Featured
-
-
Article
| Open Access Neonatal thyroxine activation modifies epigenetic programming of the liver
Neonatal thyroxine activation modifies epigenetic programming of the liver
Whether thyroid hormones affect gene expression via DNA methylation is not well known. Here the authors show that type 2 deiodinase (D2) converts T4 to produce T3, which prevents DNA methylation of discrete areas in the neonatal liver. In the absence of D2, DNA methylation occurs and is associated with reduced chromatin accessibility in promoters and enhancers and affects gene expression.
- Tatiana L. Fonseca
- , Tzintzuni Garcia
- & Antonio C. Bianco
-
Article
| Open Access Early-life social experience affects offspring DNA methylation and later life stress phenotype
Early-life social experience affects offspring DNA methylation and later life stress phenotype
Early social experience can alter epigenetic patterns and stress responses later in life. A study on wild spotted hyenas finds that maternal care and social connections after leaving the den influence DNA methylation and contribute to a developmentally plastic stress response.
- Zachary M. Laubach
- , Julia R. Greenberg
- & Wei Perng
-
Article
| Open Access Arabidopsis MORC proteins function in the efficient establishment of RNA directed DNA methylation
Arabidopsis MORC proteins function in the efficient establishment of RNA directed DNA methylation
MORC ATPases are required for transposable element silencing and heterochromatin condensation in plants and animals. Here the authors show that Arabidopsis MORCs colocalize with sites of RNA-directed DNA methylation and provide evidence that they act as molecular tethers to efficiently establish DNA methylation.
- Yan Xue
- , Zhenhui Zhong
- & Steven E. Jacobsen
-
Article
| Open Access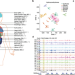 Tissue-specific 5-hydroxymethylcytosine landscape of the human genome
Tissue-specific 5-hydroxymethylcytosine landscape of the human genome
Charting the landscape of 5hmC in human tissues is fundamental to understanding its regulatory functions. Here, we systematically profiled the whole-genome 5hmC landscape at single-base resolution for 19 types of human tissues and found 5hmC shows tissue-specific patterns.
- Bo He
- , Chao Zhang
- & Chengqi Yi
-
Article
| Open Access Redirected nuclear glutamate dehydrogenase supplies Tet3 with α-ketoglutarate in neurons
Redirected nuclear glutamate dehydrogenase supplies Tet3 with α-ketoglutarate in neurons
α-ketoglutarate (αKG) is an intermediate in the tricarboxylic acid cycle that is required in the nucleus for genomic DNA demethylation by Tet3. Here, the authors show that the enzyme glutamate dehydrogenase, which converts glutamate to αKG, is redirected from the mitochondria to the nucleus.
- Franziska R. Traube
- , Dilara Özdemir
- & Thomas Carell
-
Article
| Open Access Environmental enrichment preserves a young DNA methylation landscape in the aged mouse hippocampus
Environmental enrichment preserves a young DNA methylation landscape in the aged mouse hippocampus
Decline of brain function during aging is associated with epigenetic changes, including DNA methylation. Here the authors provide evidence that environmental enrichment delays age-related DNA methylation alterations in the mouse hippocampus.
- Sara Zocher
- , Rupert W. Overall
- & Gerd Kempermann
-
Article
| Open Access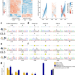 Genomic imprinting in mouse blastocysts is predominantly associated with H3K27me3
Genomic imprinting in mouse blastocysts is predominantly associated with H3K27me3
In most mammals, imprinted genes contain epigenetic marks that differ in each parental genome and control their parent-of-origin-specific expression. Here, the authors map imprinted genes in mouse preimplantation embryos and find that imprinted gene expression in blastocysts is mainly dependent on Polycomb-mediated H3K27me3-associated gene silencing.
- Laura Santini
- , Florian Halbritter
- & Martin Leeb
-
Article
| Open Access E2F6 initiates stable epigenetic silencing of germline genes during embryonic development
E2F6 initiates stable epigenetic silencing of germline genes during embryonic development
DNA methylation targets CpG island promoters of germline genes to repress their expression in mouse somatic cells. Here the authors show that a transcription factor E2F6 is required to target CpG island DNA methylation and epigenetic silencing to germline genes during early mouse development.
