Featured
-
-
Article
| Open Access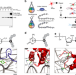 Analysis of protein-DNA interactions in chromatin by UV induced cross-linking and mass spectrometry
Analysis of protein-DNA interactions in chromatin by UV induced cross-linking and mass spectrometry
Cross-linking mass spectrometry (XLMS) allows mapping of protein-protein and protein-RNA interactions, but the analysis of protein-DNA complexes remains challenging. Here, the authors develop a UV light-based XLMS workflow to determine protein-DNA interfaces in reconstituted chromatin and isolated nuclei.
- Alexandra Stützer
- , Luisa M. Welp
- & Henning Urlaub
-
Article
| Open Access Cryo-EM structure of the deltaretroviral intasome in complex with the PP2A regulatory subunit B56γ
Cryo-EM structure of the deltaretroviral intasome in complex with the PP2A regulatory subunit B56γ
Human T-cell lymphotropic virus type 1 (HTLV-1) is a pathogenic deltaretrovirus. Here the authors resolve the cryo-EM structure of the deltaretroviral intasome in complex with the human host PP2A regulatory subunit.
- Michał S. Barski
- , Jordan J. Minnell
- & Goedele N. Maertens
-
Article
| Open Access Structural snapshots of human DNA polymerase μ engaged on a DNA double-strand break
Structural snapshots of human DNA polymerase μ engaged on a DNA double-strand break
Polymerase μ (Polμ) participates in the repair of DNA double-strand breaks (DSBs) via the nonhomologous end-joining (NHEJ) pathway. Here, the authors determine the crystal structure of a pre-catalytic ternary complex of human Polμ with a bound DSB substrate and they obtain further mechanistic insights by allowing the insertion reaction to proceed in crystallo, which enabled them to determine a Polμ structure with incomplete incorporation and the structure of the post-catalytic nicked state.
- Andrea M. Kaminski
- , John M. Pryor
- & Katarzyna Bebenek
-
Article
| Open Access Multiplex flow magnetic tweezers reveal rare enzymatic events with single molecule precision
Multiplex flow magnetic tweezers reveal rare enzymatic events with single molecule precision
Single molecule force measurements have shed light on dynamic biological events, but rare events escape notice owing to low throughput of the methods. Here, the authors combine an array of magnetic tweezers with lateral flow to increase throughput 100-fold, and detect rare DNA breaks induced by gyrase.
- Rohit Agarwal
- & Karl E. Duderstadt
-
Article
| Open Access Structural mechanism for replication origin binding and remodeling by a metazoan origin recognition complex and its co-loader Cdc6
Structural mechanism for replication origin binding and remodeling by a metazoan origin recognition complex and its co-loader Cdc6
The origin recognition complex (ORC) is essential for loading the Mcm2–7 replicative helicase onto DNA during DNA replication initiation. Here, the authors describe several cryo-electron microscopy structures of Drosophila ORC bound to DNA and its cofactor Cdc6 and also report an in vitro reconstitution system for Drosophila Mcm2–7 loading, revealing unexpected features of ORC’s DNA binding and remodeling mechanism during Mcm2–7 loading.
- Jan Marten Schmidt
- & Franziska Bleichert
-
Article
| Open Access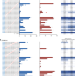 Genome-wide specificity of dCpf1 cytidine base editors
Genome-wide specificity of dCpf1 cytidine base editors
Cas12a-linked base editors can broaden the targeting scope of programmable cytidine deaminases. Here the authors assess their target specificity in an in vitro genome-wide assay.
- Daesik Kim
- , Kayeong Lim
- & Jin-Soo Kim
-
Article
| Open Access Zuo1 supports G4 structure formation and directs repair toward nucleotide excision repair
Zuo1 supports G4 structure formation and directs repair toward nucleotide excision repair
G4 structure-interacting proteins have been linked to DNA repair processes. Here the authors reveal that in yeast, Zuo1 associates with G4 structures and plays a role in repair pathway choice by promoting nucleotide excision repair and ultimately contributing to maintenance of genome stability.
