, but not in granddaughters.
- Denny Vågerö
- , Agneta Cederström
- & Gerard J. van den Berg
-
Article
| Open Access N6-methyladenosine RNA modification regulates photosynthesis during photodamage in plants
N6-methyladenosine RNA modification regulates photosynthesis during photodamage in plants
Efficient photoprotection is vital for the maintenance of photosynthetic efficiency during photodamage. Here, the authors reveal that m6A writer VIRILIZER acts as a molecular switch of photoprotection by post-transcriptional regulation in plants.
- Man Zhang
- , Yunping Zeng
- & Hong-Lei Jin
-
Article
| Open Access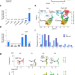 PCGF1-PRC1 links chromatin repression with DNA replication during hematopoietic cell lineage commitment
PCGF1-PRC1 links chromatin repression with DNA replication during hematopoietic cell lineage commitment
Here the authors show that a Polycomb Repressive Complex 1 (PRC1) containing PCGF1 prevents excessive loading of transcriptional activators and chromatin remodelers on nascent DNA, allowing proper deposition of nucleosomes immediately after the passage of the DNA replication fork to optimize downstream chromatin configurations in hematopoietic stem and progenitor cells (HSPCs).
- Junichiro Takano
- , Shinsuke Ito
- & Tomokatsu Ikawa
-
Article
| Open Access Chromatin accessibility dynamics dictate renal tubular epithelial cell response to injury
Chromatin accessibility dynamics dictate renal tubular epithelial cell response to injury
Renal tubular epithelial cells (TECs) can initiate an adaptive or maladaptive response after injuries of different severity. Here, the authors elucidate a chromatin-mediated mechanism underlying the responses of TECs to varying kidney injuries.
- Xinyi Cao
- , Jiuchen Wang
- & Lirong Zhang
-
Article
| Open Access DNA methylation-based classification of sinonasal tumors
DNA methylation-based classification of sinonasal tumors
Sinonasal tumour diagnosis can be complicated by the heterogeneity of disease and classification systems. Here, the authors use machine learning to classify sinonasal undifferentiated carcinomas into 4 molecular classe with differences in differentiation state and clinical outcome.
- Philipp Jurmeister
- , Stefanie Glöß
- & David Capper
-
Article
| Open Access Active DNA damage response signaling initiates and maintains meiotic sex chromosome inactivation
Active DNA damage response signaling initiates and maintains meiotic sex chromosome inactivation
Here the authors show that active DNA damage response signaling is required both to initiate and maintain meiotic sex chromosome inactivation via a dynamic and reversible process.
- Hironori Abe
- , Yu-Han Yeh
- & Satoshi H. Namekawa
-
Article
| Open Access ARID1A loss induces polymorphonuclear myeloid-derived suppressor cell chemotaxis and promotes prostate cancer progression
ARID1A loss induces polymorphonuclear myeloid-derived suppressor cell chemotaxis and promotes prostate cancer progression
The accumulation of myeloid derived suppressor cells (MDSC) has been associated with prostate cancer progression and castration resistance. Here the authors show that loss of ARID1A, a subunit of the SWI/SNF chromatin remodeling complex, results in polymorphonuclear-MDSC infiltration and cooperates with Pten loss to accelerate prostate tumorigenesis.
- Ni Li
- , Qiuli Liu
- & Jun Qin
-
Article
| Open Access Epigenetic regulation of white adipose tissue plasticity and energy metabolism by nucleosome binding HMGN proteins
Epigenetic regulation of white adipose tissue plasticity and energy metabolism by nucleosome binding HMGN proteins
White adipose tissue browning plays an important role in regulating whole-body energy homeostasis and metabolism. Here the authors show that the HMGN nucleosome binding proteins epigenetically promote white adipocyte differentiation and modulate the rate of white adipose tissue browning and energy metabolism in male mice.
- Ravikanth Nanduri
- , Takashi Furusawa
- & Michael Bustin
-
Article
| Open Access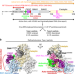 Structural basis for activation of DNMT1
Structural basis for activation of DNMT1
DNMT1 is an essential for maintaining genomic DNA methylation. Here, we report the cryo-EM structure of DNMT1 bound to ubiquitinated H3 and hemimethylated DNA, revealing structural insight into the activation mechanism of DNMT1.
- Amika Kikuchi
- , Hiroki Onoda
- & Kyohei Arita
-
Article
| Open Access PPARγ lipodystrophy mutants reveal intermolecular interactions required for enhancer activation
PPARγ lipodystrophy mutants reveal intermolecular interactions required for enhancer activation
Mutations in PPARγ lead to lipodystrophy, but the mechanisms by which the mutations affect the activity in chromatin is unknown. Here, Madsen, Broekema et al. showed that mutations affecting two intermolecular interactions compromise chromatin remodeling.