- Thomas Dahlet
- , Matthias Truss
- & Michael Weber
-
Article
| Open Access High throughput screening identifies SOX2 as a super pioneer factor that inhibits DNA methylation maintenance at its binding sites
High throughput screening identifies SOX2 as a super pioneer factor that inhibits DNA methylation maintenance at its binding sites
The functional relevance of epigenetic modifications on transcription regulation has been an important question since their discovery. Here, the authors investigate the effect of DNA methylation on Pioneer Transcription Factor (PF) binding and distinguish between PFs that protect their binding sites from methylation and those that bind to methylated DNA and induce DNA demethylation.
- Ludovica Vanzan
- , Hadrien Soldati
- & Rabih Murr
-
Article
| Open Access TET1-mediated DNA hydroxymethylation regulates adult remyelination in mice
TET1-mediated DNA hydroxymethylation regulates adult remyelination in mice
Myelin formation is regulated by epigenetic mechanisms and ensures proper neuronal function during development and after demyelination. Here, the authors show that TET1, a DNA hydroxymethylase, regulates myelin repair in adult mice, but is defective with aging.
- Sarah Moyon
- , Rebecca Frawley
- & Patrizia Casaccia
-
Article
| Open Access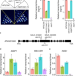 A histone H3K4me1-specific binding protein is required for siRNA accumulation and DNA methylation at a subset of loci targeted by RNA-directed DNA methylation
A histone H3K4me1-specific binding protein is required for siRNA accumulation and DNA methylation at a subset of loci targeted by RNA-directed DNA methylation
In plants, RNA-directed DNA methylation (RdDM) is a de novo DNA methylation pathway that is responsible for transcriptional silencing of repetitive elements. Here, the authors characterized a new RdDM factor, RDM15, and show that it is required for RdDM-dependent DNA methylation and siRNA accumulation at a subset of RdDM target loci.
- Qingfeng Niu
- , Zhe Song
- & Zhaobo Lang
-
Article
| Open Access Impact of DNA methylation on 3D genome structure
Impact of DNA methylation on 3D genome structure
Multi-layered epigenetic regulation in higher eukaryotes makes it challenging to disentangle the individual effects of modifications on chromatin structure and function. Here, the authors expressed mammalian DNA methyltransferases in yeast, which have no DNA methylation, to show that methylation has intrinsic effects on chromatin structure.
- Diana Buitrago
- , Mireia Labrador
- & Modesto Orozco
-
Article
| Open Access Ectopic targeting of CG DNA methylation in Arabidopsis with the bacterial SssI methyltransferase
Ectopic targeting of CG DNA methylation in Arabidopsis with the bacterial SssI methyltransferase
The ability to target DNA methylation to specific loci is important for both basic and applied research. Here, the authors fuse CG-specific methyltransferase SssI with an artificial zinc finger protein for DNA methylation targeting and show the chromatin features favorable for efficient gain of methylation.
- Wanlu Liu
- , Javier Gallego-Bartolomé
- & Steven E. Jacobsen
-
Article
| Open Access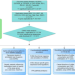 Epigenome-wide association meta-analysis of DNA methylation with coffee and tea consumption
Epigenome-wide association meta-analysis of DNA methylation with coffee and tea consumption
While coffee and tea consumption has been associated with risk of diseases, their mechanisms of action remain elusive. Here the authors present a large EWAS on coffee and tea consumption in cohorts of European and African-American ancestries, finding that coffee consumption is associated with differential DNA methylation levels at multiple CpGs.
- Irma Karabegović
- , Eliana Portilla-Fernandez
- & Mohsen Ghanbari
-
Article
| Open Access Contrasting epigenetic control of transgenes and endogenous genes promotes post-transcriptional transgene silencing in Arabidopsis
Contrasting epigenetic control of transgenes and endogenous genes promotes post-transcriptional transgene silencing in Arabidopsis
Accumulating evidences point to a discrepancy in the epigenetic behaviour of transgenes and endogenous genes. Here, via characterization of mutants impaired in histone demethylases JMJ14 and IBM1, the authors show that transgenes and endogenous genes are regulated by different epigenetic mechanisms in Arabidopsis.