- Alessio De Magis
- , Silvia Götz
- & Katrin Paeschke
-
Article
| Open Access Duplex DNA engagement and RPA oppositely regulate the DNA-unwinding rate of CMG helicase
Duplex DNA engagement and RPA oppositely regulate the DNA-unwinding rate of CMG helicase
During eukaryotic chromosome replication cells utilize ring-shaped CMG helicase that separates the two strands of the DNA double helix. Here the authors reveal that CMG helicase activity is inhibited by duplex DNA engagement at the fork, which is relieved by binding of RPA to the lagging-strand template.
- Hazal B. Kose
- , Sherry Xie
- & Hasan Yardimci
-
Article
| Open Access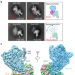 Structure of the polymerase ε holoenzyme and atomic model of the leading strand replisome
Structure of the polymerase ε holoenzyme and atomic model of the leading strand replisome
DNA polymerase epsilon (Pol ε) is responsible for leading strand synthesis during DNA replication. Here the authors use Cryo-EM to describe the architecture of the Pol ε holoenzyme and to provide an atomic model for the leading strand replisome.
- Zuanning Yuan
- , Roxana Georgescu
- & Huilin Li
-
Article
| Open Access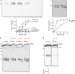 Specificity of end resection pathways for double-strand break regions containing ribonucleotides and base lesions
Specificity of end resection pathways for double-strand break regions containing ribonucleotides and base lesions
DNA double-strand break repair by homologous recombination initiates with nucleolytic resection of the 5’ DNA strand at the break ends. Here, the authors reveal that the lesion context influences the action and efficiency of the long range resection factors EXO1 and BLM-DNA2.
- James M. Daley
- , Nozomi Tomimatsu
- & Patrick Sung
-
Article
| Open Access Modular self-assembly of gamma-modified peptide nucleic acids in organic solvent mixtures
Modular self-assembly of gamma-modified peptide nucleic acids in organic solvent mixtures
Nucleic acid-based materials are usually only compatible with aqueous solutions. Here, the authors show a structural nucleic acid nanotechnology using gamma-modified peptide nucleic acids that enables formation of solvent-compatible, self-assembling nanostructures defined by Watson-Crick base pairing.
- Sriram Kumar
- , Alexander Pearse
- & Rebecca E. Taylor
-
Article
| Open Access Nucleoid remodeling during environmental adaptation is regulated by HU-dependent DNA bundling
Nucleoid remodeling during environmental adaptation is regulated by HU-dependent DNA bundling
HU is among the most conserved and abundant nucleoid-associated proteins in eubacteria. Here the authors investigate the role of histone-like proteins (HU) in the 3D organization of the bacteria DNA and show via soft X-ray tomography the process of nucleoid remodeling.
- Soumya G. Remesh
- , Subhash C. Verma
- & Michal Hammel
-
Article
| Open Access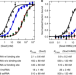 The Sox2 transcription factor binds RNA
The Sox2 transcription factor binds RNA
Some transcription factors have been proposed to functionally interact with RNA to facilitate proper regulation of gene expression. Here the authors demonstrate that human Sox2 interact directly and with high affinity to RNAs through its HMG DNA-binding domain.
- Zachariah E. Holmes
- , Desmond J. Hamilton
- & Robert T. Batey
-
Article
| Open Access In TFIIH the Arch domain of XPD is mechanistically essential for transcription and DNA repair
In TFIIH the Arch domain of XPD is mechanistically essential for transcription and DNA repair
XPD is part of the TFIIH complex which plays major roles in transcription initiation and nucleotide excision repair (NER). Here the authors present a high-resolution crystal structure of the XPD-MAT1 interface and dissect the role of this interface in transcription and NER.
