Article
|
Open Access
Featured
-
-
Article
| Open Access The juxtamembrane linker of synaptotagmin 1 regulates Ca2+ binding via liquid-liquid phase separation
The juxtamembrane linker of synaptotagmin 1 regulates Ca2+ binding via liquid-liquid phase separation
Synaptotagmin (syt) 1 is a calcium sensor for neuronal exocytosis. Here, the authors show that the juxtamembrane linker of this integral membrane protein negatively regulates its calcium sensing activity by mediating self-association via liquid-liquid phase separation.
- Nikunj Mehta
- , Sayantan Mondal
- & Edwin R. Chapman
-
Article
| Open Access Tuning parameters for polygenic risk score methods using GWAS summary statistics from training data
Tuning parameters for polygenic risk score methods using GWAS summary statistics from training data
Some polygenic risk score (PRS) methods for predicting genetic risk for common diseases require an external individual-level dataset for parameter tuning, posing privacy-related concerns. Here, the authors present an empirical Bayes method that tunes PRS models using only summary statistics from the training data.
- Wei Jiang
- , Ling Chen
- & Hongyu Zhao
-
Article
| Open Access Revealing hidden patterns in deep neural network feature space continuum via manifold learning
Revealing hidden patterns in deep neural network feature space continuum via manifold learning
Existing feature visualisation methods are not well-suited for regression tasks. Here, authors introduce a method to learn the manifold topology related to deep neural network output and target labels and provide insightful visualisations of the high-dimensional features while preserving the local geometry.
- Md Tauhidul Islam
- , Zixia Zhou
- & Lei Xing
-
Article
| Open Access Pathway centric analysis for single-cell RNA-seq and spatial transcriptomics data with GSDensity
Pathway centric analysis for single-cell RNA-seq and spatial transcriptomics data with GSDensity
Clustering-based analysis has limited power in highly dynamic single-cell data, which is a common situation in tumour samples. Here, authors introduce GSDensity, enabling pathway-centric analysis for the direct integration of data with their domain knowledge.
- Qingnan Liang
- , Yuefan Huang
- & Ken Chen
-
Article
| Open Access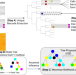 LinRace: cell division history reconstruction of single cells using paired lineage barcode and gene expression data
LinRace: cell division history reconstruction of single cells using paired lineage barcode and gene expression data
Inferring lineage trees while incorporating gene expressions and lineage barcodes is a challenging task. Here, authors present LinRace, which infers improved cell lineage trees and ancestral cell states using the proposed asymmetric division model.
- Xinhai Pan
- , Hechen Li
- & Xiuwei Zhang
-
Article
| Open Access A computational toolbox for the assembly yield of complex and heterogeneous structures
A computational toolbox for the assembly yield of complex and heterogeneous structures
Predicting the effective assembly of a set of proteins into a desired structure has traditionally been a challenging task. Here, authors demonstrate that advancements in automatic differentiation make it possible to address this problem using classical statistical mechanics.
- Agnese I. Curatolo
- , Ofer Kimchi
- & Michael P. Brenner
-
Article
| Open Access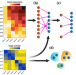 Accurate integration of single-cell DNA and RNA for analyzing intratumor heterogeneity using MaCroDNA
Accurate integration of single-cell DNA and RNA for analyzing intratumor heterogeneity using MaCroDNA
Here, the authors develop MaCroDNA, an algorithm to integrate single-cell DNA and RNA sequencing data from the same tissue. They use MaCroDNA to show—in agreement with previous studies—that copy number changes can predict progression from Barrett’s esophagus to esophageal adenocarcinoma.
- Mohammadamin Edrisi
- , Xiru Huang
- & Luay Nakhleh
-
Article
| Open Access GNTD: reconstructing spatial transcriptomes with graph-guided neural tensor decomposition informed by spatial and functional relations
GNTD: reconstructing spatial transcriptomes with graph-guided neural tensor decomposition informed by spatial and functional relations
Reconstructing transcriptome-wide spatially-resolved gene expressions requires modelling nonlinear patterns and spatial structures in RNA profiling data. Here, authors introduce a graph-guided neural hierarchical tensor decomposition model that incorporates spatial and functional relations for the task.