- Nicolas Butel
- , Agnès Yu
- & Hervé Vaucheret
-
Article
| Open Access The genomic loci of specific human tRNA genes exhibit ageing-related DNA hypermethylation
The genomic loci of specific human tRNA genes exhibit ageing-related DNA hypermethylation
The epigenome has been shown to change with age, potentially impacting on ageing-related disease. Here the authors investigate the DNA methylation state of the genomic loci of human tRNA and observe enrichment for age-related DNA hypermethylation at tRNA loci.
- Richard J. Acton
- , Wei Yuan
- & Christopher G. Bell
-
Article
| Open Access The histone variant H2A.W and linker histone H1 co-regulate heterochromatin accessibility and DNA methylation
The histone variant H2A.W and linker histone H1 co-regulate heterochromatin accessibility and DNA methylation
T-DNA mutants have been widely used for Arabidopsis gene function characterization. Here, by characterizing a null mutant created by CRISPR, the authors show that previous reported function of H2A.W is confounded by a T-DNA insertion induced chromosomal rearrangement and reveal its role in regulating heterochromatin accessibility.
- Pierre Bourguet
- , Colette L. Picard
- & Olivier Mathieu
-
Article
| Open Access Comprehensive cell type decomposition of circulating cell-free DNA with CelFiE
Comprehensive cell type decomposition of circulating cell-free DNA with CelFiE
Tissue damage and turnover lead to the release of DNA in the blood and can be used to monitor changes in tissue state. Here, the authors developed a tool to accurately estimate the proportion of cell types contributing to cell-free DNA in the blood, with an application to pregnant women and ALS patients.
- Christa Caggiano
- , Barbara Celona
- & Noah Zaitlen
-
Article
| Open Access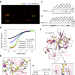 DNMT1 reads heterochromatic H4K20me3 to reinforce LINE-1 DNA methylation
DNMT1 reads heterochromatic H4K20me3 to reinforce LINE-1 DNA methylation
How histone modifications crosstalk with DNA methylation to regulate epigenomic patterning and genome stability in mammals remains elusive. Here, the authors show that DNA methyltransferase DNMT1 is a reader for histone H4K20 trimethylation via its BAH1 domain, which leads to optimal maintenance of DNA methylation at repetitive LINE-1 elements.
- Wendan Ren
- , Huitao Fan
- & Jikui Song
-
Article
| Open Access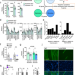 VPS39-deficiency observed in type 2 diabetes impairs muscle stem cell differentiation via altered autophagy and epigenetics
VPS39-deficiency observed in type 2 diabetes impairs muscle stem cell differentiation via altered autophagy and epigenetics
Insulin resistance and lower muscle strength in relation to mass are hallmarks of type 2 diabetes. Here, the authors report alterations in muscle stem cells from individuals with type 2 diabetes that may contribute to these phenotypes through VPS39 mediated effects on autophagy and epigenetics.
- Cajsa Davegårdh
- , Johanna Säll
- & Charlotte Ling
-
Article
| Open Access B1a and B2 cells are characterized by distinct CpG modification states at DNMT3A-maintained enhancers
B1a and B2 cells are characterized by distinct CpG modification states at DNMT3A-maintained enhancers
B cell progenitors differentiate into multiple subsets with distinct functions. Here the authors analyze the epigenetic landscapes of sorted B cell subsets using multiple platforms and show that the epigenetic regulator, DNMT3A, is essential for modulating the activity of enhancers critical for B1 and B2 lineage-determining genes.
- Vinay S. Mahajan
- , Hamid Mattoo
- & Shiv Pillai
-
Article
| Open Access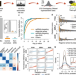 CRISPRi screens reveal a DNA methylation-mediated 3D genome dependent causal mechanism in prostate cancer
CRISPRi screens reveal a DNA methylation-mediated 3D genome dependent causal mechanism in prostate cancer
Prostate cancer risk-associated SNPs are enriched in noncoding CREs. Here the authors perform CRISPRi screens of CREs in prostate cancer cell lines to describe a causal mechanism synergistically driven by a risk SNP and DNA methylation-mediated 3D genome architecture.
