Article
|
Open Access
Featured
-
-
Article
| Open Access Interplay between chromosomal alterations and gene mutations shapes the evolutionary trajectory of clonal hematopoiesis
Interplay between chromosomal alterations and gene mutations shapes the evolutionary trajectory of clonal hematopoiesis
Patients with solid cancers have high rates of clonal haematopoiesis associated with increased risk of secondary leukemias. Here, by using peripheral blood sequencing data from patients with solid non-hematologic cancer, the authors profile the landscape of mosaic chromosomal alterations and gene mutations, defining patients at high risk of leukemia progression.
- Teng Gao
- , Ryan Ptashkin
- & Elli Papaemmanuil
-
Article
| Open Access Inferring high-resolution human mixing patterns for disease modeling
Inferring high-resolution human mixing patterns for disease modeling
The growing need for realism in addressing complex public health questions calls for accurate models of the human contact patterns that govern disease transmission. Here, the authors generate effective population-level contact matrices by using highly detailed macro (census) and micro (survey) data on key socio-demographic features.
- Dina Mistry
- , Maria Litvinova
- & Alessandro Vespignani
-
Article
| Open Access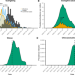 Estimating internationally imported cases during the early COVID-19 pandemic
Estimating internationally imported cases during the early COVID-19 pandemic
Sparse testing early in the SARS-CoV-2 pandemic hinders estimation of the dates and origins of initial case importations. Here, the authors show that the main source of cases imported from China shifted from Wuhan to other Chinese cities by mid-February, especially for African locations.
- Tigist F. Menkir
- , Taylor Chin
- & Rene Niehus
-
Article
| Open Access Accurate protein structure prediction with hydroxyl radical protein footprinting data
Accurate protein structure prediction with hydroxyl radical protein footprinting data
Mass spectrometry-based covalent labeling techniques such as hydroxyl radical protein footprinting (HRPF) provide information about protein tertiary structures. Here, the authors present a dynamics driven HRPF-guided algorithm for protein structure prediction that is incorporated in the Rosetta software suite and only requires the protein sequence and HRPF data as input and they demonstrate its successful application to four benchmark proteins.
- Sarah E. Biehn
- & Steffen Lindert
-
Article
| Open Access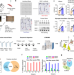 Deep muscle-proteomic analysis of freeze-dried human muscle biopsies reveals fiber type-specific adaptations to exercise training
Deep muscle-proteomic analysis of freeze-dried human muscle biopsies reveals fiber type-specific adaptations to exercise training
Skeletal muscle conveys the beneficial effects of physical exercise but due to its heterogeneity, studying the effects of exercise on muscle fibres is challenging. Here, the authors carry out proteomic analysis of myofibres from freeze-dried muscle biopsies, show fibre-type specific changes in response to exercise, and show that the oxidative and glycolytic muscle fibers adapt differentially to exercise training.
- A. S. Deshmukh
- , D. E. Steenberg
- & J. F. P. Wojtaszewski
-
Article
| Open Access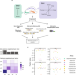 Differential contribution of transcriptomic regulatory layers in the definition of neuronal identity
Differential contribution of transcriptomic regulatory layers in the definition of neuronal identity
Post-transcriptional gene regulation is an important contributor to cell type-specific differences at the transcriptomic level. Here, the authors use a multiomics approach to characterize neuronal diversity in the mouse nervous system, analyzing the relative contributions of multiple layers of transcriptomic regulation in the specification of cell type identity.
- Kevin C. H. Ha
- , Timothy Sterne-Weiler
- & Benjamin J. Blencowe
-
Article
| Open Access Mathematical modeling of COVID-19 in 14.8 million individuals in Bahia, Brazil
Mathematical modeling of COVID-19 in 14.8 million individuals in Bahia, Brazil
Low-resource settings can face additional challenges in managing the COVID-19 pandemic. Here, the authors use mathematical modelling to investigate transmission in the state of Bahia, Brazil, and quantify control measures needed to prevent the hospital system becoming overwhelmed.
- Juliane F. Oliveira
- , Daniel C. P. Jorge
- & Roberto F. S. Andrade
-
Article
| Open Access SCISSOR: a framework for identifying structural changes in RNA transcripts
SCISSOR: a framework for identifying structural changes in RNA transcripts
Many computational tools identify mRNA variations by analyzing the transformed RNA-seq data such as collapsed reads. Here, the authors report a computational method which uses shape changes in the RNA-seq coverage profile to discover changes in mRNA expression and alternative splicing.