- Tianci Song
- , Charles Broadbent
- & Rui Kuang
-
Article
| Open Access An archetype and scaling of developmental tissue dynamics across species
An archetype and scaling of developmental tissue dynamics across species
Limb tissue dynamics until basic skeletal pattern establishment exhibit a high degree of conservation between chick and frog after proper rescaling of spacetime, suggesting the presence of a species-independent archetype of morphogenetic dynamics.
- Yoshihiro Morishita
- , Sang-Woo Lee
- & Aiko Kawasumi-Kita
-
Article
| Open Access UniKP: a unified framework for the prediction of enzyme kinetic parameters
UniKP: a unified framework for the prediction of enzyme kinetic parameters
Prediction of enzyme kinetic parameters is essential for designing and optimising enzymes for various biotechnological and industrial applications. Here, authors presented a prediction framework (UniKP), which improves the accuracy of predictions for three enzyme kinetic parameters.
- Han Yu
- , Huaxiang Deng
- & Xiaozhou Luo
-
Article
| Open Access STalign: Alignment of spatial transcriptomics data using diffeomorphic metric mapping
STalign: Alignment of spatial transcriptomics data using diffeomorphic metric mapping
Spatial transcriptomics (ST) enables gene expression characterisation within tissue sections, but comparing across sections and technologies remains challenging. Here, authors develop STalign to spatially align ST data and demonstrate applications including aligning to common coordinate frameworks.
- Kalen Clifton
- , Manjari Anant
- & Jean Fan
-
Article
| Open Access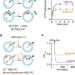 Phospholipids are imported into mitochondria by VDAC, a dimeric beta barrel scramblase
Phospholipids are imported into mitochondria by VDAC, a dimeric beta barrel scramblase
Mitochondria depend on phospholipids supplied by the endoplasmic reticulum. Here, using biochemical assays and molecular dynamics simulations, authors identify VDAC as a scramblase-type lipid transporter that catalyze lipid entry.
- Helene Jahn
- , Ladislav Bartoš
- & Anant K. Menon
-
Article
| Open Access Spatial transcriptomics deconvolution at single-cell resolution using Redeconve
Spatial transcriptomics deconvolution at single-cell resolution using Redeconve
Computational deconvolution with single-cell RNA sequencing data as a reference is pivotal for interpreting spatial transcriptomics data. Here, authors present Redeconve, which improves the resolution by more than 100-fold with higher accuracy and speed.
- Zixiang Zhou
- , Yunshan Zhong
- & Xianwen Ren
-
Article
| Open Access Engineered immunogens to elicit antibodies against conserved coronavirus epitopes
Engineered immunogens to elicit antibodies against conserved coronavirus epitopes
A pan-betacoronavirus vaccine will likely require the elicitation of antibodies against spike regions conserved across diverse coronaviruses. Here, authors computationally engineer and experimentally validate immunogens to elicit antibodies against two such spike regions.
- A. Brenda Kapingidza
- , Daniel J. Marston
- & Mihai L. Azoitei
-
Article
| Open Access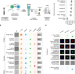 Explainable machine learning for profiling the immunological synapse and functional characterization of therapeutic antibodies
Explainable machine learning for profiling the immunological synapse and functional characterization of therapeutic antibodies
Therapeutic antibodies are crucial in treating severe diseases. Here, the authors introduce scifAI, an open-source explainable AI framework for analyzing imaging flow cytometry data, enabling rapid screening of therapeutic antibody candidates.
- Sayedali Shetab Boushehri
- , Katharina Essig
- & Fabian Schmich
-
Article
| Open Access TRS: a method for determining transcript termini from RNAtag-seq sequencing data
TRS: a method for determining transcript termini from RNAtag-seq sequencing data
TRS is a new method for determining 3’ transcript termini in bacteria, using data generated by the RNAtag-seq protocol. This methodology opens the door to study the evolution of transcription termini and their condition-dependent dynamics.
- Amir Bar
- , Liron Argaman
- & Hanah Margalit
-
Article
| Open Access Integrating spatial and single-cell transcriptomics data using deep generative models with SpatialScope
Integrating spatial and single-cell transcriptomics data using deep generative models with SpatialScope
Spatial transcriptomics (ST) is transforming tissue analysis but has limitations. Here, authors introduce SpatialScope, an integrated approach combining scRNA-seq and ST data using deep generative models, enabling comprehensive spatial characterisation at transcriptome-wide single-cell resolution.