- Maria Stahl Madsen
- , Marjoleine F. Broekema
- & Eric Kalkhoven
-
Article
| Open Access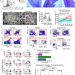 The 3D enhancer network of the developing T cell genome is shaped by SATB1
The 3D enhancer network of the developing T cell genome is shaped by SATB1
Here the authors analyze the 3D genome structure of murine thymocytes and show that SATB1, a predominantly T-cell specific protein, helps to establish a regulatory, finer-scale organizational layer built upon a pre-existing chromatin scaffold mediated by other architectural proteins, such as CTCF.
- Tomas Zelenka
- , Antonios Klonizakis
- & Charalampos Spilianakis
-
Article
| Open Access CK2-mediated phosphorylation of SUZ12 promotes PRC2 function by stabilizing enzyme active site
CK2-mediated phosphorylation of SUZ12 promotes PRC2 function by stabilizing enzyme active site
Here the authors identify SUZ12 as a cellular substate of casein kinase 2 (CK2), and show this phosphorylation changes the active site structure of Polycomb repressive complex 2 (PRC2) and promotes PRC2 function in cell identity maintenance during stem cell differentiation.
- Lihu Gong
- , Xiuli Liu
- & Xin Liu
-
Article
| Open Access Nicking mechanism underlying the DNA phosphorothioate-sensing antiphage defense by SspE
Nicking mechanism underlying the DNA phosphorothioate-sensing antiphage defense by SspE
SspABCD–SspE is a phosphorothioation-sensing bacterial defence system. Here, authors find that SspE exploits DNA phosphorothioation-binding preference to modulate its biochemical activities for the self/nonself discrimination and phage resistance.
- Haiyan Gao
- , Xinqi Gong
- & Lianrong Wang
-
Article
| Open Access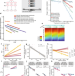 Cell division drives DNA methylation loss in late-replicating domains in primary human cells
Cell division drives DNA methylation loss in late-replicating domains in primary human cells
DNA methylation loss has been observed in aging tissues and cancers for decades. Researchers from Van Andel Institute have now provided experimental evidence that this process is directly driven by cell division.
- Jamie L. Endicott
- , Paula A. Nolte
- & Peter W. Laird
-
Article
| Open Access Parent-of-Origin inference for biobanks
Parent-of-Origin inference for biobanks
Studies on parent-of-origin effects have been limited in terms of sample size due to lack of parental genomes or known genealogies. Here, the authors develop a method to infer the parent-of-origin of an individual alleles in biobank-scale datasets, without requiring parental genomes or prior knowledge of genealogy, allowing discovery of parent-of-origin effects with an unprecedented sample size.
- Robin J. Hofmeister
- , Simone Rubinacci
- & Olivier Delaneau
-
Article
| Open Access Hi-TrAC reveals division of labor of transcription factors in organizing chromatin loops
Hi-TrAC reveals division of labor of transcription factors in organizing chromatin loops
It is currently not clear how architectural proteins orchestrate chromatin looping at different scales of genome organisation. Here the authors report Hi-TrAC as a proximity ligation-free method to profile genome-wide chromatin interactions at single nucleosome resolution among regulatory elements.
- Shuai Liu
- , Yaqiang Cao
- & Keji Zhao
-
Article
| Open Access Epigenetic Alterations of Repeated Relapses in Patient-matched Childhood Ependymomas
Epigenetic Alterations of Repeated Relapses in Patient-matched Childhood Ependymomas
While recurrence is frequent in ependymoma, the underlying molecular mechanisms remain to be explored. Here, the authors investigate epigenetic, genetic and tumorigenic changes in 30 patient-matched repeated relapses over 13 years and identify distinct patterns of DNA methylation.
- Sibo Zhao
- , Jia Li
- & Xiao-Nan Li
-
Article
| Open Access Defining cellular complexity in human autosomal dominant polycystic kidney disease by multimodal single cell analysis
Defining cellular complexity in human autosomal dominant polycystic kidney disease by multimodal single cell analysis
Autosomal dominant polycystic kidney disease (ADPKD) is a complicated disease that involves numerous cell types. Here the authors used a multiomics approach consisting of single nucleus transcriptomes and epigenomes to redefine cell states in ADPKD and to dissect the cellular interactions and molecular mechanisms of ADPKD.
- Yoshiharu Muto
- , Eryn E. Dixon
- & Benjamin D. Humphreys
-
Article
| Open Access Epigenetic control of chromosome-associated lncRNA genes essential for replication and stability
Epigenetic control of chromosome-associated lncRNA genes essential for replication and stability
Heskett et al. describe several members of a class of long non-coding RNAs, known as ASARs, which show distinct epigenetic regulation between subclonal lineages and are essential for normal DNA replication timing and stability of human autosomes.