- Stefan Peissert
- , Florian Sauer
- & Caroline Kisker
-
Article
| Open Access Structural insights into sequence-dependent Holliday junction resolution by the chloroplast resolvase MOC1
Structural insights into sequence-dependent Holliday junction resolution by the chloroplast resolvase MOC1
Holliday junctions (HJs) are DNA intermediates in genetic recombination that are resolved by nuclease, termed resolvase, to ensure genome stability. Here, the authors provide insights into the resolution process by resolving the crystal structure of the chloroplast resolvase MOC1 with HJs substrates.
- Junjie Yan
- , Sixing Hong
- & Ping Yin
-
Article
| Open Access FAM111A protects replication forks from protein obstacles via its trypsin-like domain
FAM111A protects replication forks from protein obstacles via its trypsin-like domain
DNA-protein crosslinks represent obstacles on genomic DNA that can hamper progression of replication forks. Here, the authors reveal that FAM111A, a PCNA-interacting protein, plays part in mitigating the effect of protein obstacles on replication forks.
- Yusuke Kojima
- , Yuka Machida
- & Yuichi J. Machida
-
Article
| Open Access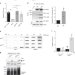 TEX264 coordinates p97- and SPRTN-mediated resolution of topoisomerase 1-DNA adducts
TEX264 coordinates p97- and SPRTN-mediated resolution of topoisomerase 1-DNA adducts
Eukaryotic topoisomerase 1 (TOP1) regulates DNA topology to ensure efficient DNA replication and transcription. Here, the authors reveal insights into the molecular resolution of topoisomerase 1-DNA adducts by TEX264, p97 and SPRTN.
- John Fielden
- , Katherine Wiseman
- & Kristijan Ramadan
-
Article
| Open Access Structure of the processive human Pol δ holoenzyme
Structure of the processive human Pol δ holoenzyme
Pol δ bound to the proliferating cell nuclear antigen (PCNA) replicates the lagging strand in eukaryotes and cooperates with flap endonuclease 1 (FEN1) to process the Okazaki fragments for their ligation. Here, the authors present a Cryo-EM structure of the human 4-subunit Pol δ bound to DNA and PCNA in a replicating state with an incoming nucleotide in the active site.
- Claudia Lancey
- , Muhammad Tehseen
- & Alfredo De Biasio
-
Article
| Open Access In vitro self-replication and multicistronic expression of large synthetic genomes
In vitro self-replication and multicistronic expression of large synthetic genomes
A main objective of synthetic biology is the creation of chemical systems capable of replication and evolution. Here, the authors demonstrate combined self-replication and expression of multipartite genomes in vitro.
- K. Libicher
- , R. Hornberger
- & H. Mutschler
-
Article
| Open Access DNA unwinding mechanism of a eukaryotic replicative CMG helicase
DNA unwinding mechanism of a eukaryotic replicative CMG helicase
The DNA duplex is known to be split apart in a steric exclusion manner during replication, but the specific mechanism has remained unclear. Here the authors present a cryo-EM structure of a eukaryotic replicative CMG helicase on forked DNA, revealing the mechanism of DNA unwinding.
- Zuanning Yuan
- , Roxana Georgescu
- & Michael E. O’Donnell
-
Article
| Open Access Eukaryotic transcription factors can track and control their target genes using DNA antennas
Eukaryotic transcription factors can track and control their target genes using DNA antennas
To carry out their function, transcription factors must efficiently recognize specific DNA sequence targets, a complex problem in the context of eukaryotic chromatin. Here the authors use single-molecule biophysical experiments, statistical mechanical theory and bioinformatics analyses to conclude that interactions with non-target sequences near promoters serve to increase overall affinity and targeting efficiency.
- Milagros Castellanos
- , Nivin Mothi
- & Victor Muñoz
-
Article
| Open Access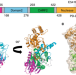 Structure and mechanism of a Type III CRISPR defence DNA nuclease activated by cyclic oligoadenylate
Structure and mechanism of a Type III CRISPR defence DNA nuclease activated by cyclic oligoadenylate
Antiviral defence type III CRISPR systems produce cyclic oligoadenylates (cOA) as second messengers that activate downstream effectors. Here the authors present the crystal structure of the type III CRISPR defence DNA nuclease Can1 in complex with cyclic tetra-adenylate (cA4) and show that Can1 nicks supercoiled DNA.