- Xiaomeng Wan
- , Jiashun Xiao
- & Can Yang
-
Article
| Open Access PhenoSV: interpretable phenotype-aware model for the prioritization of genes affected by structural variants
PhenoSV: interpretable phenotype-aware model for the prioritization of genes affected by structural variants
Here, authors present PhenoSV, a phenotype-aware machine-learning model for the functional interpretation of various types of structural variants (SVs) and genes within or outside SVs, facilitating the extraction of biological insights from coding and noncoding SVs.
- Zhuoran Xu
- , Quan Li
- & Kai Wang
-
Article
| Open Access scDREAMER for atlas-level integration of single-cell datasets using deep generative model paired with adversarial classifier
scDREAMER for atlas-level integration of single-cell datasets using deep generative model paired with adversarial classifier
Integration of single-cell datasets is essential to gain a comprehensive understanding of complex biological systems. Here, the authors develop scDREAMER, a deep generative framework for performing unsupervised and supervised atlas-level integration, demonstrating improved bio-conservation and batch-correction.
- Ajita Shree
- , Musale Krushna Pavan
- & Hamim Zafar
-
Article
| Open Access SPACEL: deep learning-based characterization of spatial transcriptome architectures
SPACEL: deep learning-based characterization of spatial transcriptome architectures
Spatial transcriptomics (ST) technologies detect transcript distribution in space. Here, authors present a deep learning based method SPACEL for cell type deconvolution, spatial domain identification and 3D alignment, showcasing it as a valuable toolkit for ST data analysis
- Hao Xu
- , Shuyan Wang
- & Kun Qu
-
Article
| Open Access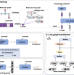 A knowledge-guided pre-training framework for improving molecular representation learning
A knowledge-guided pre-training framework for improving molecular representation learning
Accurate property prediction relies on effective molecular representation. Here, the authors introduce KPGT, a knowledge-guided self-supervised framework that improves molecular representation, leading to superior predictions of molecular properties and advancing AI-driven drug discovery.
- Han Li
- , Ruotian Zhang
- & Jianyang Zeng
-
Article
| Open Access Evaluation of the US COVID-19 Scenario Modeling Hub for informing pandemic response under uncertainty
Evaluation of the US COVID-19 Scenario Modeling Hub for informing pandemic response under uncertainty
The US COVID-19 Scenario Modeling Hub produced medium to long term projections based on different epidemic scenarios. In this study, the authors evaluate 14 rounds of projections by comparing them to the epidemic trajectories that occurred, and discuss lessons learned for future similar projects.
- Emily Howerton
- , Lucie Contamin
- & Justin Lessler
-
Article
| Open Access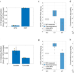 Inferring mitochondrial and cytosolic metabolism by coupling isotope tracing and deconvolution
Inferring mitochondrial and cytosolic metabolism by coupling isotope tracing and deconvolution
Studying metabolism in distinct subcellular compartments typically involves isolating organelles. Here, the authors demonstrate a quantitative approach to infer cytosolic and mitochondrial metabolic activities based on experiments with intact cells, maintaining physiological conditions.
- Alon Stern
- , Mariam Fokra
- & Tomer Shlomi
-
Article
| Open Access scReadSim: a single-cell RNA-seq and ATAC-seq read simulator
scReadSim: a single-cell RNA-seq and ATAC-seq read simulator
Benchmarking computational tools for analysis of single-cell sequencing data demands simulation of realistic sequencing reads. However, none of the few existing read simulators aim to mimic real data. Here, the authors introduce scReadSim, a single-cell RNA-seq and ATAC-seq read simulator that works by mimicking real data.
- Guanao Yan
- , Dongyuan Song
- & Jingyi Jessica Li
-
Article
| Open Access ProRefiner: an entropy-based refining strategy for inverse protein folding with global graph attention
ProRefiner: an entropy-based refining strategy for inverse protein folding with global graph attention
Inverse Protein Folding is a critical component of protein design. Here, authors introduce ProRefiner, a deep-learning model for IPF that exhibits both high performance and memory efficiency, thereby contributing to advancements in protein design.