- Musaddeque Ahmed
- , Fraser Soares
- & Housheng Hansen He
-
Article
| Open Access The epigenetic pioneer EGR2 initiates DNA demethylation in differentiating monocytes at both stable and transient binding sites
The epigenetic pioneer EGR2 initiates DNA demethylation in differentiating monocytes at both stable and transient binding sites
DNA methylation turnover is an essential epigenetic process during development. Here, the authors look at the changes in DNA methylation during the differentiation of post-mitotic human monocytes (MO), and find that EGR2 interacts with TET2 and is required for DNA demethylation at its binding sites; revealing EGR2 as an epigenetic pioneer factor in human MO.
- Karina Mendes
- , Sandra Schmidhofer
- & Michael Rehli
-
Article
| Open Access Strand-specific single-cell methylomics reveals distinct modes of DNA demethylation dynamics during early mammalian development
Strand-specific single-cell methylomics reveals distinct modes of DNA demethylation dynamics during early mammalian development
Erasure of DNA methylation from the parental genomes is critical to reset the methylome of differentiated gametes to pluripotent cells in the blastocyst. Here, the authors present a high-throughput single-cell method that enables strand-specific quantification of DNA methylation and identify distinct modes of DNA demethylation dynamics during early mammalian development.
- Maya Sen
- , Dylan Mooijman
- & Alexander van Oudenaarden
-
Article
| Open Access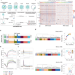 Single-cell multiomics sequencing reveals the functional regulatory landscape of early embryos
Single-cell multiomics sequencing reveals the functional regulatory landscape of early embryos
Extensive epigenetic reprogramming occurs during preimplantation embryo development. Here the authors develop a single cell multiomics sequencing technology that enables profiling of genome-wide chromatin accessibility, DNA methylation and RNA expression in the same individual cell and apply this method to study mouse preimplantation embryos.
- Yang Wang
- , Peng Yuan
- & Liying Yan
-
Article
| Open Access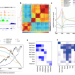 Non-CG methylation and multiple histone profiles associate child abuse with immune and small GTPase dysregulation
Non-CG methylation and multiple histone profiles associate child abuse with immune and small GTPase dysregulation
Early-life adversity is thought to increase the risk of psychopathology through epigenetic mechanisms. Here, the authors profile 6 histone marks, chromatin states and DNA methylation in the lateral amygdala in subjects with a history of early-life adversity.
- Pierre-Eric Lutz
- , Marc-Aurèle Chay
- & Gustavo Turecki
-
Article
| Open Access The genome-wide impact of trisomy 21 on DNA methylation and its implications for hematopoiesis
The genome-wide impact of trisomy 21 on DNA methylation and its implications for hematopoiesis
Down syndrome has a high co-morbidity with immune and hematopoietic disorders. Here, the authors perform an epigenome-wide association study in newborns with and without Down syndrome to find differential methylation across the genome, including in hematopoietic regulators RUNX1 and FLI1.
- Ivo S. Muskens
- , Shaobo Li
- & Adam J. de Smith
-
Article
| Open Access De novo DNA methyltransferase activity in colorectal cancer is directed towards H3K36me3 marked CpG islands
De novo DNA methyltransferase activity in colorectal cancer is directed towards H3K36me3 marked CpG islands
Aberrant gain of DNA methylation at CpG islands is frequently observed in colorectal tumours. Here the authors use ectopically integrated CpG islands in colorectal cancer cells and find that aberrantly methylated CpG islands are subject to low levels of de novo DNA methylation, and that instead de novo DNA methylation activity is targeted primarily to CpG islands marked by the histone modification H3K36me3.
- Roza H. Ali Masalmeh
- , Francesca Taglini
- & Duncan Sproul
-
Article
| Open Access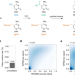 Subtraction-free and bisulfite-free specific sequencing of 5-methylcytosine and its oxidized derivatives at base resolution
Subtraction-free and bisulfite-free specific sequencing of 5-methylcytosine and its oxidized derivatives at base resolution
Specific and quantitative sequencing of cytosine modifications is challenging at base-resolution. Here the authors present TAPSβ and CAPS for subtraction-free whole genome sequencing of 5mC and 5hmC.