- Stephen A. McMahon
- , Wenlong Zhu
- & Tracey M. Gloster
-
Article
| Open Access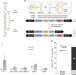 Distinct DNA repair pathways cause genomic instability at alternative DNA structures
Distinct DNA repair pathways cause genomic instability at alternative DNA structures
Z-DNA-forming CG repeats are mutagenic in mammalian cells but the mechanism has remained unknown so far. Here, the authors show that the nucleotide excision repair complex Rad10-Rad1 (ERCC1-XPF) and the mismatch repair complex Msh2-Msh3 (MSH2-MSH3) are required for Z-DNA-induced genetic instability in yeast and human cells.
- Jennifer A. McKinney
- , Guliang Wang
- & Karen M. Vasquez
-
Article
| Open Access Chromatin fibers stabilize nucleosomes under torsional stress
Chromatin fibers stabilize nucleosomes under torsional stress
Torsional stress is generated during DNA replication and transcription, however, the propagation of twist in condensed chromatin is poorly understood. Here the authors measure how force and torque impact chromatin fibers and find that the fibers fold into a left-handed superhelix that can be stabilized by positive torsion, suggesting that chromatin fibers stabilize nucleosomes under torsional stress.
- Artur Kaczmarczyk
- , He Meng
- & Nynke H. Dekker
-
Article
| Open Access Carbon-nanotube reinforcement of DNA-silica nanocomposites yields programmable and cell-instructive biocoatings
Carbon-nanotube reinforcement of DNA-silica nanocomposites yields programmable and cell-instructive biocoatings
DNA composite materials have potential for biomedical sciences; however, control over the materials can be an issue. Here, the authors report on a carbon-nanotube reinforced DNA-silica gel with controllable mechanical properties to steer the attachment, proliferation, migration and release of cells.
- Yong Hu
- , Carmen M. Domínguez
- & Christof M. Niemeyer
-
Article
| Open Access Alkyladenine DNA glycosylase associates with transcription elongation to coordinate DNA repair with gene expression
Alkyladenine DNA glycosylase associates with transcription elongation to coordinate DNA repair with gene expression
How genome stability is maintained at regions of active transcription is currently not entirely clear. Here, the authors reveal an association between base excision repair factors and transcription elongation to modulate DNA repair.
- Nicola P. Montaldo
- , Diana L. Bordin
- & Barbara van Loon
-
Article
| Open Access MutL sliding clamps coordinate exonuclease-independent Escherichia coli mismatch repair
MutL sliding clamps coordinate exonuclease-independent Escherichia coli mismatch repair
The mechanics of MMR strand specific excision that begins at a distant ssDNA break are not yet clear. Here the authors have used multiple single molecule imaging techniques to visualize the behavior of MMR components on mismatched DNA substrates and reveal an exonuclease-independent mechanism for E.coli MMR.
- Jiaquan Liu
- , Ryanggeun Lee
- & Richard Fishel
-
Article
| Open Access RuvC uses dynamic probing of the Holliday junction to achieve sequence specificity and efficient resolution
RuvC uses dynamic probing of the Holliday junction to achieve sequence specificity and efficient resolution
Holliday junctions (HJs) are four-way DNA structures that occur in DNA repair by homologous recombination which are removed by nucleases such as RuvC. Here the authors provide structural and biochemical details on the steps required for the completion of the process.
- Karolina Maria Górecka
- , Miroslav Krepl
- & Marcin Nowotny
-
Article
| Open Access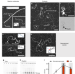 In vitro role of Rad54 in Rad51-ssDNA filament-dependent homology search and synaptic complexes formation
In vitro role of Rad54 in Rad51-ssDNA filament-dependent homology search and synaptic complexes formation
Homologous recombination uses a template to accurately repair DNA double-strand breaks and stalled replication forks to maintain genome stability. Here authors use electron microscopy to investigate the role of Rad54 in homology search and synaptic complex formation.