- Xinyi Zhou
- , Guangyong Chen
- & Pheng Ann Heng
-
Article
| Open Access Deep learning of human polyadenylation sites at nucleotide resolution reveals molecular determinants of site usage and relevance in disease
Deep learning of human polyadenylation sites at nucleotide resolution reveals molecular determinants of site usage and relevance in disease
The authors develop deep learning models to identify genome-wide polyA sites at nucleotide resolution and calculate site strength. They further examine genomic parameters regulating site usage and reveal genetic variants altering polyA activity.
- Emily Kunce Stroup
- & Zhe Ji
-
Article
| Open Access Dimension-agnostic and granularity-based spatially variable gene identification using BSP
Dimension-agnostic and granularity-based spatially variable gene identification using BSP
Identifying spatially variable genes (SVGs) is essential for linking molecular cell functions with tissue phenotypes. Here, authors introduce a non-parametric model that detects SVGs from two or three-dimensional spatial transcriptomics data by comparing gene expression patterns at granularities.
- Juexin Wang
- , Jinpu Li
- & Dong Xu
-
Article
| Open Access CamoTSS: analysis of alternative transcription start sites for cellular phenotypes and regulatory patterns from 5' scRNA-seq data
CamoTSS: analysis of alternative transcription start sites for cellular phenotypes and regulatory patterns from 5' scRNA-seq data
Five-prime single-cell RNA-seq, especially the read 1, has precise capture of transcription start sites (TSS), but such information is often overlooked. Here, authors present a computational method suite, CamoTSS, to precisely identify TSS and quantify its expression, enabling effective detection of alternative TSS usage in different biological processes.
- Ruiyan Hou
- , Chung-Chau Hon
- & Yuanhua Huang
-
Article
| Open Access trRosettaRNA: automated prediction of RNA 3D structure with transformer network
trRosettaRNA: automated prediction of RNA 3D structure with transformer network
Here, authors develop trRosettaRNA, a deep learning-based approach for predicting RNA 3D structures. Blind tests demonstrate that the automated predictions compete effectively with top human predictions on natural RNAs.
- Wenkai Wang
- , Chenjie Feng
- & Jianyi Yang
-
Article
| Open Access Probabilities of developing HIV-1 bNAb sequence features in uninfected and chronically infected individuals
Probabilities of developing HIV-1 bNAb sequence features in uninfected and chronically infected individuals
Successful induction of broadly neutralizing antibodies is a main challenge in HIV vaccine development. The authors provide a framework to determine probabilities of antibody sequence development and show that uninfected and chronically infected individuals have the same chances to develop HIV-1 neutralizing antibodies.
- Christoph Kreer
- , Cosimo Lupo
- & Florian Klein
-
Article
| Open Access A generative adversarial network model alternative to animal studies for clinical pathology assessment
A generative adversarial network model alternative to animal studies for clinical pathology assessment
Generative AI has the potential to transform the way chemical and drug safety research is conducted. Here the authors show AnimalGAN, a model developed using Generative Adversarial Networks, which simulates virtual animal experiments to generate multidimensional rat clinical pathology measurements.
- Xi Chen
- , Ruth Roberts
- & Weida Tong
-
Article
| Open Access Dynamic characterization and interpretation for protein-RNA interactions across diverse cellular conditions using HDRNet
Dynamic characterization and interpretation for protein-RNA interactions across diverse cellular conditions using HDRNet
Predicting dynamic RNA-RBP interactions in diverse cell lines is an important challenge in unravelling RNA function and post-transcriptional regulatory mechanisms. Here, authors develop HDRNet, an end-to-end deep-learning-based framework for accurately predicting dynamic RBP binding events across various cellular conditions.
- Haoran Zhu
- , Yuning Yang
- & Xiangtao Li
-
Article
| Open Access Deep flanking sequence engineering for efficient promoter design using DeepSEED
Deep flanking sequence engineering for efficient promoter design using DeepSEED
Designing promoters with desired properties is crucial in synthetic biology. Here, authors introduce DeepSEED, an AI-aided flanking sequence optimisation framework which combines expert knowledge with deep learning techniques to efficiently design promoters in both eukaryotic and prokaryotic cells.