- Michael B. Heskett
- , Athanasios E. Vouzas
- & Mathew J. Thayer
-
Article
| Open Access DNA methyltransferase 3A controls intestinal epithelial barrier function and regeneration in the colon
DNA methyltransferase 3A controls intestinal epithelial barrier function and regeneration in the colon
DNA methyltransferase 3 A (DNMT3A) is involved in DNA methylation, and genetic variants in the DNMT3 locus have been associated with inflammatory bowel disease. Here the authors report that DNMT3A controls intestinal epithelial barrier function and restoration of the gut barrier function after intestinal epithelial perturbation.
- Antonella Fazio
- , Dora Bordoni
- & Philip Rosenstiel
-
Article
| Open Access Acute deletion of TET enzymes results in aneuploidy in mouse embryonic stem cells through decreased expression of Khdc3
Acute deletion of TET enzymes results in aneuploidy in mouse embryonic stem cells through decreased expression of Khdc3
Inducible disruption of TET dioxygenases in mouse embryonic stem cells results in chromosome mis-segregation and aneuploidies, due to Khdc3 downregulation suggesting a role for TET enzymes and DNA methylation patterns in maintaining genome stability.
- Romain O. Georges
- , Hugo Sepulveda
- & Anjana Rao
-
Article
| Open Access DNA methylation at birth in monozygotic twins discordant for pediatric acute lymphoblastic leukemia
DNA methylation at birth in monozygotic twins discordant for pediatric acute lymphoblastic leukemia
The role of DNA methylation in predisposing to pediatric acute lymphoblastic leukemia remains unknown. Here, the authors utilize a discordant twin model to investigate how DNA methylation variation contributes to disease risk in genetically identical subjects.
- Eric M. Nickels
- , Shaobo Li
- & Joseph L. Wiemels
-
Article
| Open Access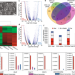 Allele-specific differential regulation of monoallelically expressed autosomal genes in the cardiac lineage
Allele-specific differential regulation of monoallelically expressed autosomal genes in the cardiac lineage
The authors use allele-specific single cell transcriptomic analysis to elucidate the establishment of monoallelic gene expression in the cardiac lineage. The findings emphasize the importance of allele-specific insight into gene regulation in development, homeostasis and disease.
- Gayan I. Balasooriya
- & David L. Spector
-
Article
| Open Access The immune factors driving DNA methylation variation in human blood
The immune factors driving DNA methylation variation in human blood
Many studies assess epigenetic marks in white blood cells, but it is unclear how much immune factors affect the epigenome. Here, the authors show that fine-scale blood cell composition and cytomegalovirus infection affect the DNA methylome of adults.
- Jacob Bergstedt
- , Sadoune Ait Kaci Azzou
- & Lluis Quintana-Murci
-
Article
| Open Access Vitamin C epigenetically controls osteogenesis and bone mineralization
Vitamin C epigenetically controls osteogenesis and bone mineralization
For decades vitamin C’s primary function in bone has been attributed to its involvement in collagen synthesis. Here, the authors uncover that vitamin C’s central role in bone is to globally orchestrate osteogenesis via epigenetic mechanisms.
- Roman Thaler
- , Farzaneh Khani
- & Andre J. van Wijnen
-
Article
| Open Access MYPT1-PP1β phosphatase negatively regulates both chromatin landscape and co-activator recruitment for beige adipogenesis
MYPT1-PP1β phosphatase negatively regulates both chromatin landscape and co-activator recruitment for beige adipogenesis
How β-AR signaling coordinates epigenetic and transcriptional pathways is unknown. Here the authors show that cold-induced β-AR signaling negatively regulates MYPT1-PP1β phosphatase activity to orchestrate both pathways for beige adipogenesis.
- Hiroki Takahashi
- , Ge Yang
- & Juro Sakai
-
Article
| Open Access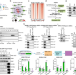 The transcriptional coactivator RUVBL2 regulates Pol II clustering with diverse transcription factors
The transcriptional coactivator RUVBL2 regulates Pol II clustering with diverse transcription factors
RNA polymerase II (Pol II) transcription factories play a central role in gene expression and 3D chromatin organization. Here, the authors demonstrate that RUVBL2 directly regulates Pol II clustering at active gene promoters.
- Hui Wang
- , Boyuan Li
- & Xiong Ji
-
Article
| Open Access Phosphorylation of Jhd2 by the Ras-cAMP-PKA(Tpk2) pathway regulates histone modifications and autophagy
Phosphorylation of Jhd2 by the Ras-cAMP-PKA(Tpk2) pathway regulates histone modifications and autophagy
How cells transduce nutrient availability to appropriate gene expression remains poorly understood. Here the authors show that the nutrient sensor, protein kinase A modulates histone modifications and gene transcription by phosphorylating histone demethylase.
- Qi Yu
- , Xuanyunjing Gong
- & Shanshan Li

 Arabidopsis TRB proteins function in H3K4me3 demethylation by recruiting JMJ14
Arabidopsis TRB proteins function in H3K4me3 demethylation by recruiting JMJ14