- Yibin Liu
- , Zhiyuan Hu
- & Chun-Xiao Song
-
Article
| Open Access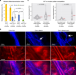 Male fertility in Arabidopsis requires active DNA demethylation of genes that control pollen tube function
Male fertility in Arabidopsis requires active DNA demethylation of genes that control pollen tube function
Active DNA demethylation is required for sexual reproduction in plants, but the underlying mechanisms are unknown. Here, the authors show that the DNA glycosylases DEMETER and REPRESSOR OF SILENCING 1 enable the DNA demethylation-dependent activation of genes involved in pollen tube progression.
- Souraya Khouider
- , Filipe Borges
- & Daniel Bouyer
-
Article
| Open Access Targeting aberrant DNA methylation in mesenchymal stromal cells as a treatment for myeloma bone disease
Targeting aberrant DNA methylation in mesenchymal stromal cells as a treatment for myeloma bone disease
Mesenchymal stromal cells (MSCs) have been shown to support multiple myeloma (MM) development. Here, MSCs isolated from the bone marrow of MM patients are shown to have altered DNA methylation patterns and a methyltransferase inhibitor reverts MM-associated bone loss and reduces tumour burden in MM murine models.
- Antonio Garcia-Gomez
- , Tianlu Li
- & Esteban Ballestar
-
Article
| Open Access Cellular Heterogeneity–Adjusted cLonal Methylation (CHALM) improves prediction of gene expression
Cellular Heterogeneity–Adjusted cLonal Methylation (CHALM) improves prediction of gene expression
Here, the authors introduce Cell Heterogeneity–Adjusted cLonal Methylation (CHALM) as a methylation quantification method that considers the heterogeneity of sequenced bulk cells. They apply CHALM to methylation datasets to detect differentially methylated genes that exhibit distinct biological functions supporting underlying mechanisms.
- Jianfeng Xu
- , Jiejun Shi
- & Wei Li
-
Article
| Open Access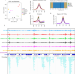 H2AK121ub in Arabidopsis associates with a less accessible chromatin state at transcriptional regulation hotspots
H2AK121ub in Arabidopsis associates with a less accessible chromatin state at transcriptional regulation hotspots
Polycomb Group complexes maintain gene repression through the incorporation of H2AK121ub and H3K27me3. Here, the authors show that H2AK121ub marks less accessible but transcriptionally permissive chromatin, while H3K27me3 enforces a repressed transcriptionally less-permissive state.
- Xiaochang Yin
- , Francisco J. Romero-Campero
- & Yue Zhou
-
Article
| Open Access The molecular basis for recognition of 5′-NNNCC-3′ PAM and its methylation state by Acidothermus cellulolyticus Cas9
The molecular basis for recognition of 5′-NNNCC-3′ PAM and its methylation state by Acidothermus cellulolyticus Cas9
Acidothermus cellulolyticus CRISPR-Cas9 (AceCas9) is a Type II-C enzyme that cleaves DNA in a Protospacer Adjacent Motif (PAM) methylation sensitive fashion. Biochemical analysis and crystal structures of AceCas9 in complex with sgRNA and DNA bearing the correct and incorrect PAM offer insight into the structural basis for the recognition of PAM and its methylation.
- Anuska Das
- , Travis H. Hand
- & Hong Li
-
Article
| Open Access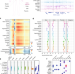 A human tissue map of 5-hydroxymethylcytosines exhibits tissue specificity through gene and enhancer modulation
A human tissue map of 5-hydroxymethylcytosines exhibits tissue specificity through gene and enhancer modulation
DNA 5-hydroxymethylcytosine (5hmC) modification is associated with gene transcription and used as a mark of mammalian development. Here the authors report a comprehensive 5hmC tissue map and analysis of 5hmC genomic distributions in 19 human tissues derived from 10 organ systems, thus providing insights into the role of 5hmC in tissue-specific development.
- Xiao-Long Cui
- , Ji Nie
- & Chuan He
-
Article
| Open Access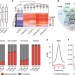 Recent evolution of a TET-controlled and DPPA3/STELLA-driven pathway of passive DNA demethylation in mammals
Recent evolution of a TET-controlled and DPPA3/STELLA-driven pathway of passive DNA demethylation in mammals
Active and passive demethylation pathways have been implicated in the genome-wide erasure of 5mC accompanying mammalian preimplantation development. Here the authors reveal a recently evolved, mammalian-specific pathway in which global hypomethylation is achieved by the coupling of active and passive demethylation.