- Eliana Moreira Tavares
- , William Douglass Wright
- & Pauline Dupaigne
-
Article
| Open Access Satb1 integrates DNA binding site geometry and torsional stress to differentially target nucleosome-dense regions
Satb1 integrates DNA binding site geometry and torsional stress to differentially target nucleosome-dense regions
Satb1 is a master regulator of multiple cellular processes. Here the authors find that Satb1 preferentially targets nucleosome dense regions and combinatorially uses multiple selection criteria including DNA torsion, flanking DNA shape, motif density and periodicity to streamline binding choices.
- Rajarshi P. Ghosh
- , Quanming Shi
- & Jan T. Liphardt
-
Article
| Open Access Poly(ADP-ribose) polymerase-1 antagonizes DNA resection at double-strand breaks
Poly(ADP-ribose) polymerase-1 antagonizes DNA resection at double-strand breaks
Poly(ADP-ribose) polymerase-1 (PARP-1) facilitates local chromatin relaxation and the recruitment of DNA repair factors at double strand breaks site (DSBs). Here the authors reveal that PARP-1 acts as a critical regulator of DNA end resection of DSBs.
- Marie-Christine Caron
- , Ajit K. Sharma
- & Jean-Yves Masson
-
Article
| Open Access Design and fabrication of flexible DNA polymer cocoons to encapsulate live cells
Design and fabrication of flexible DNA polymer cocoons to encapsulate live cells
The ability to encapsulate living cells could lead to many applications. Here, the authors present a flexible method to graft DNA polymers onto bacteria, yeast and mammalian cells, polymerize them into DNA cocoons and use these to manipulate and select cells based on the encoded polymer sequences on DNA cocoons.
- Tao Gao
- , Tianshu Chen
- & Genxi Li
-
Article
| Open Access Comprehensive study of nuclear receptor DNA binding provides a revised framework for understanding receptor specificity
Comprehensive study of nuclear receptor DNA binding provides a revised framework for understanding receptor specificity
The type II nuclear receptors (NRs) and the retinoid X receptor (RXR) form heterodimeric transcription factors to regulate development, metabolism, and inflammation. Here the authors employ protein-binding microarrays to comprehensively analyze the DNA binding of 12 NR:RXRα heterodimers, and report promiscuous NR-DNA binding.
- Ashley Penvose
- , Jessica L. Keenan
- & Trevor Siggers
-
Article
| Open Access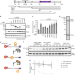 Regulation of PCNA cycling on replicating DNA by RFC and RFC-like complexes
Regulation of PCNA cycling on replicating DNA by RFC and RFC-like complexes
Replication-Factor-C (RFC) and RFC-like complexes (RLCs) mediate chromatin engagement of the proliferating cell nuclear antigen (PCNA). Here authors use biochemical and single molecule measurements to show that ATAD5-RLC has the most potent PCNA unloading activity and forms structurally distinct intermediates compared to RFC-PCNA.
- Mi-Sun Kang
- , Eunjin Ryu
- & Kyungjae Myung
-
Article
| Open Access Atomic resolution cryo-EM structure of a native-like CENP-A nucleosome aided by an antibody fragment
Atomic resolution cryo-EM structure of a native-like CENP-A nucleosome aided by an antibody fragment
CENP-A histone variants replace histones H3 at centromeres. Here the authors use a single-chain antibody fragment (scFv) to stabilize human CENP-A nucleosome containing a native α-satellite DNA and solved its structure by cryo-EM to 2.6 Å resolution, providing insight into the structure and function of the CENP-A nucleosome.