- Hyo Young Choi
- , Heejoon Jo
- & D. Neil Hayes
-
Article
| Open Access Bidirectional contact tracing could dramatically improve COVID-19 control
Bidirectional contact tracing could dramatically improve COVID-19 control
Contact tracing is critical to controlling COVID-19, but most protocols only “forward-trace” to notify people who were recently exposed. Using a stochastic branching-process model, the authors show that “bidirectional” tracing to identify infector individuals and their other infectees robustly improves outbreak control.
- William J. Bradshaw
- , Ethan C. Alley
- & Kevin M. Esvelt
-
Article
| Open Access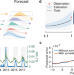 Optimizing respiratory virus surveillance networks using uncertainty propagation
Optimizing respiratory virus surveillance networks using uncertainty propagation
Lack of a widespread surveillance network hampers accurate infectious disease forecasting. Here the authors provide a framework to optimize the selection of surveillance site locations and show that accurate forecasting of respiratory diseases for locations without surveillance is feasible.
- Sen Pei
- , Xian Teng
- & Jeffrey Shaman
-
Article
| Open Access Anti-senescent drug screening by deep learning-based morphology senescence scoring
Anti-senescent drug screening by deep learning-based morphology senescence scoring
Cellular senescence is a hallmark of ageing and is important for the pathogenesis of ageing-related diseases. Here, the authors develop a morphology-based deep learning system to identify senescent cells and a quantitative scoring system to evaluate the state of endothelial cells to evaluate the effects of anti-senescent reagents.
- Dai Kusumoto
- , Tomohisa Seki
- & Shinsuke Yuasa
-
Article
| Open Access Bayesian genome scale modelling identifies thermal determinants of yeast metabolism
Bayesian genome scale modelling identifies thermal determinants of yeast metabolism
While temperature impacts the function of all cellular components, it’s hard to rule out how the temperature dependence of cell phenotypes emerged from the dependence of individual components. Here, the authors develop a Bayesian genome scale modelling approach to identify thermal determinants of yeast metabolism.
- Gang Li
- , Yating Hu
- & Jens Nielsen
-
Article
| Open Access Lossless integration of multiple electronic health records for identifying pleiotropy using summary statistics
Lossless integration of multiple electronic health records for identifying pleiotropy using summary statistics
Thus far, pleiotropy analysis using individual-level Electronic Health Records data has been limited to data from one site. Here, the authors introduce Sum-Share, a method designed to efficiently and losslessly integrate EHR and genetic data from multiple sites to perform pleiotropy analysis.
- Ruowang Li
- , Rui Duan
- & Jason H. Moore
-
Article
| Open Access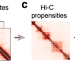 High-resolution single-cell 3D-models of chromatin ensembles during Drosophila embryogenesis
High-resolution single-cell 3D-models of chromatin ensembles during Drosophila embryogenesis
Balancing high resolution and broad genome coverage in single-cell Hi-C approaches remains challenging. Here, the authors describe a computational method for the reconstruction of a large 3D-ensemble of single-cell chromatin conformations from population Hi-C measurements and apply this model to study embryogenesis in Drosophila.
- Qiu Sun
- , Alan Perez-Rathke
- & Jie Liang
-
Article
| Open Access Integrated analysis of telomerase enzymatic activity unravels an association with cancer stemness and proliferation
Integrated analysis of telomerase enzymatic activity unravels an association with cancer stemness and proliferation
Telomerase activity correlates with distinct cell states, but can be challenging to quantify. Here the authors quantify telomerase activity across a range of biological samples using the expression of 13 genes, and show it correlates with cancer cell proliferation and stemness.
- Nighat Noureen
- , Shaofang Wu
- & Siyuan Zheng
-
Article
| Open Access Survey data and human computation for improved flu tracking
Survey data and human computation for improved flu tracking
Digital trace data from search engines lacks information about the experiences of the individuals generating the data. Here the authors link search data and human computation to build a tracking model of influenza-like illness.
- Stefan Wojcik
- , Avleen S. Bijral
- & David Lazer
-
Article
| Open Access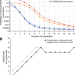 Optimal COVID-19 quarantine and testing strategies
Optimal COVID-19 quarantine and testing strategies
Safely reducing the necessary duration of quarantine for COVID-19 could lessen the economic impacts of the pandemic. Here, the authors demonstrate that testing on exit from quarantine is more effective than testing on entry, and can enable quarantine to be reduced from fourteen to seven days.