- Christopher B. Mulholland
- , Atsuya Nishiyama
- & Heinrich Leonhardt
-
Article
| Open Access Maternal DNMT3A-dependent de novo methylation of the paternal genome inhibits gene expression in the early embryo
Maternal DNMT3A-dependent de novo methylation of the paternal genome inhibits gene expression in the early embryo
The paternal genome in mice undergoes widespread DNA methylation loss post-fertilization. Here, the authors apply allele-specific analysis of WGBS data to show that a number of genomic regions are simultaneously de novo methylated on the paternal genome dependent on maternal DNMT3A activity, which induces transcriptional silencing of this allele in the early embryo.
- Julien Richard Albert
- , Wan Kin Au Yeung
- & Matthew Lorincz
-
Article
| Open Access Globally altered epigenetic landscape and delayed osteogenic differentiation in H3.3-G34W-mutant giant cell tumor of bone
Globally altered epigenetic landscape and delayed osteogenic differentiation in H3.3-G34W-mutant giant cell tumor of bone
The histone variant mutation H3.3-G34W occurs in the majority of giant cell tumor of bone (GCTB). By profiling patient-derived GCTB tumor cells, the authors show that this mutation associates with epigenetic alterations in heterochromatic and bivalent regions that contribute to an impaired osteogenic differentiation and the osteolytic phenotype of GCTB.
- Pavlo Lutsik
- , Annika Baude
- & Christoph Plass
-
Article
| Open Access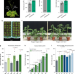 MSH1-induced heritable enhanced growth vigor through grafting is associated with the RdDM pathway in plants
MSH1-induced heritable enhanced growth vigor through grafting is associated with the RdDM pathway in plants
The meiotic transmissibility and progeny phenotypic influence of graft-mediated epigenetic changes remain unclear. Here, the authors use the msh1 mutant in the rootstock to trigger heritable enhanced growth vigor in Arabidopsis and tomato, and show it is associated with the RNA-directed DNA methylation pathway.
- Hardik Kundariya
- , Xiaodong Yang
- & Sally A. Mackenzie
-
Article
| Open Access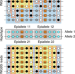 Detection of haplotype-dependent allele-specific DNA methylation in WGBS data
Detection of haplotype-dependent allele-specific DNA methylation in WGBS data
Allele-specific measurements can reveal differences in DNA methylation between homologous alleles associated with changes in genetic sequence. Here, the authors develop a method for detecting allele specific methylation events within haplotypes of linked SNPs, compare it with existing methods, and show it identifies haplotypes for which the genetic variant carries significant information about the methylation state of the allele of origin.
- J. Abante
- , Y. Fang
- & J. Goutsias
-
Article
| Open Access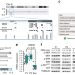 Haploinsufficiency of RREB1 causes a Noonan-like RASopathy via epigenetic reprogramming of RAS-MAPK pathway genes
Haploinsufficiency of RREB1 causes a Noonan-like RASopathy via epigenetic reprogramming of RAS-MAPK pathway genes
Mutations in RAS-MAPK pathway genes are implicated in Noonan-spectrum, yet up to 20% of cases have unknown cause. Here, the authors identify RREB1 underlying a 6p microdeletion RASopathy-like syndrome and show that RREB1, SIN3A and KDM1A form a transcriptional repressive complex to control methylation of MAPK pathway genes.
- Oliver A. Kent
- , Manipa Saha
- & Robert Rottapel
-
Article
| Open Access DNA methylation study of Huntington’s disease and motor progression in patients and in animal models
DNA methylation study of Huntington’s disease and motor progression in patients and in animal models
Although Huntington’s disease (HD) is a well-studied genetic disorder, less is known about the epigenetic changes underlying it. Here, the authors characterize DNA methylation levels in tissues from patients, a mouse huntingtin (Htt) gene knock-in model, and a transgenic HTT sheep model, and provide evidence that HD is accompanied by DNA methylation changes in these three species.
- Ake T. Lu
- , Pritika Narayan
- & Steve Horvath
-
Article
| Open Access Transgenerational inheritance of impaired larval T cell development in zebrafish
Transgenerational inheritance of impaired larval T cell development in zebrafish
Evidence for transgenerational inheritance of epigenetic information in vertebrates is scarce. Here the authors report that homozygous dnmt1 mutant zebrafish are essentially normal, with the exception of impaired lymphopoiesis, with impaired larval (but not adult) T cell development being transmitted to subsequent generations by genotypically wildtype fish.