- Bing-Rui Zhou
- , K. N. Sathish Yadav
- & Ping Zhang
-
Article
| Open Access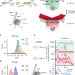 Direct observation of coordinated DNA movements on the nucleosome during chromatin remodelling
Direct observation of coordinated DNA movements on the nucleosome during chromatin remodelling
Chromatin remodelling enzymes (remodellers) regulate DNA accessibility of eukaryotic genomes, which rely in large part on an ability to reposition nucleosomes. Here the authors use three-colour single-molecule FRET to simultaneously monitor remodeller-induced DNA movements on both sides of the nucleosome in real-time.
- Anton Sabantsev
- , Robert F. Levendosky
- & Sebastian Deindl
-
Article
| Open Access Identification of human D lactate dehydrogenase deficiency
Identification of human D lactate dehydrogenase deficiency
D-lactic acidosis typically occurs in the context of short bowel syndrome; excess D-lactate is produced by intestinal bacteria. Here, the authors identify two point mutations in the human lactate dehydrogenase D (LDHD) gene that cause enzymatic loss of function and are associated with elevated plasma D-lactate.
- Glen R. Monroe
- , Albertien M. van Eerde
- & Judith J. Jans
-
Article
| Open Access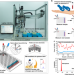 Measuring luteinising hormone pulsatility with a robotic aptamer-enabled electrochemical reader
Measuring luteinising hormone pulsatility with a robotic aptamer-enabled electrochemical reader
Assessment of luteinising hormone pulsatility is important in the diagnosis of reproductive disorders. Here the authors develop a DNA aptamer-based electrochemical analysis integrated into a robotic platform for high-throughput and sensitive analysis.
- Shaolin Liang
- , Andrew B. Kinghorn
- & Julian A. Tanner
-
Article
| Open Access Copy-choice recombination during mitochondrial L-strand synthesis causes DNA deletions
Copy-choice recombination during mitochondrial L-strand synthesis causes DNA deletions
Large-scale deletions of mitochondrial DNA (mtDNA) are associated with different human mitochondrial diseases and normal human ageing. Here the authors present a model for mtDNA formation based on generation sequencing analysis of patients samples and in vitro reconstituted mtDNA deletion using purified proteins.
- Örjan Persson
- , Yazh Muthukumar
- & Maria Falkenberg
-
Article
| Open Access Conformations and cryo-force spectroscopy of spray-deposited single-strand DNA on gold
Conformations and cryo-force spectroscopy of spray-deposited single-strand DNA on gold
Cryo-electron microscopy can determine the structure but not the nanomechanics of biological matter. Here the authors combine force spectroscopy in cryogenic conditions with computer simulations to characterize the properties of DNA simultaneously down to the sub-nm level.
- Rémy Pawlak
- , J. G. Vilhena
- & Ernst Meyer
-
Article
| Open Access A recurrent cancer-associated substitution in DNA polymerase ε produces a hyperactive enzyme
A recurrent cancer-associated substitution in DNA polymerase ε produces a hyperactive enzyme
Somatic alterations in the exonuclease domain of DNA polymerase ɛ have been linked to the development of highly mutated cancers. Here, the authors report that a major consequence of the most common cancer-associated Polɛ variant is a dramatically increased DNA polymerase activity.
- Xuanxuan Xing
- , Daniel P. Kane
- & Polina V. Shcherbakova
-
Article
| Open Access Structural consequence of the most frequently recurring cancer-associated substitution in DNA polymerase ε
Structural consequence of the most frequently recurring cancer-associated substitution in DNA polymerase ε
Mutations in the human POLE gene are associated with tumours with high mutational loads. Here the authors provide a structural rationale for the mutagenic activity of the cancer-associated DNA polymerase ε P286R variant.
- Vimal Parkash
- , Yashraj Kulkarni
- & Erik Johansson
-
Article
| Open Access Promotion of homology-directed DNA repair by polyamines
Promotion of homology-directed DNA repair by polyamines
The maintenance polyamines homeostasis is important for cell growth, and several cancers harbor elevated levels of polyamines that may contribute to sustained proliferative potential. Here the authors demonstrate that polyamines participate in DNA double-strand break repair through the stimulation of RAD51-mediated homologous DNA pairing and strand exchange.