- Chad R. Wells
- , Jeffrey P. Townsend
- & Alison P. Galvani
-
Article
| Open Access Benchmarking joint multi-omics dimensionality reduction approaches for the study of cancer
Benchmarking joint multi-omics dimensionality reduction approaches for the study of cancer
Advances in omics technology have resulted in the generation of multi-view data for cancer samples. Here, the authors compare dimensionality reduction techniques using simulated and TCGA data and identify the features of the methods with superior performance.
- Laura Cantini
- , Pooya Zakeri
- & Anaïs Baudot
-
Article
| Open Access The role of water in host-guest interaction
The role of water in host-guest interaction
Computational approaches to predict water’s role in host-ligand binding attract a great deal of attention. Here the authors use a metadynamics enhanced sampling method and machine learning to compute binding energies for host-guest systems from the SAMPL5 challenge and provide details of water structural changes.
- Valerio Rizzi
- , Luigi Bonati
- & Michele Parrinello
-
Article
| Open Access Multi-domain translation between single-cell imaging and sequencing data using autoencoders
Multi-domain translation between single-cell imaging and sequencing data using autoencoders
Integration of single cell data modalities increases the richness of information about the heterogeneity of cell states, but integration of imaging and transcriptomics is an open challenge. Here the authors use autoencoders to learn a probabilistic coupling and map these modalities to a shared latent space.
- Karren Dai Yang
- , Anastasiya Belyaeva
- & Caroline Uhler
-
Article
| Open Access Global computational alignment of tumor and cell line transcriptional profiles
Global computational alignment of tumor and cell line transcriptional profiles
The determination of whether cancer cell lines recapitulate the molecular features of corresponding patient tumours remains essential for the selection of appropriate cell line models for preclinical studies. The method developed here, Celligner, integrates cancer cell line and tumour RNA-seq datasets and reveals large differences in their concordance across cell lines and cancer types.
- Allison Warren
- , Yejia Chen
- & James M. McFarland
-
Article
| Open Access Efficient assembly of nanopore reads via highly accurate and intact error correction
Efficient assembly of nanopore reads via highly accurate and intact error correction
Nanopore reads have been advantageous for de novo genome assembly; however these reads have high error rates. Here, the authors develop an error correction and de novo assembly tool, NECAT, which produces efficient, high quality assemblies of nanopore reads.
- Ying Chen
- , Fan Nie
- & Chuan-Le Xiao
-
Article
| Open Access The aging transcriptome and cellular landscape of the human lung in relation to SARS-CoV-2
The aging transcriptome and cellular landscape of the human lung in relation to SARS-CoV-2
Age is one of the strongest risk factors for severe illness from COVID-19. By integrating human lung transcriptomes with experimental data on SARS-CoV-2, the authors pinpoint specific age-associated factors that could contribute to the heightened severity of COVID-19 in older populations.
- Ryan D. Chow
- , Medha Majety
- & Sidi Chen
-
Article
| Open Access Long first exons and epigenetic marks distinguish conserved pachytene piRNA clusters from other mammalian genes
Long first exons and epigenetic marks distinguish conserved pachytene piRNA clusters from other mammalian genes
The pachytene piRNA loci are transcribed by RNA polymerase II in the male germline of placental mammals. Here the authors show that a long first exon or a long unspliced transcript correlates with germline-specific production of piRNA precursor transcripts and mature piRNAs.
- Tianxiong Yu
- , Kaili Fan
- & Zhiping Weng
-
Article
| Open Access Error correction enables use of Oxford Nanopore technology for reference-free transcriptome analysis
Error correction enables use of Oxford Nanopore technology for reference-free transcriptome analysis
Nanopore sequencing technologies applied to transcriptome analysis suffer from high error rates, limiting them largely to reference-based analyses. Here, the authors develop a computational error correction method for transcriptome analysis that reduces the median error rate from ~7% to ~1%.
- Kristoffer Sahlin
- & Paul Medvedev
-
Article
| Open Access Humanizing the yeast origin recognition complex
Humanizing the yeast origin recognition complex
In most model yeast species the Origin Recognition Complex (ORC) binds defined and species-specific base sequences while in humans what determines the binding appears to be more complex. Here the authors reveal that the yeast’s ORC complex binding specificity is dependent on a 19-amino acid insertion helix in the Orc4 subunit which is lost in human.