- Pengcheng Zhang
- , Haochen Wang
- & Xiaowo Wang
-
Article
| Open Access MetaCC allows scalable and integrative analyses of both long-read and short-read metagenomic Hi-C data
MetaCC allows scalable and integrative analyses of both long-read and short-read metagenomic Hi-C data
The authors develop an integrative and scalable framework to eliminate systematic biases and retrieve high-quality metagenome-assembled genomes using either long-read or short-read metagenomic Hi-C data.
- Yuxuan Du
- & Fengzhu Sun
-
Article
| Open Access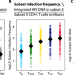 Estimating the contribution of CD4 T cell subset proliferation and differentiation to HIV persistence
Estimating the contribution of CD4 T cell subset proliferation and differentiation to HIV persistence
The authors used mathematical modeling of human data to study how HIV persists despite suppressive antiretroviral therapy. They found that when latently infected CD4+ T cells proliferate or differentiate, they can create HIV DNA and passage it into other subsets. More mature CD4 cell subsets then clear HIV DNA faster.
- Daniel B. Reeves
- , Charline Bacchus-Souffan
- & Peter W. Hunt
-
Article
| Open Access Patient-specific models link neurotransmitter receptor mechanisms with motor and visuospatial axes of Parkinson’s disease
Patient-specific models link neurotransmitter receptor mechanisms with motor and visuospatial axes of Parkinson’s disease
Neurotransmitter receptor distributions help explain structural and functional brain alterations in Parkinson’s disease. Distinct multi-receptor profiles are associated with the severity of motor, and visuospatial, psychiatric and memory symptoms.
- Ahmed Faraz Khan
- , Quadri Adewale
- & Yasser Iturria-Medina
-
Article
| Open Access TAGET: a toolkit for analyzing full-length transcripts from long-read sequencing
TAGET: a toolkit for analyzing full-length transcripts from long-read sequencing
Accurate long-read RNA sequencing facilitates analysis of full-length transcripts. Here the authors develop an integrative toolkit, optimised for Iso-Seq data analysis, that includes transcript alignment, annotation, quantification and gene fusion detection.
- Yuchao Xia
- , Zijie Jin
- & Ruibin Xi
-
Article
| Open Access Thermodynamic forces from protein and water govern condensate formation of an intrinsically disordered protein domain
Thermodynamic forces from protein and water govern condensate formation of an intrinsically disordered protein domain
In this work, the authors report atomistic molecular dynamics simulations showing that solvation entropy and protein-protein interactions are the main thermodynamic driving forces for the formation of condensates of the intrinsically disordered domain of the protein FUS.
- Saumyak Mukherjee
- & Lars V. Schäfer
-
Article
| Open Access epiAneufinder identifies copy number alterations from single-cell ATAC-seq data
epiAneufinder identifies copy number alterations from single-cell ATAC-seq data
'Here the authors present epiAneufinder, an algorithm for the identification of single-cell copy number alterations from scATAC-seq data, and explore the clonal heterogeneity in cell populations.
- Akshaya Ramakrishnan
- , Aikaterini Symeonidi
- & Maria Colomé-Tatché
-
Article
| Open Access SEPepQuant enhances the detection of possible isoform regulations in shotgun proteomics
SEPepQuant enhances the detection of possible isoform regulations in shotgun proteomics
Protein isoform quantification in shotgun proteomics is challenging due to the mapping of many peptides to multiple protein isoforms. Here, the authors present a computational method SEPepQuant and demonstrate its utility in revealing protein isoform level regulation in shotgun proteomics.
- Yongchao Dou
- , Yuejia Liu
- & Bing Zhang
-
Article
| Open Access The tumor microenvironment shows a hierarchy of cell-cell interactions dominated by fibroblasts
The tumor microenvironment shows a hierarchy of cell-cell interactions dominated by fibroblasts
The tumor microenvironment (TME) is complex and heterogenous, with cancer cells and diverse non-malignant cells interacting with each other. Here the authors define the network of interactions between different cell types in the TME of breast cancer, identifying and characterizing a two-cell circuit of cancer associated fibroblasts and macrophages.