- Chih-Ying Lee
- , Guan-Chin Su
- & Peter Chi
-
Article
| Open Access Experimental evidence of symmetry breaking of transition-path times
Experimental evidence of symmetry breaking of transition-path times
Microscopic transition mechanisms impact many biophysical systems. In this work, the authors explore transition path times between thermodynamic states experimentally, and show symmetry breaking in the transition times under an external force that drives the system out of equilibrium.
- J. Gladrow
- , M. Ribezzi-Crivellari
- & U. F. Keyser
-
Article
| Open Access Nucleotide-dependent DNA gripping and an end-clamp mechanism regulate the bacteriophage T4 viral packaging motor
Nucleotide-dependent DNA gripping and an end-clamp mechanism regulate the bacteriophage T4 viral packaging motor
Packaging of viral DNA depends on strong molecular motors that are powered by ATP hydrolysis. Here, the authors develop a single-molecule assay to monitor how nucleotide binding regulates motor-DNA interactions and reveal a generic mechanism that prevents exit of the whole DNA from the viral capsid during packaging.
- Mariam Ordyan
- , Istiaq Alam
- & Douglas E. Smith
-
Article
| Open Access Structural basis of meiotic telomere attachment to the nuclear envelope by MAJIN-TERB2-TERB1
Structural basis of meiotic telomere attachment to the nuclear envelope by MAJIN-TERB2-TERB1
The meiotic telomere complex (MAJIN, TERB1, TERB2) tethers telomere ends to the nuclear envelope. Here the authors present the crystal structure of human MAJIN-TERB2 and combine biophysical approaches and structured illumination microscopy analysis of mouse meiotic chromosomes to characterize the molecular architecture of the wider MAJIN-TERB2-TERB1 complex and its interactions with TRF1.
- James M. Dunce
- , Amy E. Milburn
- & Owen R. Davies
-
Article
| Open Access Structural basis for the recognition of sulfur in phosphorothioated DNA
Structural basis for the recognition of sulfur in phosphorothioated DNA
DNA phosphorothioation (PT-DNA) is a DNA backbone sulfur modification that is recognized by the type-IV restriction endonuclease ScoMcrA. Here the authors provide insights into sulfur recognition by solving the crystal structure of the PT-DNA bound sulfur-binding domain (SBD) from ScoMcrA and they further show that SBD homologs are widely spread among prokaryotes.
- Guang Liu
- , Wencheng Fu
- & Xinyi He
-
Article
| Open Access PWWP2A binds distinct chromatin moieties and interacts with an MTA1-specific core NuRD complex
PWWP2A binds distinct chromatin moieties and interacts with an MTA1-specific core NuRD complex
PWWP2A is a chromatin-binding transcriptional regulator that mediates mitosis-progression. Here, the authors provide evidence that PWWP2A directly interacts with H2A.Z nucleosomes, DNA and H3K36me3, binds to an MTA1-specific subcomplex of the NuRD complex (M1HR) and promotes changes to histone acetylation.
- Stephanie Link
- , Ramona M. M. Spitzer
- & Sandra B. Hake
-
Article
| Open Access Pol μ dGTP mismatch insertion opposite T coupled with ligation reveals promutagenic DNA repair intermediate
Pol μ dGTP mismatch insertion opposite T coupled with ligation reveals promutagenic DNA repair intermediate
Incorporation of mismatched nucleotides during DNA replication or repair can lead to mutagenesis. Here the authors reveal that DNA ligase can ligate NHEJ intermediates following incorporation of 8-oxodGTP or dGTP opposite T by DNA Polymerase mu (Pol mu) in vitro, which suggests that Pol mu could cause promutagenic mismatches during DSB repair.
- Melike Çağlayan
- & Samuel H. Wilson

 The interaction of DNA repair factors ASCC2 and ASCC3 is affected by somatic cancer mutations
The interaction of DNA repair factors ASCC2 and ASCC3 is affected by somatic cancer mutations