- Norimasa Iwanami
- , Divine-Fondzenyuy Lawir
- & Thomas Boehm
-
Article
| Open Access Disease-associated KIF3A variants alter gene methylation and expression impacting skin barrier and atopic dermatitis risk
Disease-associated KIF3A variants alter gene methylation and expression impacting skin barrier and atopic dermatitis risk
Genetic variants in KIF3A are associated with atopic dermatitis (AD). Here, the authors identify two AD-risk alleles that show high methylation resulting in lower KIF3A expression. Mice with epidermis-specific loss of Kif3a show disrupted skin barrier homeostasis and increased AD susceptibility.
- Mariana L. Stevens
- , Zhonghua Zhang
- & Gurjit K. Khurana Hershey
-
Article
| Open Access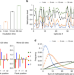 DNA sequence-dependent activity and base flipping mechanisms of DNMT1 regulate genome-wide DNA methylation
DNA sequence-dependent activity and base flipping mechanisms of DNMT1 regulate genome-wide DNA methylation
DNA methylation is one of the major epigenetic mechanisms that critically influence gene expression, genomic stability and cell differentiation. Here, the authors study DNMT1 in complex with DNA substrates and systematically analyze the mechanism and specificity of DNMT1.
- Sabrina Adam
- , Hiwot Anteneh
- & Albert Jeltsch
-
Article
| Open Access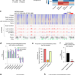 Transition to naïve human pluripotency mirrors pan-cancer DNA hypermethylation
Transition to naïve human pluripotency mirrors pan-cancer DNA hypermethylation
Epigenetic reprogramming is a hallmark of cancer. Here the authors find that resetting primed human embryonic stem cells to naïve state results in the acquisition of a DNA methylation landscape that mirrors the cancer DNA methylome and provides evidence that the transition to naïve pluripotency and oncogenic transformation share common epigenetic trajectories.
- Hemalvi Patani
- , Michael D. Rushton
- & Gabriella Ficz
-
Article
| Open Access Comprehensive structure-function characterization of DNMT3B and DNMT3A reveals distinctive de novo DNA methylation mechanisms
Comprehensive structure-function characterization of DNMT3B and DNMT3A reveals distinctive de novo DNA methylation mechanisms
In mammals, DNA methylation patterns are established by two de novo DNA methyltransferases, DNMT3A and DNMT3B. Here the authors report the crystal structures of DNMT3B in complex with both CpG and CpA DNA, providing insight into the substrate-recognition mechanism underpinning the divergent genomic methylation activities of DNMT3A and DNMT3B.
- Linfeng Gao
- , Max Emperle
- & Jikui Song
-
Article
| Open Access Epigenetic regulation of spurious transcription initiation in Arabidopsis
Epigenetic regulation of spurious transcription initiation in Arabidopsis
Epigenetic regulation can silence transposons and maintain gene expression. Here the authors survey Arabidopsis mutants defective in epigenetic regulation and show ectopic activation of thousands of cryptic TSSs and altered expression of nearby genes demonstrating the importance of suppressing spurious transcription.
- Ngoc Tu Le
- , Yoshiko Harukawa
- & Hidetoshi Saze
-
Article
| Open Access Identification of distinct loci for de novo DNA methylation by DNMT3A and DNMT3B during mammalian development
Identification of distinct loci for de novo DNA methylation by DNMT3A and DNMT3B during mammalian development
De novo DNA methylation is carried out by DNMT3A and DNMT3B, but the distinct functions of these two enzymes is poorly understood. Here the authors present a comprehensive, genome-wide identification of target sites for de novo DNA methylation by the DNMT3A and DNMT3B in mouse ES cells and embryos, identifying unique de novo DNA methylation target sites for both DNMT3 enzymes.
- Masaki Yagi
- , Mio Kabata
- & Yasuhiro Yamada

 A machine learning approach to brain epigenetic analysis reveals kinases associated with Alzheimer’s disease
A machine learning approach to brain epigenetic analysis reveals kinases associated with Alzheimer’s disease