- Clare S. K. Lee
- , Ming Fung Cheung
- & Bik-Kwoon Tye
-
Article
| Open Access Loss-of-function genomic variants highlight potential therapeutic targets for cardiovascular disease
Loss-of-function genomic variants highlight potential therapeutic targets for cardiovascular disease
Drugs targeting cardiovascular disease (CVD) can have negative consequences for liver function. Here, the authors combine genome wide analyses on 69,479 individuals to identify loss-of-function variants with beneficial effects on CVD-related traits without negative impacts on liver function.
- Jonas B. Nielsen
- , Oren Rom
- & Kristian Hveem
-
Article
| Open Access A collection of bacterial isolates from the pig intestine reveals functional and taxonomic diversity
A collection of bacterial isolates from the pig intestine reveals functional and taxonomic diversity
The authors present a public collection of 117 bacterial isolates from the pig gut, including the description of 38 novel taxa. Interesting functions discovered in these organisms include a new fucosyltransferease and sactipeptide-like molecules encoded by biosynthetic gene clusters.
- David Wylensek
- , Thomas C. A. Hitch
- & Thomas Clavel
-
Article
| Open Access A meta-learning approach for genomic survival analysis
A meta-learning approach for genomic survival analysis
RNA-sequencing data from tumours can be used to predict the prognosis of patients. Here, the authors show that a neural network meta-learning approach can be useful for predicting prognosis from a small number of samples.
- Yeping Lina Qiu
- , Hong Zheng
- & Olivier Gevaert
-
Article
| Open Access Deep learning-based cross-classifications reveal conserved spatial behaviors within tumor histological images
Deep learning-based cross-classifications reveal conserved spatial behaviors within tumor histological images
Histopathological images are a rich but incompletely explored data type for studying cancer. Here the authors show that convolutional neural networks can be systematically applied across cancer types, enabling comparisons to reveal shared spatial behaviors.
- Javad Noorbakhsh
- , Saman Farahmand
- & Jeffrey H. Chuang
-
Article
| Open Access Machine learning uncovers independently regulated modules in the Bacillus subtilis transcriptome
Machine learning uncovers independently regulated modules in the Bacillus subtilis transcriptome
The systems-level regulatory structure underlying gene expression in bacteria can be inferred using machine learning algorithms. Here we show this structure for Bacillus subtilis, present five hypotheses gleaned from it, and analyse the process of sporulation from its perspective.
- Kevin Rychel
- , Anand V. Sastry
- & Bernhard O. Palsson
-
Article
| Open Access Uptake of monoaromatic hydrocarbons during biodegradation by FadL channel-mediated lateral diffusion
Uptake of monoaromatic hydrocarbons during biodegradation by FadL channel-mediated lateral diffusion
The uptake of hydrophobic molecules by bacterial FadL channels is implicated in quorum sensing, interactions with eukaryotic hosts and biodegradation of many pollutants. Insights into monoaromatic hydrocarbon uptake by TodX and CymD channels suggest that all FadL channels mediate substrate uptake via lateral diffusion.
- Kamolrat Somboon
- , Anne Doble
- & Bert van den Berg
-
Article
| Open Access Scalable multiple whole-genome alignment and locally collinear block construction with SibeliaZ
Scalable multiple whole-genome alignment and locally collinear block construction with SibeliaZ
Multiple whole-genome alignment is a challenging problem in bioinformatics, especially when computational resources are limited. Here the authors present SibeliaZ, an algorithm and software based on analysis of de Bruijn graphs, which provides improved computational efficiency and scalability.
- Ilia Minkin
- & Paul Medvedev
-
Article
| Open Access A machine learning toolkit for genetic engineering attribution to facilitate biosecurity
A machine learning toolkit for genetic engineering attribution to facilitate biosecurity
The potential for accidental or deliberate misuse of biotechnology is of concern for international biosecurity. Here the authors apply machine learning to DNA sequences and associated phenotypic data to facilitate genetic engineering attribution and identify country-of-origin and ancestral lab of engineered DNA sequences.