- Shimrit Mayer
- , Tomer Milo
- & Ruth Scherz-Shouval
-
Article
| Open Access Natural plant growth and development achieved in the IPK PhenoSphere by dynamic environment simulation
Natural plant growth and development achieved in the IPK PhenoSphere by dynamic environment simulation
The PhenoSphere is a unique plant cultivation facility in which field-like environments can be simulated. Here, the authors find that a single season simulation is superior to an averaged season and to a climatized glasshouse cultivation to elicit field-like phenotypes evaluated in 11 maize lines.
- Marc C. Heuermann
- , Dominic Knoch
- & Thomas Altmann
-
Article
| Open Access Integrating end-to-end learning with deep geometrical potentials for ab initio RNA structure prediction
Integrating end-to-end learning with deep geometrical potentials for ab initio RNA structure prediction
Here the authors developed an open-source program (DRfold) for RNA tertiary structure prediction from sequence. Through a unique combination of end-to-end learning and geometry restraint guided simulations, the method demonstrates advantage over peer methods.
- Yang Li
- , Chengxin Zhang
- & Yang Zhang
-
Article
| Open Access Performance of tumour microenvironment deconvolution methods in breast cancer using single-cell simulated bulk mixtures
Performance of tumour microenvironment deconvolution methods in breast cancer using single-cell simulated bulk mixtures
Multiple computational approaches have been developed for the deconvolution of cells in the tumour microenvironment (TME) using bulk RNA-seq data. Here, the authors use breast cancer single-cell RNA-seq data to produce simulated bulk data, with which they compare the performance of nine TME deconvolution methods.
- Khoa A. Tran
- , Venkateswar Addala
- & Nicola Waddell
-
Article
| Open Access Local flux coordination and global gene expression regulation in metabolic modeling
Local flux coordination and global gene expression regulation in metabolic modeling
Genome-scale metabolic networks (GSMs) are a representation of a cell’s stoichiometrically balanced reactions. Here the authors report Decrem, a GSM reconstruction method, by integrating locally coupled reactions and global transcriptional regulation of metabolism by cell state.
- Gaoyang Li
- , Li Liu
- & Huansheng Cao
-
Article
| Open Access SiGra: single-cell spatial elucidation through an image-augmented graph transformer
SiGra: single-cell spatial elucidation through an image-augmented graph transformer
Recent advances have pushed spatial transcriptomics to subcellular resolution. Here, the authors propose SiGra, a graph artificial intelligence model designed for high-throughput spatial molecular imaging.
- Ziyang Tang
- , Zuotian Li
- & Qianqian Song
-
Article
| Open Access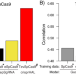 A generalizable Cas9/sgRNA prediction model using machine transfer learning with small high-quality datasets
A generalizable Cas9/sgRNA prediction model using machine transfer learning with small high-quality datasets
Current bacterial sgRNA activity models struggle with accurate predictions and generalizations. Here the authors report crisprHAL, a machine learning architecture that can be trained on existing datasets, and shows good sgRNA activity prediction accuracy can generalize predictions to different bacteria.
- Dalton T. Ham
- , Tyler S. Browne
- & David R. Edgell
-
Article
| Open Access Interpreting biologically informed neural networks for enhanced proteomic biomarker discovery and pathway analysis
Interpreting biologically informed neural networks for enhanced proteomic biomarker discovery and pathway analysis
Deep neural networks hold significant promise in capturing the complexity of biological systems. However, they suffer from a lack of interpretability. Here, authors present a generalizable method for developing, interpreting, and visualizing biologically informed neural networks for proteomics data.
- Erik Hartman
- , Aaron M. Scott
- & Johan Malmström
-
Article
| Open Access Addressing mechanism bias in model-based impact forecasts of new tuberculosis vaccines
Addressing mechanism bias in model-based impact forecasts of new tuberculosis vaccines
The complex transmission chain of tuberculosis (TB) forces mathematical modelers to make mechanistic assumptions when modelling vaccine effects. Here, authors posit a Bayesian formalism that unlocks mechanism-agnostic impact forecasts for TB vaccines.
- M. Tovar
- , Y. Moreno
- & J. Sanz

 MENDER: fast and scalable tissue structure identification in spatial omics data
MENDER: fast and scalable tissue structure identification in spatial omics data