- Ethan C. Alley
- , Miles Turpin
- & Kevin M. Esvelt
-
Article
| Open Access Improving the informativeness of Mendelian disease-derived pathogenicity scores for common disease
Improving the informativeness of Mendelian disease-derived pathogenicity scores for common disease
Pathogenicity scores are instrumental in prioritizing variants for Mendelian disease, yet their application to common disease is largely unexplored. Here, the authors assess the utility of pathogenicity scores for 41 complex traits and develop a framework to improve their informativeness for common disease.
- Samuel S. Kim
- , Kushal K. Dey
- & Alkes L. Price
-
Article
| Open Access Establishment of a morphological atlas of the Caenorhabditis elegans embryo using deep-learning-based 4D segmentation
Establishment of a morphological atlas of the Caenorhabditis elegans embryo using deep-learning-based 4D segmentation
The systematic characterization of C. elegans morphology during development has yet to be performed. Here, the authors produce a 3D atlas of C. elegans morphology from 17 embryos and 54 developmental stages, using an automated pipeline, CShaper (combining segmentation of fluorescently labeled membranes with automated cell lineage tracing).
- Jianfeng Cao
- , Guoye Guan
- & Hong Yan
-
Article
| Open Access Nonlinear mechanics of lamin filaments and the meshwork topology build an emergent nuclear lamina
Nonlinear mechanics of lamin filaments and the meshwork topology build an emergent nuclear lamina
Mechanical strength of in situ assembled nuclear lamin filaments arranged in a 3D meshwork is unclear. Here, using mechanical, structural and simulation tools, the authors report the hierarchical organization of the lamin meshwork that imparts strength and toughness to lamin filaments at par with silk and Kevlar®
- K. Tanuj Sapra
- , Zhao Qin
- & Ohad Medalia
-
Article
| Open Access Deciphering functional redundancy in the human microbiome
Deciphering functional redundancy in the human microbiome
Here, the authors develop a genome evolution model to investigate the origin of functional redundancy in the human microbiome by analyzing its genomic content network and illustrate potential ecological and evolutionary processes that may contribute to its resilience.
- Liang Tian
- , Xu-Wen Wang
- & Yang-Yu Liu
-
Article
| Open Access Integrative modeling of membrane-associated protein assemblies
Integrative modeling of membrane-associated protein assemblies
Most approaches for modeling the membrane protein complexes are not capable of incorporating the topological information provided by the membrane. Here authors present an integrative computational protocol for the modeling of membrane-associated protein assemblies, specifically complexes consisting of a membrane-embedded protein and a soluble partner.
- Jorge Roel-Touris
- , Brian Jiménez-García
- & Alexandre M. J. J. Bonvin
-
Article
| Open Access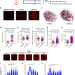 TGFβ promotes widespread enhancer chromatin opening and operates on genomic regulatory domains
TGFβ promotes widespread enhancer chromatin opening and operates on genomic regulatory domains
The TGFβ signaling pathway has been shown to regulate transcription by regulating enhancer activity. Here, the authors perform a comprehensive analysis of enhancers in normal mammary epithelial gland cells to elucidate how TGFβ-dependent enhancers control gene transcription in these cells.
- Jose A. Guerrero-Martínez
- , María Ceballos-Chávez
- & Jose C. Reyes
-
Article
| Open Access State-level tracking of COVID-19 in the United States
State-level tracking of COVID-19 in the United States
High numbers of COVID-19-related deaths have been reported in the United States, but estimation of the true numbers of infections is challenging. Here, the authors estimate that on 1 June 2020, 3.7% of the US population was infected with SARS-CoV-2, and 0.01% was infectious, with wide variation by state.
- H. Juliette T. Unwin
- , Swapnil Mishra
- & Seth Flaxman
-
Article
| Open Access Structural modularity of the XIST ribonucleoprotein complex
Structural modularity of the XIST ribonucleoprotein complex
The long noncoding RNA XIST plays a central role in sex-specific gene expression in humans by silencing one of two X chromosomes in female cells. Here the authors show that higher order secondary structure creates the modular domain structure of XIST ribonucleoprotein complex and spatial separation of functions.
- Zhipeng Lu
- , Jimmy K. Guo
- & Howard Y. Chang
-
Article
| Open Access Identification and characterization of constrained non-exonic bases lacking predictive epigenomic and transcription factor binding annotations
Identification and characterization of constrained non-exonic bases lacking predictive epigenomic and transcription factor binding annotations
Genome-wide maps of evolutionary constraint and large-scale compendia of epigenomic and transcription factor data provide complementary information for genome annotation. Here, the authors develop the Constrained Non-Exonic Predictor (CNEP) that enables better understanding of their relationship.
- Olivera Grujic
- , Tanya N. Phung
- & Jason Ernst
-
Perspective
| Open Access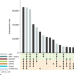 Towards a unified open access dataset of molecular interactions
Towards a unified open access dataset of molecular interactions
The IMEx consortium provides one of the largest resources of curated, experimentally verified molecular interaction data. Here, the authors review how IMEx evolved into a fundamental resource for life scientists and describe how IMEx data can support biomedical research.
- Pablo Porras
- , Elisabet Barrera
- & Sandra Orchard
-
Article
| Open Access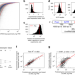 Deep learning suggests that gene expression is encoded in all parts of a co-evolving interacting gene regulatory structure
Deep learning suggests that gene expression is encoded in all parts of a co-evolving interacting gene regulatory structure
Regulatory and coding regions of genes are shaped by evolution to control expression levels. Here, the authors use deep learning to identify rules controlling gene expression levels and suggest that all parts of the gene regulatory structure interact in this.
- Jan Zrimec
- , Christoph S. Börlin
- & Aleksej Zelezniak
-
Article
| Open Access Leveraging multi-way interactions for systematic prediction of pre-clinical drug combination effects
Leveraging multi-way interactions for systematic prediction of pre-clinical drug combination effects
Combinatorial treatments have become a standard of care for various complex diseases including cancers. Here, the authors show that combinatorial responses of two anticancer drugs can be accurately predicted using factorization machines trained on large-scale pharmacogenomic data for guiding precision oncology studies.
- Heli Julkunen
- , Anna Cichonska
- & Juho Rousu
-
Article
| Open Access An investigation of irreproducibility in maximum likelihood phylogenetic inference
An investigation of irreproducibility in maximum likelihood phylogenetic inference
Replicate runs of maximum likelihood phylogenetic analyses can generate different tree topologies due to differences in parameters, such as random seeds. Here, Shen et al. demonstrate that replicate runs can generate substantially different tree topologies even with identical data and parameters.
- Xing-Xing Shen
- , Yuanning Li
- & Antonis Rokas
-
Article
| Open Access muscat detects subpopulation-specific state transitions from multi-sample multi-condition single-cell transcriptomics data
muscat detects subpopulation-specific state transitions from multi-sample multi-condition single-cell transcriptomics data
Single-cell transcriptomics enhanced our ability to profile heterogeneous cell populations. It is not known which statistical frameworks are performant to detect subpopulation-level responses. Here, the authors developed a simulation framework to evaluate various methods across a range of scenarios.
- Helena L. Crowell
- , Charlotte Soneson
- & Mark D. Robinson
-
Article
| Open Access A clinically applicable deep-learning model for detecting intracranial aneurysm in computed tomography angiography images
A clinically applicable deep-learning model for detecting intracranial aneurysm in computed tomography angiography images
Interpretation of Computed Tomography Angiography for intracranial aneurysm diagnosis can be time-consuming and challenging. Here, the authors present a deep-learning-based framework achieving improved performance compared to that of radiologists and expert neurosurgeons.
- Zhao Shi
- , Chongchang Miao
- & Long Jiang Zhang
Browse broader subjects
Browse narrower subjects
- Biochemical reaction networks
- Cellular signalling networks
- Classification and taxonomy
- Communication and replication
- Computational models
- Computational neuroscience
- Computational platforms and environments
- Data acquisition
- Data integration
- Data mining
- Data processing
- Data publication and archiving
- Databases
- Functional clustering
- Gene ontology
- Gene regulatory networks
- Genome informatics
- Hardware and infrastructure
- High-throughput screening
- Image processing
- Literature mining
- Machine learning
- Microarrays
- Network topology
- Phylogeny
- Power law
- Predictive medicine
- Probabilistic data networks
- Programming language
- Protein analysis
- Protein design
- Protein folding
- Protein function predictions
- Protein structure predictions
- Proteome informatics
- Quality control
- Scale invariance
- Sequence annotation
- Software
- Standards
- Statistical methods
- Virtual drug screening

 isoCirc catalogs full-length circular RNA isoforms in human transcriptomes
isoCirc catalogs full-length circular RNA isoforms in human transcriptomes